CHEMICAL BIOLOGY
Taste: Topics in Chemical Biology
Maik Behrens, Frauke Stahler, Peng Shi, Bernd Bute and Wolfgang Meyerhof, Department of Molecular Genetics, German Institute of Human Nutrition Potsdam-Rehbruecke, Nuthetal, Germany
doi: 10.1002/9780470048672.wecb592
The mammalian sense of taste is crucial for evaluating food palatability and nutritional quality. To achieve the detection of relevant chemicals, evolution has shaped a set of receptor molecules that allow the detection of five basic taste qualities: sweet, umami, bitter, salty, and sour. Each taste modality has unique characteristics and serves a distinct function for an animal's nutrition. Therefore, we address the characteristics, the recent advances, the persisting difficulties, and the future perspectives of the basic taste qualities in separate paragraphs. Enormous progress has been made in the identification and the characterization of taste receptor molecules, and in the growing number of animal genomes accessible from databases, which has inspired us to devote a section of this review to discuss a series of sophisticated evolutionary studies on taste receptor molecules. Finally, evidence is accumulating to show that taste receptor and signal transduction molecules have nongustatory functions as well. The extragustatory expression of such genes and the resulting implications are summarized in the final section.
The chemical senses of olfaction and gustation were developed from phylogenetically old systems that enabled organisms to detect chemicals in their environment. The detectors were linked to behavioral patterns that enabled the organisms to escape noxious substances or to approach potential nutrients. These stereotypic mechanisms are still present in higher organisms, including mammals, although they compose more complex regulatory loops. The chemical sense of taste allows mammals to evaluate the food they consume. Each of the five basic taste modalities fulfils a particular task. Sweet and umami (glutamate and 5'-ribonucleotides) taste detects calorie-rich food that contains carbohydrates or protein. Therefore, both sweet and unami tastes are linked to pleasant feelings and to behaviors that facilitate food intake. Salty taste is part of a control loop that underlies electrolyte homeostasis. Salt intake compensates for salt loss through sweating and elimination. Like umami and sweet taste, salty taste is linked to liking and attraction promoting intake. Sour and bitter tastes are repulsive and seem to be part of a warning system. Sour taste prevents excessive intake of protons and balances the acid and bases in the body; also it prevents intoxication through consumption of spoiled food or unripe fruits. Bitter taste prevents ingestion of noxious compounds.
Taste sensation is initiated on contact of chemicals dissolved in saliva with cognate taste receptor molecules on the apical side of specialized epithelial cells (1). These taste receptor cells seem to be dedicated to only one of the basic taste modalities (2). They are assembled into groups of ~100 cells referred to as taste buds, which are structures embedded in the epithelium. On the tongue, taste buds are a part of morphologically, clearly visible epithelial protrusions and/or invaginations known as papillae. The contact of taste stimuli activates signal transduction cascades that result in the depolarization of the receptor cells and the release of the neurotransmitter ATP. ATP excites afferent nerves and allows taste information to be transmitted to the cerebral cortex, where neuronal activity creates the sensory perception (3, 4). In this scenario the taste receptor molecules have the important task of chemical recognition and discrimination. Organisms use these receptor molecules to convert chemical structures into biochemical reactions and, ultimately, to perceive taste.
In recent years, impressive progress has been made in the field of gustation, because of the discovery of the receptors for sweet, umami, bitter, and sour taste and the experimental tools that were created. Our objective here is to review the recent developments in the field with emphasis on taste receptors and their associated biochemical signal transduction cascades.
Sweet Taste
Sweet taste is elicited by many compounds of various chemical classes (Fig. 1). Sweeteners include monosaccharides and disaccharides such as glucose and sucrose; amino acids such as D-tryptophane, alanine, and glycine; proteins such as monellin and thaumatin; and many chemically diverse artificial sweeteners such as saccharin, cyclamate, aspartame, and alitame (5). This observation has fostered long-lasting speculations about how many receptors are necessary to detect the many structurally divergent compounds. Finally, an answer to this question was provided by the discovery of the TAS1R genes that encode a new family of the putative taste receptors (6). The gene family consists of the three members: TAS1R1, TAS1R2, and TAS1R3. TAS1R is the gene symbol proposed by the human genome project nomenclature committee for the gene family previously referred to as T1Rs; the corresponding mouse gene symbol is Tas1r. Genes are written in italics, whereas the protein is printed in normal letters. Many studies have demonstrated convincingly that the TAS1R2/TAS1R3 heteromer mediates the majority of human sweet taste perception.
The TAS1Rs are subclass 3 G-protein-coupled receptors that are related distantly to the Ca2+ sensing receptor, the metabotropic glutamate receptors, GABAB, and the V2R pheromone receptors. Consistent with their proposed role as taste receptors, behavioral experiments and neuronal recordings of Tas1r2 and Tas1r3 knockout mice showed that the deletion of either gene reduced strongly the nerve responses and the attractiveness of various sweeteners, whereas the deletion of the third family member Tas1r1 affected umami taste (7). Moreover, in situ hybridizations in rodents showed that Tas1r3 is co-expressed with either Tas1r2 or Tas1r1 in two nonoverlapping subsets of taste receptor cells (8). This finding led to the hypothesis that the functional sweet receptor could be a heteromer of two Tas1r subunits. Indeed, expression studies in HEK293 cells demonstrated subsequently that cells cotransfected with Tas1r2 and Tas1r3 or its human counterparts responded to various sweeteners (8, 9) and thereby confirmed that Tas1r2 and Tas1r3 form a functional sweet taste receptor. Both the human and the rodent receptors are activated by chemically diverse sweeteners such as monosaccharides, disaccharides, sweet amino acids, and artificial sweeteners (9). Notably, all tested compounds that taste sweet to humans activate the human TAS1R2/TAS1R3 receptor (9).
Interestingly, the human TAS1R2/TAS1R3, but not its mouse counterpart, are sensitive to the sweet proteins monellin, thaumatin, and brazzein, and to the artificial sweeteners neotame, cyclamate, and aspartame (9-11). This difference provides a molecular explanation for the previous observation that these compounds are sweet for humans but not attractive to rodents (9). The species difference also applies to the inhibitor lactisole that blocks the sweet taste in humans but not in rats, and only inhibits the response of human TAS1R2/TAS1R3 to sweet stimuli (9).
Recently, these functional differences between the human and the rodent sweet receptor have been exploited to obtain insight into how this receptor can be activated by so many structurally different sweeteners. Replacement of the large, extracellular domain at the N-terminus of rat Tas1r2 by its human counterpart was sufficient to create a chimeric receptor that could be activated by the dipeptide derivates aspartame, neotame, and the sweet-tasting protein monellin (10, 11). Similarly, replacement of the cysteine-rich region in mouse Tas1r3, which connects the N-terminal extracellular domain to the heptahelical domain by its human counterpart, created a receptor chimera that could be activated by the sweet protein brazzein (11). These findings suggest that the binding sites for aspartame, neotame, and monellin are located in the large extracellular domain of TAS1R2, whereas the binding site for brazzein may be located in the cysteine-rich domain of the TAS1R3 subunit. Additional analyses of receptor chimeras in combination with mutational studies and molecular modeling revealed that the sweet inhibitor lacti-sole and the sweetener cyclamate share an overlapping binding site in the heptahelical domain of the human TAS1R3 subunit (12). Moreover, tryptophan fluorescence spectroscopy analysis of the purified extracellular N-terminal domains of Tas1r3 and Tas1r2 support the notion that sucrose, glucose, and sucralose interact with both domains (13). In summary, these results provide evidence that structurally diverse sweeteners use multiple binding sites to activate the sweet receptor (Fig. 1).

