Cracking the AP Biology Exam 2018


Editorial
Rob Franek, Editor-in-Chief
Casey Cornelius, VP Content Development
Mary Beth Garrick, Director of Production
Selena Coppock, Managing Editor
Meave Shelton, Senior Editor
Colleen Day, Editor
Sarah Litt, Editor
Aaron Riccio, Editor
Orion McBean, Associate Editor
Penguin Random House Publishing Team
Tom Russell, VP, Publisher
Alison Stoltzfus, Publishing Director
Jake Eldred, Associate Managing Editor
Ellen Reed, Production Manager
Suzanne Lee, Designer
The Princeton Review
555 W. 18th Street
New York, NY 10011
Email: editorialsupport@review.com
Copyright © 2017 by TPR Education IP Holdings, LLC. All rights reserved.
Published in the United States by Penguin Random House LLC, New York, and in Canada by Random House of Canada, a division of Penguin Random House Ltd., Toronto.
Terms of Service: The Princeton Review Online Companion Tools (“Student Tools”) for retail books are available for only the two most recent editions of that book. Student Tools may be activated only twice per eligible book purchased for two consecutive 12-month periods, for a total of 24 months of access. Activation of Student Tools more than twice per book is in direct violation of these Terms of Service and may result in discontinuation of access to Student Tools Services.
Trade Paperback ISBN 9781524710606
Ebook ISBN 9781524710637
AP and Advanced Placement Program are registered trademarks of the College Board, which does not sponsor or endorse this product.
The Princeton Review is not affiliated with Princeton University.
Editor: Sarah Litt
Production Editors: Dallin Law and Jim Melloan
Production Artist: Craig Patches
Cover art by Sebastian Kaulitzki / Alamy Stock Photo
Cover design by Suzanne Lee
v4.1
a
Acknowledgments
The Princeton Review would like to thank Katie Chamberlain, Ph.D, Craig Patches, Jim Melloan, and Dallin Law for their hard work on revisions to this edition.
Contents
Cover
Title Page
Copyright
Acknowledgments
Register Your Book Online!
Part I: Using This Book to Improve Your AP Score
Part II: Practice Test 1
Part III: About the AP Biology Exam
Part IV: Test-Taking Strategies for the AP Biology Exam
1 How to Approach Multiple-Choice Questions
2 How to Approach Free-Response Questions
3 Using Time Effectively to Maximize Points
Part V: Content Review for the AP Biology Exam
4 Chemistry of Life
5 Cells
6 Cellular Energetics
7 Molecular Biology
8 Cell Reproduction
9 Heredity
10 Evolutionary Biology
11 Animal Structure and Function
12 Behavior and Ecology
13 Quantitative Skills and Biostatistics
14 Sample Free-Response Questions
15 Laboratory
16 Chapter Drill Answers and Explanations
AP Biology Equations and Formulas
Part VI: Additional Practice Tests
Register Your Book Online!
1 Go to PrincetonReview.com/cracking
2 You’ll see a welcome page where you can register your book using the following ISBN: 9781524710637.
3 After placing this free order, you’ll either be asked to log in or to answer a few simple questions in order to set up a new Princeton Review account.
4 Finally, click on the “Student Tools” tab located at the top of the screen. It may take an hour or two for your registration to go through, but after that, you’re good to go.
If you have noticed potential content errors, please email EditorialSupport@review.com with the full title of the book, its ISBN (located above), and the page number of the error.
Experiencing technical issues? Please email TPRStudentTech@review.com with the following information:
• your full name
• email address used to register the book
• full book title and ISBN
• your computer OS (Mac or PC) and Internet browser (Firefox, Safari, Chrome, etc.)
• description of technical issue
Once you’ve registered, you can…
• Access the fifth full-length AP Biology Test (or click here to access as a PDF)
• Find any late-breaking information released about the AP Biology Exam
• Take a full-length practice SAT and ACT
• Get valuable advice about the college application process, including tips for writing a great essay and where to apply for financial aid
• Sort colleges by whatever you’re looking for (such as Best Theater or Dorm), learn more about your top choices, and see how they all rank according to The Best 382 Colleges
• Access comprehensive study guides and a variety of printable resources, including bubble sheets and the AP Biology equation and formula tables
• Check to see if there have been any corrections or updates to this edition
Look For These Icons Throughout The Book
 Premium Portal
Premium Portal
 Online Practice Tests
Online Practice Tests
 College Advisor
College Advisor
 Proven Techniques
Proven Techniques
 Applied Strategies
Applied Strategies
 More Great Books
More Great Books
Part I
Using This Book to Improve Your AP Score
• Preview: Your Knowledge, Your Expectations
• Your Guide to Using This Book
• How to Begin
PREVIEW: YOUR KNOWLEDGE, YOUR EXPECTATIONS
Your route to a high score on the AP Biology Exam depends a lot on how you plan to use this book. Respond to the following questions.
