THE LIVING WORLD
Unit Three. The Continuity of Life
12.9. Complex Regulation of Gene Expression
As you have seen, the eukaryotic gene is structurally more complex than the prokaryotic gene, and the regulation of gene expression is also more complex. Eukaryotic gene expression is controlled at many stages, reviewed in figure 12.20. Chromatin structure can affect gene expression by determining if a gene will be accessible to RNA polymerase. The availability of many factors influences the rate at which a particular gene is transcribed. Once transcribed, a gene’s expression can be altered by alternative splicing, or it can be silenced by RNA interference. Although gene regulation often occurs earlier in the process of gene expression, some control mechanisms act later. The availability of translational proteins can affect protein synthesis, and a protein can also be chemically modified after it is produced.
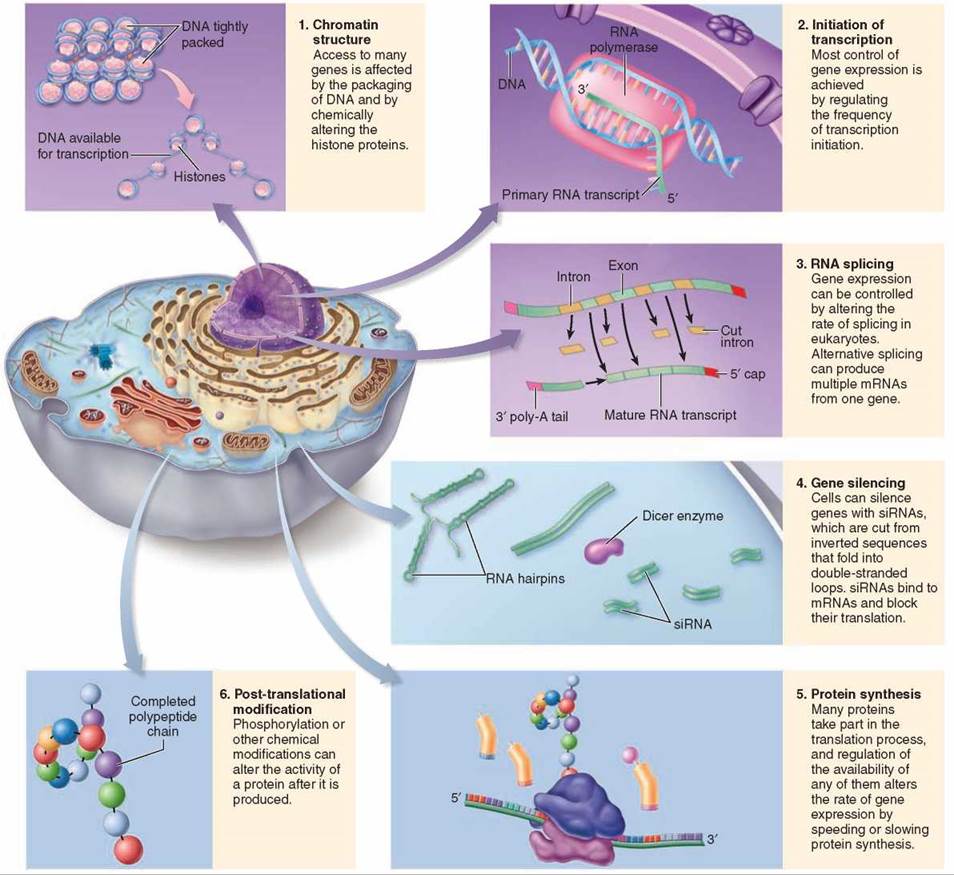
Figure 12.20. The control of gene expression in eukaryotes.
Key Learning Outcome 12.9. A eukaryotic gene is controlled at many points during gene expression.
Inquiry & Analysis
Building Proteins in a Test Tube
The complex mechanisms used by cells to build proteins were not discovered all at once.
Our understanding came slowly, accumulating through a long series of experiments, each telling us a little bit more. To gain some sense of the incremental nature of this experimental journey, and to appreciate the excitement that each step gave, it is useful to step into the shoes of an investigator back when little was known and the way forward was not clear.
The shoes we will step into are those of Paul Zamecnik, an early pioneer in protein synthesis research. Working with colleagues at Massachusetts General Hospital in the early 1950s, Zamecnik first asked the most direct of questions: Where in the cell are proteins synthesized? To find out, they injected radioactive amino acids into rats. After a few hours, the labeled amino acids could be found as part of newly made proteins in the livers of the rats. And, if the livers were removed and checked only minutes after injection, radioactive-labeled proteins were found only associated with small particles in the cytoplasm. Composed of protein and RNA, these particles, later named ribosomes, had been discovered years earlier by electron microscope studies of cell components. This experiment identified them as the sites of protein synthesis in the cell.
After several years of trial-and-error tinkering, Zamecnik and his colleagues had worked out a "cell-free” protein-synthesis system that would lead to the synthesis of proteins in a test tube. It included ribosomes, mRNA, and ATP to provide energy. It also included a collection of required soluble "factors” isolated from homogenized rat cells that somehow worked with the ribosome to get the job done. When Zamecnik's team characterized these required factors, they found most of them to be proteins, as expected, but also present in the mix was a small RNA, very unexpected.