Figure 1. Schematic presentation of proposed binding sites for structurally different sweet tasting compounds at the human sweet taste receptor. Identification of the binding sites for aspartame, neotame, monnellin, brazzein, cyclamate, and lactisole are based on functional analysis of the rodent and human sweet receptor, chimeras created of them and specific receptor mutants, as well as data obtained through molecular modeling. Informations about the sucrose, glucose, and sucralose binding sites have been derived from measuring agonist-induced conformational changes of the purified N-terminal ectodomains of the sweet receptor (see text for further details).
Umami Taste
In humans, umami taste (also referred to as amino acid taste) is elicited predominantly by L-glutamate and L-aspartate (14), whereas rodents respond to most L-amino acids (7). Interestingly, umami taste is enhanced by 5'-ribonucleotides such as inosine-5'-monophosphat (IMP) and guanosin-5'- monophosphate (GMP) (15). Thus, a genuine umami receptor should reflect these properties. Umami compounds are enriched during the ripening processes in many foods, including fruits, vegetables, cheese, and meat. Therefore, this taste quality helps us to choose the ripest fruits and the most palatable cheese for our meal.
In humans, some metabotropic glutamate receptor agonists such as ibotenate and L-AP4 elicit umami taste (15). Moreover, studies have demonstrated the expression of mGluR1-4 in taste buds (16-21). These observations are consistent with the hypothesis that metabotropic glutamate receptors contribute to umami taste. In line with this assumption, the cDNA of an N-terminally truncated “taste” variant (mGluR4t) of the mGluR4 was isolated from rodent tongue tissue (20). Functional studies showed that it could be activated by L-AP4 and glutamate at concentrations that are typical for umami taste (20). Based on these data the truncated mGluR4 variant initially seemed to be an attractive candidate for an umami taste receptor, although several inconsistencies exist [c.f. (7)]. Most prominently, mGluR4 knockout mice show an increased preference for glutamate (22) instead of a reduced response as one would expect. Therefore, receptor activities cannot be enhanced by ribonucleotides.
Studies of Tas1r1 and Tas1r3 knockout mice provide clear evidence for their involvement in umami taste. The deletion of either receptor gene reduced strongly the attractiveness of umami compounds in mice and in the corresponding nerve responses (7). In situ hybridizations in rodents demonstrated that Tas1r3 is coexpressed with Taslrl in a subset of taste receptor cells, which suggests that the umami receptor is a heteromer of TAS1R1 and TAS1R3. In vitro expression studies showed that cells cotransfected with cDNAs for human TAS1R1 and TAS1R3 responded to glutamate, aspartame, and L-AP4 (9), whereas cells transfected with the counterparts from mice acquired general sensitivity for L-amino acids (23). Remarkably, 5'-ribonucleotides such as IMP and GMP enhanced strongly the receptor responses, which is a hallmark of umami taste (9, 15). Thus, these functional properties of the TAS1R1/TAS1R3 receptor dimer of humans and rodents explain some of the most important properties of umami taste. It should be pointed out, however, that the response profiles of all the umami receptor candidates in transfected cells, i.e., TAS1R1/TAS1R3 and the various mGluRs found in taste tissue, differ from those observed in native taste cells (24). A complete description of umami taste transduction may involve combinations of the candidate receptors and/or as yet-undiscovered taste receptors (24), or umami taste may be a delicious flavor formed by neuronal mechanisms in the brain (25).
Bitter Taste
Bitter compounds are numerous and structurally diverse (26). Estimates count thousands of these compounds in the human environment. Known bitter-tasting substances include fatty acids, peptides, amino acids, amines, azacycloalkanes, N-heterocyclic compounds, amides, ureas, thioureas, esters, lactones, carbonyl compounds, phenols, crown ethers, alkaloids, and metal ions. In mammals, these compounds are recognized by approximately 30 G-protein-coupled receptors belonging to the TAS2R gene family (27). These comparably few receptors face the enormous challenge to sense the numerous synthetic and natural bitter substances. One of the central questions in bitter taste research is how these few receptors enable the detection of so many different bitter-tasting compounds. Detailed knowledge about the molecular basis of TAS2R-tastant interaction is required to answer this question. As a first step toward a better understanding, the identification of as many bitter receptor-bitter agonist pairs as possible is necessary to establish a fundament for detailed structure-function analyses. The successful development of a variety of functional expression assays led to an enormous boost of deorphanizations of bitter taste receptors during the last few years. One of the difficulties in setting up efficient screening assays is the insufficient cell-surface targeting properties of bitter taste receptor proteins (28), which have been observed for other chemosensory receptor gene families, such as odorant receptors (29) and pheromone receptors of the V2R type (30). These problems are circumvented commonly by the amino terminal extensions of taste receptors with amino termini of other GPCRs such as bovine rhodopsin (28) or rat somatostatin receptor 3 (31). The physiologic cell-surface targeting properties of TAS2Rs seem to be individual and may depend on various cofactors (32). With such assays in place, nine human (see also Reference 33), two mouse, and one rat TAS2Rs have been de- orphanized (see Table 1) by various laboratories and different experimental approaches to date.