1. Rate your level of confidence about your knowledge of the content tested by the AP Biology Exam.
A. Very confident—I know it all
B. I’m pretty confident, but there are topics for which I could use help
C. Not confident—I need quite a bit of support
D. I’m not sure
2. If you have a goal score in mind, highlight your goal score for the AP Biology Exam.
| 5 | 4 | 3 | 2 | 1 | I’m not sure yet |
3. What do you expect to learn from this book? Highlight all that apply to you.
A. A general overview of the test and what to expect
B. Strategies for how to approach the test
C. The content tested by this exam
D. I’m not sure yet
YOUR GUIDE TO USING THIS BOOK
This book is organized to provide as much—or as little—support as you need, so you can use this book in whatever way will be most helpful to improving your score on the AP Biology Exam.
• The remainder of Part I will provide guidance on how to use this book and help you determine your strengths and weaknesses.
• Part II of this book contains Practice Test 1 along with the answers and explanations. (Bubble sheets can be found before each test for easy reference.) We recommend that you take this test before going any further in order to realistically determine:
 your starting point right now
your starting point right now
 which question types you’re ready for and which you might need to practice
which question types you’re ready for and which you might need to practice
 which content topics you are familiar with and which you will want to carefully review
which content topics you are familiar with and which you will want to carefully review
Once you have nailed down your strengths and weaknesses with regard to this exam, you can focus your test preparation, build a study plan, and use your time efficiently.
• Part III of this book will
 provide information about the structure, scoring, and content of the AP Biology Exam
provide information about the structure, scoring, and content of the AP Biology Exam
 help you to make a study plan
help you to make a study plan
 point you towards additional resources
point you towards additional resources
• Part IV of this book will explore
 how to attack multiple-choice questions
how to attack multiple-choice questions
 how to write high-scoring free-response answers
how to write high-scoring free-response answers
 how to manage your time to maximize the number of points available to you
how to manage your time to maximize the number of points available to you
• Part V of this book covers the content you need for your exam.
• Part VI of this book contains Practice Tests 2, 3, and 4, and their answers and explanations. (Again, bubble sheets can be found before each test for easy reference.) Compare your progress between these tests and with Practice Test 1. If you get a certain type of question wrong several times, you probably need to review it. If you only got it wrong once, you may have run out of time or been distracted. In either case, this will allow you to focus on the factors that caused the discrepancy in scores and to be as prepared as possible on the day of the test.
You may choose to prioritize some parts of this book over others, or you may work through the entire book. This will depend on your needs and how much time you have. Let’s now look how to make this determination.
HOW TO BEGIN
1. Take Practice Test 1
Before you can decide how to use this book, you need to take a practice test. Doing so will give you insight into your strengths and weaknesses, and the test will also help you make an effective study plan. If you’re feeling test-phobic, remind yourself that a practice test is just a tool for diagnosing yourself—it’s not how well you do that matters. As long as you try your best, you can glean invaluable information from your performance to guide your preparation.
So, before you read further, take Practice Test 1 starting at this page of this book. Be sure to finish in one sitting, following the instructions that appear before the test.
2. Check Your Answers
Using the answer key on this page, count how many multiple-choice questions you got right and how many you missed. Don’t worry about the explanations for now and don’t worry about why you missed questions. We’ll get to that soon.
3. Reflect on the Test
After you take your first test, respond to the following questions:
• How much time did you spend on the multiple-choice questions?
• How much time did you spend on each free-response question?
• How many multiple-choice questions did you miss?
• Do you feel you had the knowledge to address the subject matter of the free-response questions?
• Do you feel you wrote well-organized, thoughtful responses to the free-response questions?
4. Read Part III of this Book and Complete the Self-Evaluation
As discussed in the Guide section above, Part III will provide information on how the test is structured and scored. It will also set out areas of content that are tested.