To see what this small RNA was doing, they performed the following experiment. In a test tube, they added various amounts of 14C-leucine (that is, the radioactively labeled amino acid leucine) to the cell-free system containing the soluble factors, ribosomes, and ATP. After waiting a bit, they then isolated the small RNA from the mixture and checked it for radioactivity. You can see the results in graph (a).
In a follow-up experiment, they mixed the radioactive leucine-small RNA complex that this experiment had generated with cell extracts containing intact endoplasmic reticulum (that is, a cell system of ribosomes on membranes quite capable of making protein). Looking to see where the radioactive label now went, they then isolated the newly made protein as well as the small RNA [see graph (b)].
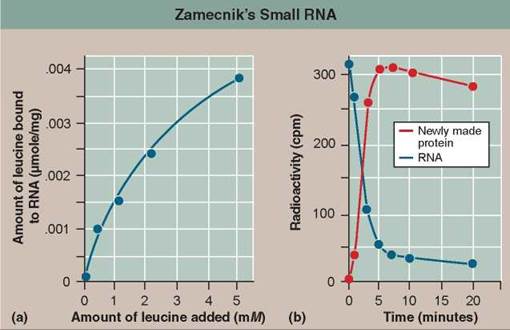
EXPERIMENT A, shown in graph (a)
1. Applying Concepts. What is the dependent variable?
2. Interpreting Data. Does the amount of leucine added to the test tube have an effect on the amount of leucine found bound to the small RNA?
3. Making Inferences. Is the amount of leucine bound to small RNA proportional to the amount of leucine added to the mixture?
4. Drawing Conclusions. Can you reasonably conclude from this result that the amino acid leucine is binding to the small RNA?
EXPERIMENT B, shown in graph (b)
1. Applying Concepts. What is the dependent variable?
2. Interpreting Data
a. Monitoring radioactivity for 20 minutes after the addition of the radioactive leucine-small RNA complex to the cell extract, what happens to the level of radioactivity in the small RNA (blue)?
b. Over the same period, what happens to the level of radioactivity in the newly made protein (red)?
3. Making Inferences. Is the same amount of radioactivity being lost from the small RNA that is being gained by the newly made protein?
4. Drawing Conclusions
a. Is it reasonable to conclude that the small RNA is donating its amino acid to the growing protein? [Hint: As a result of this experiment, the small RNA was called transfer RNA.]
b. If you were to isolate the protein from this experiment made after 20 minutes, which amino acids would be radioactively labeled? Explain.
1. Which of the following is not a type of RNA?
a. nRNA (nuclear RNA)
b. mRNA (messenger RNA)
c. rRNA (ribosomal RNA)
d. tRNA (transfer RNA)
2. Each amino acid in a polypeptide is specified by
a. an enhancer.
b. a promoter.
c. an rRNA molecule.
d. a codon.
3. The three-nucleotide codon system can be arranged into ______ combinations.
a. 16
b. 20
c. 64
d. 128
4. The process of obtaining a copy of the information in a gene as a strand of messenger RNA is called
a. polymerase.
b. expression.
c. transcription.
d. translation.
5. The site where RNA polymerase attaches to the DNA molecule to start the formation of an RNA molecule is called a(n)
a. promoter.
b. exon.
c. intron.
d. enhancer.
6. The process of taking the information on a strand of messenger RNA and building an amino acid chain, which will become all or part of a protein molecule, is called
a. polymerase.
b. expression.
c. transcription.
d. translation.
7. If an mRNA codon reads UAC, its complementary anticodon will be
a. TUC.
b. ATG.
c. AUG.
d. CAG.
8. Which of the following accurately describes gene expression in prokaryotic cells?
a. All genes are on all the time in all cells, making the needed amino acid sequences.
b. Some genes are always off unless a promoter turns them on.
c. Some genes are always on unless a promoter turns them off.
d. Some genes remain off as long as a repressor is bound.
9. Which of the following statements is correct about eukaryotic gene expression?
a. mRNAs must have introns spliced out.
b. mRNAs contain the transcript of only one gene.
c. Enhancers act from a distance.
d. All of the above.
10. Which of the following is not a mechanism of controlling gene expression in eukaryotic cells?
a. blocking translation with siRNA
b. activating an enhancer
c. translating a gene as it is being transcribed
d. alternative splicing of the primary RNA transcript
Synthesize What You Have Learned
1. On the television program The X Files, Agent Scully discovers an extraterrestrial life form that has a DNA genome like ours, but with a four-letter genetic code instead of the triplet genetic code that we earthly inhabitants possess. How many different amino acids would this extraterrestrial code be able to specify (assuming there were that many kinds of amino acids available on the extraterrestrial planet)? Why do you think the terrestrial code allows 64 combinations, when only 22 amino acids are common here on earth? Why do you think only 20 amino acids are common in proteins here on earth?
2. What would happen if all the genes in a cell were always active?
3. The nucleotide sequence of a hypothetical gene is: TACATACTTAGTTACGTCGCCCGGAAATAT
a. What will be the sequence on the mRNA when it is transcribed?
b. What will be the amino acid sequence of the protein when it’s translated?
c. What would happen to the amino acid chain if the highlighted nucleotide underwent a mutation and was changed to an A nucleotide?