The first mammalian receptor-bitter agonist combinations identified was human TAS2R4 and mouse T2R5 (28). Both receptors responded selectively to one compound of a large panel of known bitter substances best. Whereas mT2R5 was only activated by cycloheximide, hTAS2R4 responded to denato- nium benzoate and high concentrations of 6-n-propyl-thiouracil. This study indicated that mammalian TAS2Rs exhibit a strong selectivity for agonists, although, in the case of hTAS2R4, limited promiscuity might occur at high concentrations. The first hTAS2R identified to be activated by natural bitter substances was hTAS2R16 (31). Systematic testing of substances demonstrated that this receptor responded selectively to an entire group of chemically related compounds, the P-D-glucopyranosides. Thus, hTAS2R16 combines selectivity, even stereoselectivity for the β-D-conformation of the pyranose moiety, with flexibility for other substructures of its agonists. On the other hand, two additional receptors that were deorphanized in the same study, hTAS2R10 and rT2R9, the closest rat homolog of mT2R5, responded only to strychnine and cycloheximide, respectively. The recent discovery of hTAS2R38 as the receptor for PROP and PTC, two synthetic compounds that were known for decades to separate the human population into tasters and nontasters for these chemicals, showed for the first time that genetic polymorphisms in hTAS2R genes account for individual bitter taste perception among humans (34). With respect to agonist specificity, hTAS2R38 exhibits some similarities with hTAS2R16 in combining specificity and flexibility. The taster variant of this receptor recognizes a variety of compounds that have the N-C=S group in common (35). Currently, hTAS2R14 exhibits the highest flexibility for structurally diverse agonists as about one quarter of 33 tested compounds activated this receptor (36). A recent study identified aristolochic acid as an additional agonist for hTAS2R14 and deorphanized hTAS2R7, which also seems to be tuned broadly (37). One might speculate that during evolution different functional constraints shaped bitter taste receptors to face different challenges. More selective receptors might provide sensitivity for the most prominent toxic plant metabolites in a familiar environment, whereas broadly tuned receptors may be more important during exploratory phases in evolution. The characterization of members of a subfamily of closely related hTAS2Rs has been addressed independently by two studies. In one publication, the activation of the closely related receptors, hTAS2R43 and hTAS2R44, by the same subset of agonists although with different pharmacological properties was demonstrated (38), whereas the results of a second study show that hTAS2R43, hTAS2R44, and hTAS2R47 respond selectively to some chemicals (39). Interestingly, hTAS2R43 and hTAS2R44 not only responded to the purely bitter aristolochic acid, but also to the two artificial sweeteners saccharin and acesulfame K accounting for the bitter off-taste observed at high concentrations for these sweeteners (38). Pronin et al. used their discovery of selective agonists for hTAS2R43 (6-nitrosaccharin, IMNB) and hTAS2R47 (6-nitrosaccharin, denatonium) to perform a first structure-function analysis of hTAS2R43, hTAS2R44, and hTAS2R47, which indicates that extracellular as well as transmembrane regions contain residues involved in agonist activation (39).
The deophanization studies also revealed a strong correlation between the sensitivities of the hTAS2Rs for their cognate bitter compounds determined in vitro and the sensitivities of human subjects tasting that compounds. These observations suggest that the receptors report to the brain in the actual concentrations of chemicals and that this information is not modified robustly by neuronal computation. This conclusion is supported strongly by experiments in transgenic mice. Mice are indifferent to β-glucopyranosides, such as salicin, but they taste this compound with similar sensitivity as humans do when they express the human cognate bitter taste receptor hTAS2R16 as transgene.
The more detailed structure-function analyses of several TAS2Rs together with their agonists will provide detailed insight into bitter receptor-agonist interactions required to understand how such few TAS2Rs can recognize so many bitter compounds and might pave the way for the development of bitter antagonists. The recent availability of computer modeling studies of bitter taste receptors with identified agonists docked into these structures increases our knowledge about structure-function relations (40, 41) and may help to guide future mutagenesis analyses.
Table 1. List of deorphanized mammalian bitter taste receptors with their cognate agonists
|
TAS2R |
Agonists |
Agonist structure |
Reference |
|
hTAS2R4 |
denatonium benzoate, 6-n-propyl-2-thiouracil (PROP) |
|
28 |
|
hTAS2R7 |
chloroquine, papavarine, quinacrine, strychnine |
|
37 |
|
hTAS2R10 |
strychnine |
|
31 |
|
hTAS2R14 |
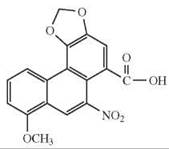
aristolochic acid, 1,8-naphthalaldehydic acid, 1-naphthoic acid, 1-nitronaphthalene, picrotin, picrotoxinin, piperonylic acid, sodium benzoate, (—)-α-thujone |
|
36, 37 |
|
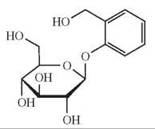
hTAS2R16 |
phenyl-β-D-glucopyranoside, salicin, helicin, arbutin, 2-nitrophenyl-β-D-glucopyranoside, naphtyl-β-D-glucopyranoside, methyl-β-D-glucopyranoside, amygdalin, esculin |
|
31 |
|
hTAS2R38 |
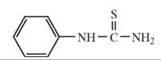
Phenylthiourea (PTC), diphenylthiourea, acetylthiourea, propylthiouracil (PROP), methylthiouracil |
|
34, 35 |
|
hTAS2R43 |
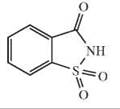
Acesulfame K, aristolochic acid, saccharin, 6-nitrosaccharin, N-isopropyl-2-methyl-5-nitrobenzene sulfonamide (IMNB) |
|
38, 39 |
|
hTAS2R44 |
Acesulfame K, aristolochic acid, saccharin |
|
38 |
|
hTAS2R47 |
Denatonium, 6-nitrosaccharin |
|
39 |
|
mT2R5 |
cycloheximide |
|
28 |
|
rT2R9 |
cycloheximide |
|
31 |
The depicted chemical structures correspond to the agonists printed in bold.
Salt Taste
Two pathways for salt taste have been reported. Nerve recordings performed in rodents showed that the chorda tympani nerve, which innervates the fungiform papillae of the anterior tongue, responded strongly to stimulation with NaCl and that this effect was highly sensitive to amiloride (42, 43). The amiloride-sensitive response was selective for Na+ ions. Based on these observations, the non-voltage-gated, sodium-permeable, heteromeric (α2βy) epithelial sodium channel (ENaC) has been suggested to be a good candidate. In rodents, ENaC subunits are expressed in a specific subset of fungiform taste receptor cells. Whole-cell patch clamp analysis of isolated fungiform taste receptor cells demonstrated that amiloride inhibited Na+-induced currents in micromolar concentrations as expected for ENaC-mediated currents (44, 45). Furthermore, behavioral studies in mice and in rats showed that sodium taste is inhibited partly by adding amiloride to a sodium salt solution, without affecting responses to other taste modalities (46, 47). In humans, the situation is less clear. Psychophysical analysis revealed only a limited reduction of salt taste by amiloride, and this seemed to be restricted to few subjects (48, 49).
In vallate and foliate papillae of rodents only α-ENaC could be easily detected, whereas β- and γ-ENaC are less abundant (50-52), which raises questions about the identity of the salt taste receptor of the posterior tongue. Moreover, NaCl-induced responses of the glossopharyngeal nerve that innervates the vallate and foliate papillae of the posterior tongue seemed to be almost insensitive to amiloride (43, 53). The amiloride-insensitive salt taste receptor is a constitutively active ion channel that is blocked by cetylpyridinium-chloride. It is not selective for sodium ions but mediates NH4+ and K+ currents (54). Based on its sensitivity to the TRPV1 antagonists SB-366791, it has been suggested that the amiloride-insensitive salt taste receptor is a variant of the vanilloid receptor 1, TrpVlt (55). But because not all properties of amiloride-insensitive salt taste receptor are replicated by TRPV1 and TRPV1 gene-targeted mice preferred NaCl over water at concentrations avoided by wild types and salt taste in these animals was less blocked by amiloride (55, 56), the role of TrpVlt in salt taste remains questionable. Taken together, the molecular identity of the salt taste receptor or receptors cannot be taken for granted.