As you read Part III, re-evaluate your answers to the questions above. At the end of Part III, you will revisit and refine the questions you answer above. You will then be able to make a study plan, based on your needs and time available, that will allow you to use this book most effectively.
5. Engage with Parts IV and V as Needed
Notice the word engage. You’ll get more out of this book if you use it intentionally than if you read it passively, hoping for an improved score through osmosis.
The strategy chapters in Part IV will help you think about your approach to the question types on this exam. Part IV will open with a reminder to think about how you approach questions now and then close with a reflection section asking you to think about how or whether you will change your approach in the future.
The content chapters in Part V are designed to provide a review of the content tested on the AP Biology Exam, including the level of detail you need to know and how the content is tested. You will have the opportunity to assess your mastery of the content of each chapter through test-appropriate questions.
6. Take Another Test and Assess Your Performance
Once you feel you have developed the strategies you need and gained the knowledge you lacked, you should take Practice Test 2, which starts on this page of this book. You should finish in one sitting, following the instructions at the beginning of the test.
When you are done, check your answers to the multiple-choice sections. See if a teacher will read your essays and provide feedback.
Once you have taken the test, reflect on what areas you still need to work on and revisit the chapters in this book that address those deficiencies. Once you feel confident, take Practice Test 3, and repeat the process. Take Practice Test 4, and repeat the process again. Through this type of reflection and engagement, you will continue to improve.
7. Keep Working
After you have revisited certain chapters in this book, continue the process of testing, reflection, and engaging with the second practice test in this book. Consider what additional work you need to do and how you will change your strategic approach to different parts of the test.
As we will discuss in Part III, there are other resources available to you, including a wealth of information at AP Students, the official site of the AP Exams. You can continue to explore areas that can stand to improve and engage in those areas right up to the day of the test.
Part II
Practice Test 1
• Practice Test 1
• Practice Test 1: Answers and Explanations
Practice Test 1
Click here to download a PDF of Practice Test 1
The Exam
AP® Biology Exam
SECTION I: Multiple-Choice Questions
DO NOT OPEN THIS BOOKLET UNTIL YOU ARE TOLD TO DO SO.
At a Glance
Total Time
1 hour and 30 minutes
Number of Questions
69
Percent of Total Score
50%
Writing Instrument
Pencil required
Instructions
Section I of this examination contains 69 multiple-choice questions. These are broken down into Part A (63 multiple-choice questions) and Part B (6 grid-in questions).
Indicate all of your answers to the multiple-choice questions on the answer sheet. No credit will be given for anything written in this exam booklet, but you may use the booklet for notes or scratch work. After you have decided which of the suggested answers is best, completely fill in the corresponding oval on the answer sheet. Give only one answer to each question. If you change an answer, be sure that the previous mark is erased completely. Here is a sample question and answer.
Sample Question
Chicago is a
(A) state
(B) city
(C) country
(D) continent
Sample Answer

Use your time effectively, working as quickly as you can without losing accuracy. Do not spend too much time on any one question. Go on to other questions and come back to the ones you have not answered if you have time. It is not expected that everyone will know the answers to all the multiple-choice questions.
About Guessing
Many candidates wonder whether or not to guess the answers to questions about which they are not certain. Multiple choice scores are based on the number of questions answered correctly. Points are not deducted for incorrect answers, and no points are awarded for unanswered questions. Because points are not deducted for incorrect answers, you are encouraged to answer all multiple-choice questions. On any questions you do not know the answer to, you should eliminate as many choices as you can, and then select the best answer among the remaining choices.
BIOLOGY
SECTION I
69 Questions
Time—90 minutes
Directions: Each of the questions or incomplete statements below is followed by four suggested answers or completions. Select the one that is best in each case and then fill in the corresponding oval on the answer sheet.