Sour Taste
Sour taste detects acids, i.e., protons. Several different sour taste receptor candidates such as acid-sensing ion channels (ASICs) (57), hyperpolarization-activated cyclic nucleotide-gated channels (HCNs) (58), and two pore domain potassium channels (K2Ps) (59, 60) have been described in the past. In addition, recent research identified two members of the polycystic kidney disease (PKD) family of the transient receptor potential superfamily (TRP) as strong sour taste “receptor” candidates or as part thereof. Immunohistochemistry and in situ hybridization revealed the presence of the polycystic-kidney-disease-like ion channel PKD2L1 in subsets of taste receptor cells of mouse fungiform, vallate, and foliate papillae. These cells differ from those expressing receptors for bitter, sweet, and umami taste (61, 62). Moreover, genetic ablation in mice of the cells expressing PKD1L2 was associated with a loss of response to acidic stimuli in both electrophysiologic recordings and behavioral experiments, whereas other taste modalities remained unaffected, which suggests that PKD2L1 cells are necessary for sour taste perception (62). Previous studies demonstrated that PKD2 polypeptides need to interact with PKD1 polypeptides for proper cell-surface expression (63, 64). Search for interaction partners for PKD2L1 in taste cells identified PKD1L3 in mouse vallate and foliate papillae being coexpressed with PKD2L1 in the same cells (61, 62, 65). PKD1L3 enhanced significantly the expression of PKD2L1 at the cell surface in vitro (61). In fungiform papillae and in the palate, expression of PKD1L3 could not be detected, which suggests that another polypeptide interacts with PKD2L1 in these structures. In accordance with the proposed involvement of PKD2L1/PKD1L3 in sour taste transduction, cells that express the two polypeptides responded to stimulation with acids (61). The data were consistent with the observation that sour taste in humans and rodents is stronger for weak acids, such as citric acid, than for strong acids, such as HCl (66, 67). However, subtle, important differences between the heterologously expressed PKD channel and the sour taste responses in native taste cells (68) require additional work to determine the precise role of the PKDs in sour taste transduction.
Molecular Evolution of Taste Receptor Genes
Taste reception varies enormously across vertebrates. Because the taste perception of a species is related essentially to its diet and environment, the studies of the variation of genes, which control taste reception among vertebrates will contribute greatly to our understanding of the relationship between the adaptation of organisms and the diversity of environmental chemicals.
Evolution dynamics of TAS2R gene family
To date, the complete repertoire of the TAS2R gene family has been reported in human, mouse, rat, dog, cow, opossum, chicken, frog, and several fish (69-72), in addition to a small number of TAS2R genes described in several primates (73-76). Comparisons of these gene repertoires revealed extremely high variation in the sizes of the TAS2R repertoire among species, ranging from 3 genes in chicken to 20-50 genes in mammals and amphibians (71, 72). This finding is consistent with the fact that bitter taste perception, as a warning sensor for bitter toxin intake, varies enormously across vertebrates with different diets and environments. Most interestingly, cows were found to have the highest proportion of TAS2R pseudogenes (44%), which may suggest that detecting poisons in diet is not as important in ruminants as in other animals because of the high detoxification capacity of cow’s rumen microbes (72).
In terms of long-term evolution of the TAS2R gene family, the phylogenetic analysis showed several interesting evolutionary patterns (Fig. 2). First, the TAS2R gene family evolves after the birth-and-death process (71), which is characterized by frequent gene duplication and gene deactivation, similar to that found in the olfactory receptor gene family (77). Second, there might be multiple TAS2R genes in the common ancestor of tetropods and teleosts (72). Third, the TAS2R repertoire expanded considerably in the common ancestor of tetrapods, followed by additional independent expansions in frogs and mammals, and contractions of TAS2R repertoire occurred in chicken (71, 72). Last but not least, based on a comparison of human and mouse TAS2R gene repertoires, some TAS2R genes exhibit one-to-one orthologous pairing, whereas other genes are part of lineage-specific (species-specific) clusters, in which the genes from the same species cluster together in the phylogenetic tree (78). These species-specific genes are also located closer to each other on chromosomes, which indicates that these newly duplicated genes resulted from tandem gene duplications (70, 78). Also, these genes seem to be under positive selection, which suggests that they are used for species-specific bitter tastants (78). On the other hand, one-to-one orthologous genes are subject to more selective constraints than lineage-specific (species-specific) genes, which indicates that each of the one-to-one orthologous genes possibly is detecting one or several distinct bitter tastants that are encountered by a wide range of animals (78). Although two recent evolutionary studies (71, 72), which extended the study of TAS2Rs outside of human and mouse to an additional nine vertebrate species, supports this hypothesis, it still waits to be scrutinized additionally by functional research.
The comparative analysis of the TAS2R gene family between several primate species with rodents revealed that primate genes were under less selective pressure than rodent genes (73-76). First, the comparison of the gene birth/death rate between primates and mice shows that the proportion of pseudogenes in the TAS2R repertoire is lower in mice (15%) than in apes (21%-28%), which is in turn lower than that in humans (31%) (73, 74). Moreover, the prevalence of lineage-specific pseudogenes in primates supports this conclusion (74). Second, based on the equal levels of nonsynonymous/synonymous substitution rate ratios for the TAS2R genes in primates, the functional constraints were more relaxed in the primate lineage than in the mouse lineage (73, 74, 76). This evolutionary pattern could be caused by the reduced effective population sizes in primates, which might cause less-effective purifying selection (73). The alternative explanation is that the reduced functional constraints in primates might be caused by reduced bitter taste needs because of a change of the environment and the diet (74). In fact, some ecologic studies support this explanation. For example, meat accounts for 2-13% of diet in chimpanzees, whereas it has never been found in other apes’ diet (76). Furthermore, this explanation has been strengthened by the findings that there were significant changes in human diet, such as increasing food from animal sources while decreasing food from plant sources, and the controlled use of fire to detoxify the food (76). Both factors may have caused a reduction in the importance of bitter taste and consequently triggered a functional relaxation in humans.
Evidence for the relaxation of selective constraints on TAS2R genes in apes and humans does not preclude the possibility that positive selection occurred on a few specific genes. Positive selection has been found in the gene for the human TAS2R16, the beta-glucopyranoside receptor (31). By analyzing the sequences from 60 human populations, Soranzo et al. detected signatures of positive selection on a more sensitive derived allele, which was found in all human populations except for African populations (79). This result might reflect the increased sensitivity of the derived TAS2R16 allele under the positive selection through an increased protection against harmful cyanogenic plant foods and natural toxins (79). In addition to the selective relaxation and positive selection, the most complex scenario of evolutionary forces has been observed in TAS2R38 gene, which is responsible largely for the human polymorphism in tasting phenylth- iocarbamide (PTC) (34). Interestingly, chimpanzees are also known to have tasters and nontasters of PTC (80). Although humans and chimpanzees shared phenotypic polymorphism, they did not share the same evolutionary forces of maintaining nontasters’ alleles (80, 81). Balancing natural selection has been suggested to maintain functional nontaster TAS2R38 alleles in human populations (81). By contrast, the nontaster allele was lost in chimpanzees, which favors the selective relaxation hypothesis in this lineage (80). As more human TAS2R genes are being studied, the understanding of the evolutionary forces behind each TAS2R will increase considerably.

Figure 2. Phylogenetic tree of vertebrate TAS2R genes. The arrow points to where the tree is rooted with vertebrate V1Rs. Image is adapted from Reference 72.
Evolution dynamics of TAS1R gene family
In contrast to the TAS2R gene family, the TAS1R family is remarkably well conserved during evolution both in gene family size and in sequence divergence (72). In terms of gene family size, the number of TAS1R gene repertoire changes rarely in mammals, which might reflect the necessity of both sweet and umami tastes among mammals (72). But the number of TAS1R gene repertoire varies in some nonmammalian vertebrates, both with a few gene duplications observed in pufferfish and fugu and with gene loss events found in western clawed frog and chicken (72). In addition to the western clawed frog, which does not have any TAS1R genes, a loss of the TAS1R2 gene was identified in the chicken genome (72). In addition, cats and closely related carnivores are also known to lack the TAS1R2 genes (82), which might reflect the insensitivity to sweet taste stimuli in these species (72, 82). Thus, pseudogenization of TAS1R2 occurred multiple times independently in evolution.
In sequence divergence level, TAS1R genes evolve more slowly than TAS2R genes whatever the comparison between species or within species. For interspecies comparison, the sequence divergence distance of orthologous pairs among human, mouse, rat, and opossum is significantly lower for TAS1R genes than TAS2R genes (72). Similarly in human populations, the mean pairwise differences per nucleotide between sequences of TAS2R are greater than those of TAS1R, which reveals the lower levels of nucleotide diversity in TAS1R family (83, 84). The positive selection has been suggested to operate separately on paralogous TAS1R genes and different alleles (72, 84), which is also the case in TAS2R genes, as mentioned above.
Extragustatory Expression of Taste Receptors
The role of taste perception in the oral cavity is to analyze the composition of food for its nutritional value and for the presence of potentially harmful substances prior to ingestion. However, evidence is accumulating that elements of the taste perception machinery, including taste receptor proteins, are expressed at several extragustatory sites as well. The anatomical organization of chemosensory structures is variable, apparently becoming less complex with growing distance from the primary gustatory areas. Taste buds located in gustatory papillae on the surface of the tongue, on the soft palate, and on the oro-pharynx are well-organized groups of about 60 to 100 cells (1). Laryngeal taste buds, however, are smaller than lingual buds. Sbarbati et al. (86) observed that the sizes and shapes in rats changed from the most rostral part of the laryngeal inlet, where mostly buds were found, to structures called “chemosensory clusters” distally. These chemosensory clusters contained only 2-3 cells staining positive for PLCβ2, which is a molecule involved critically in sweet, umami, and bitter taste transduction (85). Rarely, solitary chemosensory cells were found in this part of the larynx but they become more numerous distally (86) and extend into the airway epithelium (87). Solitary chemosensory cells are also present in respiratory nasal epithelium and the vomeronasal epithelium (88, 89). Another type of cells that express components of taste signal transduction are the brush cells, which line the stomach, the duodenum (90), and the pancreatic duct system (91). It should be noted here that, by anatomical criteria, brush cells might be related to, but they are not solitary chemosensory cells (92). Recently, secretory cells of the airway (93) and spermatozoa (94) have been identified to express taste signaling components as well.
Within the gastrointestinal tract of rodents, brush cells of the stomach, duodenum (90), and the pancreatic duct system (91) express the G-protein subunit α-gustducin, which has been demonstrated to be important for bitter, sweet, and umami taste transduction (95). By RT-PCR analyses of gastrointestinal tissues of rat and mice, a considerable number of TAS2R genes have been detected, although the cellular origin of the detected mRNAs was not directly addressed (96). Another study identified TAS1R1, TAS1R2, TAS1R3, PLCβ2, and TRPM5 along with α-gustducin. In case of TAS1R2, expression was only weak and more restricted, which suggests that most components of the canonical sweet and umami taste transduction are present in the GI tract. However, using the transgenic expression of GFP under the control of the TRPM5 5'-flanking region for colocalization, experiments with the other taste transduction molecules revealed a less clear picture. Although the colocalization of TRPM5-driven GFP expression with PLCβ2 is limited, both molecules are indispensable for bitter, sweet, and umami taste transduction as shown by knockout mouse models, which suggests that at least in part a variant transduction mechanism acts in these cells. Moreover, this study identified that, in addition to brush cells, enteroendocrine cells are positive for TRPM5-driven GFP expression (97). With respect to bitter taste receptor-mediated signal transduction, however, the mouse intestinal cell line STC-1, which expresses bitter taste receptor genes along with α-gustducin and α-transducin, reacts selectively to various bitter stimuli with transient calcium signals, which indicates the canonical transduction mechanism (96). As chemosensory cells of the gut are not innervated directly (90), and do not express the presynaptic marker SNAP25 (97), it is speculated that they might communicate via NO (98) or a currently unidentified messenger with neighboring cells/nerve fibers to control appetitive behavior or food passage through the gastrointestinal tract.
Morphologically, the α-gustducin expressing cells found in the nasal and vomeronasal epithelia resemble solitary chemosensory cells (SCCs) described in nonmammalian vertebrates (88). In addition to α-gustducin, some bitter taste receptor genes are expressed in SCCs, whereas the TAS1R1 and TAS1R2 subunits, which specifically constitute the sweet and umami receptor heteromers TAS1R2/TAS1R3 and TAS1R1/TAS1R3, respectively, are absent from nasal chemosensory cells. It indicated already that, analogous to the warning function of bitter taste receptor cells of the oral cavity, the function of TAS2R expressing cells in nasal respiratory epithelium might protect the animal from the aspiration of noxious substances. Indeed, intranasal irrigation with bitter compounds not only elicited trigeminal responses but also resulted in pronounced respiratory effects (88). Another recent example for extragustatory expression and multiple functions of taste receptor molecules aside from pure gustation is PKD2L1, a mammalian sour taste sensor, which is also expressed in a discrete population of neurons surrounding the central canal of the spinal cord, perhaps involved in monitoring the pH of the cerebrospinal fluid (62).
In perspective, the growing number of reports on the extragustatory expression of components of the taste transduction cascade requires the careful analyses of taste-specific knock out models for nongustatory deficits.
Perspectives
During the last few years, the field of taste research has been progressing with enormous speed. Current research addresses structural details on receptor agonist interactions for bitter and sweet taste receptors. For the other taste modalities, including salt taste, where definite proof for the proposed roles of candidate sensor molecules is still missing, more questions still need to be answered. A next big step in taste research will be to understand how taste information is processed along its pathway from the periphery into higher order brain centers. In view of emerging extragustatory functions of taste transduction molecules, it will be critical to understand how the different functions and cellular environments have modified the involved proteins and signaling mechanisms to serve multiple functions. More practically, the cloning of the taste receptors enabled recombinant receptor assays to be established. With these tools at hand, the food and flavor industry will be able to design high throughput screens of complex compound libraries in order to identify new taste-active compounds. Sweet taste enhancers, glutamate substitutes and salt replacers, and specific bitter blockers might be the primary targets.
References
1. Miller IJ.Jr. (ed). Anatomy of the peripheral taste system. In: Handbook of Olfaction and Gustation. Doty RL, ed. 1995. Dekker, New York. pp. 521-547.