1. The resting membrane potential depends on which of the following?
I. Active transport
II. Selective permeability
III. Differential distribution of ions across the axonal membrane
(A) III only
(B) I and II only
(C) II and III only
(D) I, II, and III
2. The Krebs cycle in humans occurs in the
(A) mitochondrial matrix
(B) inner mitochondrial membrane
(C) outer mitochondrial membrane
(D) intermembrane space
3. A heterotroph
(A) obtains its energy from sunlight, harnessed by pigments
(B) obtains its energy by catabolizing organic molecules
(C) makes organic molecules from CO2
(D) obtains its energy by consuming exclusively autotrophs
4. Regarding meiosis and mitosis, one difference between the two forms of cellular reproduction is that in meiosis
(A) there is one round of cell division, whereas in mitosis there are two rounds of cell division
(B) separation of sister chromatids occurs during the second division, whereas in mitosis separation of sister chromatids occurs during the first division
(C) chromosomes are replicated during interphase, whereas in mitosis chromosomes are replicated during the first phase of mitosis
(D) spindle fibers form during prophase, whereas in mitosis the spindle fibers form during metaphase
5. A feature of amino acids that is NOT found in carbohydrates is the presence of
(A) carbon atoms
(B) oxygen atoms
(C) nitrogen atoms
(D) hydrogen atoms
6. Which of the following is NOT a characteristic of bacteria?
(A) Circular double-stranded DNA
(B) Membrane-bound cellular organelles
(C) Plasma membrane consisting of lipids and proteins
(D) Ribosomes that synthesize polypeptides
7. Which of the following best explains why a population is described as the evolutionary unit?
(A) Genetic changes can only occur at the population level.
(B) The gene pool in a population remains fixed over time.
(C) Natural selection affects individuals, not populations.
(D) Individuals cannot evolve, but populations can.
8. The endocrine system maintains homeostasis using many feedback mechanisms. Which of the following is an example of positive feedback?
(A) Infant suckling causes a mother’s brain to release oxytocin, which in turn stimulates milk production.
(B) An enzyme is allosterically inhibited by the product of the reaction it catalyzes.
(C) When ATP is abundant the rate of glycolysis decreases.
(D) When blood sugar levels decrease to normal after a meal, insulin is no longer secreted.
9. A scientist carries out a cross between two guinea pigs, both of which have black coats. Black hair coat is dominant over white hair coat. Three quarters of the offspring have black coats, and one quarter have white coats. The genotypes of the parents were most likely
(A) bb × bb
(B) Bb × Bb
(C) Bb × bb
(D) BB × Bb
10. A large island is devastated by a volcanic eruption. Most of the horses die except for the heaviest males and heaviest females of the group. They survive, reproduce, and perpetuate the population. If weight is a highly heritable trait, which graph represents the change in population before and after the eruption?
(A) A higher mean weight compared with their parents
(B) A lower mean weight compared with their parents
(C) The same mean weight as members of the original population
(D) A higher mean weight compared with members of the original population
11. All of the following play a role in morphogenesis EXCEPT
(A) apoptosis
(B) homeotic genes
(C) operons
(D) inductive effects
12. During the period when life is believed to have begun, the atmosphere on primitive Earth contained abundant amounts of all the following gases EXCEPT
(A) oxygen
(B) hydrogen
(C) ammonia
(D) methane
Questions 13–14 refer to the following passage.
The digestive system in humans can be divided into two parts: the alimentary canal and the accessory organs. The canal is where the food actually passes during its transition into waste. The accessory organs are any organs that aid in the digestion by supplying the organs in the alimentary canal with digestive hormones and enzymes.
13. The small intestine is the main site of absorption. It can accomplish it so efficiently because of villi and microvilli that sculpt the membrane into hair-like projections. They likely aid in reabsorption by
(A) increasing the surface area of the small intestine
(B) decreasing the surface area of the small intestine
(C) making the small intestine more hydrophilic
(D) making the small intestine more hydrophobic
14. The pancreas is a major accessory organ in the digestive system. What is its likely function?
(A) Moistening the food as it passes through
(B) Removing the excess water from the food waste
(C) Making enzymes that breakdown macromolecules
(D) Preventing the food from entering the small intestine after it leaves the stomach
15. In animal cells, which of the following represents the most likely pathway that a secretory protein takes as it is synthesized in a cell?