2. Chandrashekar J, Hoon MA, Ryba NJ, Zuker CS. The receptors and cells for mammalian taste. Nature 2006; 444:288-294.
3. Finger TE, Danilova V, Barrows J, Bartel DL, Vigers AJ, Stone L, Hellekant G, Kinnamon SC. ATP signaling is crucial for communication from taste buds to gustatory nerves. Science 2005; 310:1495-1499.
4. Romanov RA, Rogachevskaja OA, Bystrova MF, Jiang P, Mar- golskee RF, Kolesnikov SS. Afferent neurotransmission mediated by hemichannels in mammalian taste cells. Embo. J. 2007; 26:657-667.
5. Schiffman SS, Gatlin CA. Sweeteners: state of knowledge review. Neurosci. Biobehav. Rev. 1993; 17:313-345.
6. Hoon MA, Adler E, Lindemeier J, Battey JF, Ryba NJ, Zuker CS. Putative mammalian taste receptors: a class of taste-specific GPCRs with distinct topographic selectivity. Cell 1999; 96: 541-551.
7. Zhao GQ, Zhang Y, Hoon MA, Chandrashekar J, Erlenbach I, Ryba NJ, Zuker CS. The receptors for mammalian sweet and umami taste. Cell 2003; 115:255-266.
8. Nelson G, Hoon MA, Chandrashekar J, Zhang Y, Ryba NJ, Zuker CS. Mammalian sweet taste receptors. Cell 2001; 106:381-390.
9. Li X, Staszewski L, Xu H, Durick K, Zoller M, Adler E. Human receptors for sweet and umami taste. Proc. Natl. Acad. Sci. U. S. A. 2002; 99:4692-4696.
10. Xu H, Staszewski L, Tang H, Adler E, Zoller M, Li X. Different functional roles of T1R subunits in the heteromeric taste receptors. Proc. Natl. Acad. Sci. U. S. A. 2004; 101:14258-14263.
11. Jiang P, Ji Q, Liu Z, Snyder LA, Benard LM, Margolskee RF, Max M. The cysteine-rich region of T1R3 determines responses to intensely sweet proteins. J. Biol. Chem. 2004; 279:45068-45075.
12. Jiang P, Cui M, Zhao B, Snyder LA, Benard LM, Osman R, Max M, Margolskee RF. Identification of the cyclamate interaction site within the transmembrane domain of the human sweet taste receptor subunit T1R3. J. Biol. Chem. 2005; 280:34296-34305.
13. Nie Y, Vigues S, Hobbs JR, Conn GL, Munger SD. Distinct contributions of T1R2 and T1R3 taste receptor subunits to the detection of sweet stimuli. Curr. Biol. 2005; 15:1948-1952.
14. Kawai M, Okiyama A, Ueda Y. Taste enhancements between various amino acids and IMP. Chem. Senses 2002; 27:739-745.
15. Kurihara K, Kashiwayanagi M. Physiological studies on umami taste. J. Nutr. 2000; 130:931S-934S.
16. Toyono T, Kataoka S, Seta Y, Shigemoto R, Toyoshima K. Expression of group II metabotropic glutamate receptors in rat gustatory papillae. Cell Tissue Res. 2007; 328:57-63.
17. Toyono T, Seta Y, Kataoka S, Kawano S, Shigemoto R, Toyoshima K. Expression of metabotropic glutamate receptor group I in rat gustatory papillae. Cell Tissue Res. 2003;313:29-35.
18. Yang H, Wanner IB, Roper SD, Chaudhari N. An optimized method for in situ hybridization with signal amplification that allows the detection of rare mRNAs. J. Histochem. Cytochem. 1999; 47:431-446.
19. Chaudhari N, Yang H, Lamp C, Delay E, Cartford C, Than T, Roper S. The taste of monosodium glutamate: membrane receptors in taste buds. J. Neurosci. 1996; 16:3817-3826.
20. Chaudhari N, Landin AM, Roper SD. A metabotropic glutamate receptor variant functions as a taste receptor. Nat. Neurosci. 2000; 3:113-119.
21. San Gabriel A, Uneyama H, Yoshie S, Torii K. Cloning and characterization of a novel mGluR1 variant from vallate papillae that functions as a receptor for L-glutamate stimuli. Chem. Senses 2005; 30:i2-i26.
22. Chaudhari N, Roper SD. Molecular and physiological evidence for glutamate (umami) taste transduction via a G protein-coupled receptor. Ann. N. Y. Acad. Sci. 1998; 855:398-406.
23. Nelson G, Chandrashekar J, Hoon MA, Feng L, Zhao G, Ryba NJ, Zuker CS. An amino-acid taste receptor. Nature 2002; 416:199-202.
24. Maruyama Y, Pereira E, Margolskee RF, Chaudhari N, Roper SD. Umami responses in mouse taste cells indicate more than one receptor. J. Neurosci. 2006; 26:2227-2234.
25. McCabe C, Rolls ET. Umami: a delicious flavor formed by convergence of taste and olfactory pathways in the human brain. Eur. J. Neurosci. 2007; 25:1855-1864.
26. Belitz H-D, Wieser H. Bitter compounds: occurrence and structure-activity relationship. Food Reviews International 1985; 1:271-354.
27. Meyerhof W. Elucidation of mammalian bitter taste. Rev Physiol Biochem Pharmacol 2005; 154:37-72.
28. Chandrashekar J, Mueller KL, Hoon MA, Adler E, Feng L, Guo W, Zuker CS, Ryba NJ. T2Rs function as bitter taste receptors. Cell 2000; 100:703-711.
29. McClintock TS, Landers TM, Gimelbrant AA, Fuller LZ, Jackson BA, Jayawickreme CK, Lerner MR. Functional expression of olfactory-adrenergic receptor chimeras and intracellular retention of heterologously expressed olfactory receptors. Brain Res Mol Brain Res 1997; 48:270-278.
30. Loconto J, Papes F, Chang E, Stowers L, Jones EP, Takada T, Kumanovics A, Fischer Lindahl K, Dulac C. Functional expression of murine V2R pheromone receptors involves selective association with the M10 and M1 families of MHC class Ib molecules. Cell 2003; 112:607-618.
31. Bufe B, Hofmann T, Krautwurst D, Raguse JD, Meyerhof W. The human TAS2R16 receptor mediates bitter taste in response to beta-glucopyranosides. Nat. Genet. 2002; 32:397-401.
32. Behrens M, Bartelt J, Reichling C, Winnig M, Kuhn C, Meyerhof W. Members of RTP and REEP gene families influence functional bitter taste receptor expression. J. Biol. Chem. 2006; 281:20650-20659.
33. Behrens M, Meyerhof W. Bitter taste receptors and human bitter taste perception. Cell. Mol. Life Sci. 2006; 63:1501-1509.
34. Kim UK, Jorgenson E, Coon H, Leppert M, Risch N, Drayna D. Positional cloning of the human quantitative trait locus underlying taste sensitivity to phenylthiocarbamide. Science 2003; 299: 1221-1225.
35. Bufe B, Breslin PA, Kuhn C, Reed DR, Tharp CD, Slack JP, Kim UK, Drayna D, Meyerhof W. The molecular basis of individual differences in phenylthiocarbamide and propylthiouracil bitterness perception. Curr. Biol. 2005; 15:322-327.
36. Behrens M, Brockhoff A, Kuhn C, Bufe B, Winnig M, Meyerhof W. The human taste receptor hTAS2R14 responds to a variety of different bitter compounds. Biochem. Biophys. Res. Commun. 2004; 319:479-485.
37. Sainz E, Cavenagh MM, Gutierrez J, Battey JF, Northup JK, Sullivan SL. Functional characterization of human bitter taste receptors. Biochem. J. 2007; 403:537-543.
38. Kuhn C, Bufe B, Winnig M, Hofmann T, Frank O, Behrens M, Lewtschenko T, Slack JP, Ward CD, Meyerhof W. Bitter taste receptors for saccharin and acesulfame K. J. Neurosci. 2004; 24:10260-10265.
39. Pronin AN, Tang H, Connor J, Keung W. Identification of ligands for two human bitter T2R receptors. Chem. Senses 2004; 29:583-593.
40. Floriano WB, Hall S, Vaidehi N, Kim U, Drayna D, Goddard WA 3rd. Modeling the human PTC bitter-taste receptor interactions with bitter tastants. J. Mol. Model. 2006; 12:931-941.
41. Miguet L, Zhang Z, Grigorov MG. Computational studies of ligand-receptor interactions in bitter taste receptors. J. Recept. Signal. Transduct. Res. 2006; 26:611-630.
42. Brand JG, Teeter JH, Silver WL. Inhibition by amiloride of chorda tympani responses evoked by monovalent salts. Brain Res. 1985; 334:207-214.
43. Kitada Y, Mitoh Y, Hill DL. Salt taste responses of the IXth nerve in Sprague-Dawley rats: lack of sensitivity to amiloride. Physiol. Behav. 1998; 63:945-949.
44. Avenet P, Lindemann B. Amiloride-blockable sodium currents in isolated taste receptor cells. J. Membr. Biol. 1988; 105:245-255.
45. Doolin RE, Gilbertson TA. Distribution and characterization of functional amiloride-sensitive sodium channels in rat tongue. J. Gen. Physiol. 1996; 107:545-554.
46. Eylam S, Spector AC. Oral amiloride treatment decreases taste sensitivity to sodium salts in C57BL/6J and DBA/2J mice. Chem. Senses 2003; 28:447-458.
47. Spector AC, Guagliardo NA, St John SJ. Amiloride disrupts NaCl versus KCl discrimination performance: implications for salt taste coding in rats. J. Neurosci. 1996; 16:8115-8122.
48. Feldman GM, Mogyorosi A, Heck GL, DeSimone JA, Santos CR, Clary RA, Lyall V. Salt-evoked lingual surface potential in humans. J. Neurophysiol. 2003; 90:2060-2064.
49. Halpern BP, Darlington RB. Effects of amiloride on gustatory quality descriptions and temporal patterns produced by NaCl. Chem. Senses 1998; 23:501-511.
50. Kretz O, Barbry P, Bock R, Lindemann B. Differential expression of RNA and protein of the three pore-forming subunits of the amiloride-sensitive epithelial sodium channel in taste buds of the rat. J. Histochem. Cytochem. 1999; 47:51-64.
51. Lin W, Finger TE, Rossier BC, Kinnamon SC. Epithelial Na+ channel subunits in rat taste cells: localization and regulation by aldosterone. J. Comp. Neurol. 1999; 405:406-420.
52. Shigemura N, Islam AA, Sadamitsu C, Yoshida R, Yasumatsu K, Ninomiya Y. Expression of amiloride-sensitive epithelial sodium channels in mouse taste cells after chorda tympani nerve crush. Chem. Senses 2005; 30:531-538.
53. Formaker BK, Hill DL. Lack of amiloride sensitivity in SHR and WKY glossopharyngeal taste responses to NaCl. Physiol. Behav. 1991; 50:765-769.
54. DeSimone JA, Lyall V, Heck GL, Phan TH, Alam RI, Feldman GM, Buch RM. A novel pharmacological probe links the amiloride-insensitive NaCl, KCl, and NH(4)Cl chorda tympani taste responses. J. Neurophysiol. 2001; 86:2638-2641.
55. Lyall V, Heck GL, Vinnikova AK, Ghosh S, Phan TH, Alam RI, Russell OF, Malik SA, Bigbee JW, DeSimone JA. The mammalian amiloride-insensitive non-specific salt taste receptor is a vanilloid receptor-1 variant. J. Physiol. 2004; 558:147-159.
56. Ruiz C, Gutknecht S, Delay E, Kinnamon S. Detection of NaCl and KCl in TRPV1 Knockout Mice. Chem. Senses 2006; 31:813-820.
57. Ugawa S, Minami Y, Guo W, Saishin Y, Takatsuji K, Yamamoto T, Tohyama M, Shimada S. Receptor that leaves a sour taste in the mouth. Nature 1998; 395:555-556.
58. Stevens DR, Seifert R, Bufe B, Muller F, Kremmer E, Gauss R, Meyerhof W, Kaupp UB, Lindemann B. Hyperpolarization- activated channels HCN1 and HCN4 mediate responses to sour stimuli. Nature 2001; 413:631-635.
59. Richter TA, Dvoryanchikov GA, Chaudhari N, Roper SD. Acid- sensitive two-pore domain potassium (K2P) channels in mouse taste buds. J. Neurophysiol. 2004; 92:1928-1936.
60. Lin W, Burks CA, Hansen DR, Kinnamon SC, Gilbertson TA. Taste receptor cells express pH-sensitive leak K+ channels. J. Neurophysiol. 2004; 92:2909-2919.
61. Ishimaru Y, Inada H, Kubota M, Zhuang H, Tominaga M, Mat- sunami H. Transient receptor potential family members PKD1L3 and PKD2L1 form a candidate sour taste receptor. Proc. Natl. Acad. Sci. U. S. A. 2006; 103:12569-12574.
62. Huang AL, Chen X, Hoon MA, Chandrashekar J, Guo W, Trankner D, Ryba NJ, Zuker CS. The cells and logic for mammalian sour taste detection. Nature 2006; 442:934-938.
63. Hanaoka K, Qian F, Boletta A, Bhunia AK, Piontek K, Tsiokas L, Sukhatme VP, Guggino WB, Germino GG. Co-assembly of polycystin-1 and -2 produces unique cation-permeable currents. Nature 2000; 408:990-994.
64. Murakami M, Ohba T, Xu F, Shida S, Satoh E, Ono K, Miyoshi I, Watanabe H, Ito H, Iijima T. Genomic organization and functional analysis of murine PKD2L1. J. Biol. Chem. 2005; 280:5626-5635.
65. LopezJimenez ND, Cavenagh MM, Sainz E, Cruz-Ithier MA, Battey JF, Sullivan SL. Two members of the TRPP family of ion channels, Pkd1l3 and Pkd2l1, are co-expressed in a subset of taste receptor cells. J. Neurochem. 2006; 98:68-77.
66. Ogiso K, Shimizu Y, Watanabe K, Tonosaki K. Possible involvement of undissociated acid molecules in the acid response of the chorda tympani nerve of the rat. J. Neurophysiol. 2000; 83:2776- 2779.
67. Ganzevles PG, Kroeze JH. Effects of adaptation and crossadaptation to common ions on sourness intensity. Physiol. Behav. 1987; 40:641-646.
68. Lindemann B. Taste reception. Physiol. Rev. 1996; 76:718-766.
69. Conte C, Ebeling M, Marcuz A, Nef P, Andres-Barquin PJ. Identification and characterization of human taste receptor genes belonging to the TAS2Rfamily. Cytogenet. Genome Res. 2002; 98:45-53.
70. Conte C, Ebeling M, Marcuz A, Nef P, Andres-Barquin PJ. Evolutionary relationships of the Tas2r receptor gene families in mouse and human. Physiol. Genomics 2003; 14:73-82.
71. Go Y. Proceedings of the SMBE Tri-National Young Investigators’ Workshop Lineage-specific expansions and contractions of the bitter taste receptor gene repertoire in vertebrates. Mol. Biol. Evol. 2006; 23:964-972.
72. Shi P, Zhang J. Contrasting modes of evolution between vertebrate sweet/umami receptor genes and bitter receptor genes. Mol. Biol. Evol. 2006; 23:292-300.
73. Fischer A, Gilad Y, Man O, Paabo S. Evolution of bitter taste receptors in humans and apes. Mol. Biol. Evol. 2005; 22:432-436.
74. Go Y, Satta Y, Takenaka O, Takahata N. Lineage-specific loss of function of bitter taste receptor genes in humans and nonhuman primates. Genetics 2005; 170:313-326.
75. Parry CM, Erkner A, le Coutre J. Divergence of T2R chemosensory receptor families in humans, bonobos, and chimpanzees. Proc. Natl. Acad. Sci. U. S. A. 2004; 101:14830-14834.
76. Wang X, Thomas SD, Zhang J. Relaxation of selective constraint and loss of function in the evolution of human bitter taste receptor genes. Hum. Mol. Genet. 2004; 13:2671-2678.
77. Zhang X, Firestein S. The olfactory receptor gene superfamily of the mouse. Nat. Neurosci. 2002; 5:124-133.
78. Shi P, Zhang J, Yang H, Zhang YP. Adaptive diversification of bitter taste receptor genes in Mammalian evolution. Mol. Biol. Evol. 2003; 20:805-814.
79. Soranzo N, Bufe B, Sabeti PC, Wilson JF, Weale ME, Marguerie R, Meyerhof W, Goldstein DB. Positive selection on a high-sensitivity allele of the human bitter-taste receptor TAS2R16. Curr. Biol. 2005; 15:1257-1265.
80. Wooding S, Bufe B, Grassi C, Howard MT, Stone AC, Vazquez M, Dunn DM, Meyerhof W, Weiss RB, Bamshad MJ. Independent evolution of bitter-taste sensitivity in humans and chimpanzees. Nature 2006; 440:930-934.
81. Wooding S, Kim UK, Bamshad MJ, Larsen J, Jorde LB, Drayna D. Natural selection and molecular evolution in PTC, a bitter-taste receptor gene. Am. J. Hum. Genet. 2004; 74:637-646.
82. Li X, Li W, Wang H, Cao J, Maehashi K, Huang L, Bachmanov AA, Reed DR, Legrand-Defretin V, Beauchamp GK, Brand JG. Pseudogenization of a sweet-receptor gene accounts for cats’ indifference toward sugar. PLoS Genet. 2005; 1:27-35.
83. Kim U, Wooding S, Ricci D, Jorde LB, Drayna D. Worldwide haplotype diversity and coding sequence variation at human bitter taste receptor loci. Hum. Mutat. 2005; 26:199-204.
84. Kim UK, Wooding S, Riaz N, Jorde LB, Drayna D. Variation in the human TAS1R taste receptor genes. Chem. Senses 2006; 31:599-611.
85. Zhang Y, Hoon MA, Chandrashekar J, Mueller KL, Cook B, Wu D, Zuker CS, Ryba NJ. Coding of sweet, bitter, and umami tastes: different receptor cells sharing similar signaling pathways. Cell 2003; 112:293-301.
86. Sbarbati A, Merigo F, Benati D, Tizzano M, Bernardi P, Crescimanno C, Osculati F. Identification and characterization of a specific sensory epithelium in the rat larynx. J. Comp. Neurol. 2004; 475:188-201.
87. Merigo F, Benati D, Tizzano M, Osculati F, Sbarbati A. alpha-Gustducin immunoreactivity in the airways. Cell Tissue Res. 2005; 319:211-219.
88. Finger TE, Bottger B, Hansen A, Anderson KT, Alimohammadi H, Silver WL. Solitary chemoreceptor cells in the nasal cavity serve as sentinels of respiration. Proc. Natl. Acad. Sci. U. S. A. 2003; 100:8981-8986.
89. Zancanaro C, Caretta CM, Merigo F, Cavaggioni A, Osculati F. alpha-Gustducin expression in the vomeronasal organ of the mouse. Eur. J. Neurosci. 1999; 11:4473-4475.
90. Hofer D, Puschel B, Drenckhahn, D. Taste receptor-like cells in the rat gut identified by expression of alpha-gustducin. Proc. Natl. Acad. Sci. U. S. A. 1996; 93:6631-6634.
91. Hofer D, Drenckhahn D. Identification of the taste cell G-protein, alpha-gustducin, in brush cells of the rat pancreatic duct system. Histochem. Cell. Biol. 1998; 110:303-309.
92. Sbarbati A, Osculati F. A new fate for old cells: brush cells and related elements. J. Anat. 2005; 206:349-358.
93. Merigo F, Benati D, Di Chio M, Osculati F, Sbarbati A. Secretory cells of the airway express molecules of the chemoreceptive cascade. Cell Tissue Res. 2007; 327:231-247.
94. Fehr J, Meyer D, Widmayer P, Borth HC, Ackermann F, Wilhelm B, Gudermann T, Boekhoff I. Expression of the G-protein alpha-subunit gustducin in mammalian spermatozoa. J. Comp. Physiol. A Neuroethol. Sens Neural Behav. Physiol. 2007; 193:21-34.
95. McLaughlin SK, McKinnon PJ, Margolskee RF. Gustducin is a taste-cell-specific G protein closely related to the transducins. Nature 1992; 357:563-569.
96. Wu SV, Rozengurt N, Yang M, Young SH, Sinnett- Smith J, Rozengurt E. Expression of bitter taste receptors of the T2R family in the gastrointestinal tract and enteroendocrine STC-1 cells. Proc. Natl. Acad. Sci. U. S. A. 2002; 99:2392-2397.
97. Bezencon C, le Coutre J, Damak S. Taste-signaling proteins are coexpressed in solitary intestinal epithelial cells. Chem Senses 2006; 32:41-49.
98. Kugler P, Drenckhahn D. Intrinsic source of stomach NO. Nature 1994; 370:25-26.