(A) Plasma membrane–Golgi apparatus–ribosome–secretory vesicle–rough ER
(B) Ribosome–Golgi apparatus–rough ER–secretory vesicle–plasma membrane
(C) Plasma membrane–Golgi apparatus–ribosome–secretory vesicle–rough ER
(D) Ribosome–rough ER–Golgi apparatus–secretory vesicle–plasma membrane
16. All of the following statements are correct regarding alleles EXCEPT
(A) alleles are alternative forms of the same gene
(B) alleles are found on corresponding loci of homologous chromosomes
(C) a gene can have more than two alleles
(D) an individual with two identical alleles is said to be heterozygous with respect to that gene
17. Once specific genes, such as the gene coding for ampicillin, have been incorporated into a plasmid, the plasmid may be cloned by
(A) inserting it into a virus to generate multiple copies
(B) treating it with a restriction enzyme in order to cut the molecule into small pieces
(C) inserting it into a suitable bacterium in order to produce multiple copies
(D) running it on a gel electrophoresis in order to determine the size of the gene of interest
18. Although mutations occur at a regular and predictable rate, which of the following statements is the LEAST likely reason the frequency of mutation often appears to be low?
(A) Some mutations produce alleles that are recessive and may not be expressed.
(B) Some undesirable phenotypic traits may be prevented from reproducing.
(C) Some mutations cause such drastic phenotypic changes that they are soon removed from the gene pool.
(D) The predictable rate of mutation results in ongoing variability in a gene pool.
19. A scientist wants to test the effect of temperature on seed germination. Which of the following should be part of the experimental design?
(A) Use temperature as the dependent variable and alter the germination times
(B) Use temperature as the independent variable and measure the rate of germination
(C) Use temperature as the controlled variable and keep everything identical between groups
(D) Use the variable natural outside temperature as a control group
20. Which of the following best accounts for the ability of legumes to grow well in nitrogen-poor soils?
(A) These plants make their own proteins.
(B) These plants have a mutualistic relationship with nitrogen-fixing bacteria.
(C) These plants are capable of directly converting nitrogen gas into nitrates.
(D) These plants do not require nitrogen to make plant proteins.
21. Which of the following is most correct concerning cell differentiation in vertebrates?
(A) Cells in different tissues contain different sets of genes, leading to structural and functional differences.
(B) Differences in the timing and expression levels of different genes lead to structural and functional differences.
(C) Differences in the reading frame of mRNA lead to structural and functional differences.
(D) Differences between tissues result from spontaneous morphogenesis.
Questions 22–23 refer to the following passage.
Pumping blood through the human heart must be carefully organized for maximal efficiency and to prevent backflow. In the figure below, the blood enters the heart through the vena cava, passes through the right atrium and right ventricle and then goes through the pulmonary artery towards the lungs. After the lungs, the blood returns through the pulmonary vein and then passes into the left atrium and the left ventricle before leaving the heart via the aorta.
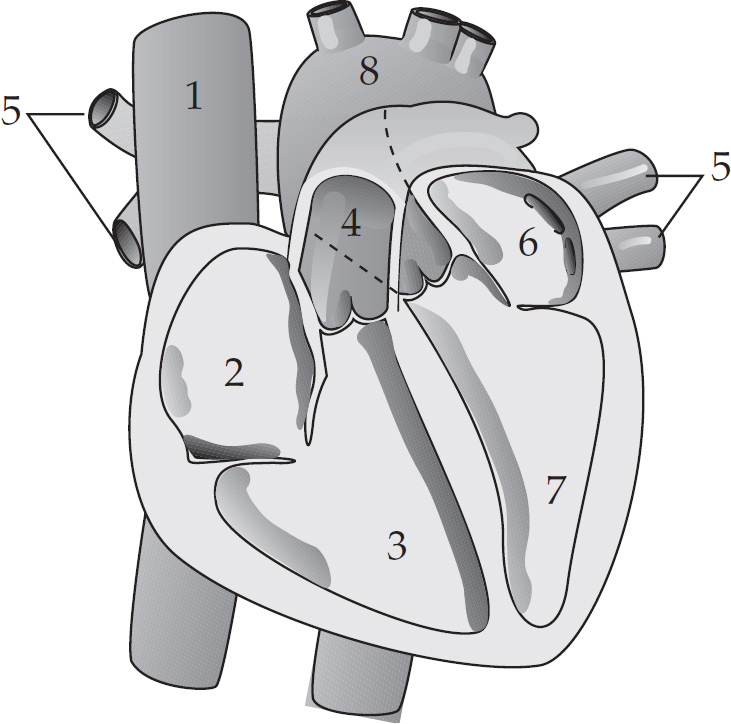
22. Which of the following chambers or vessels carry deoxygenated blood in the human heart?
(A) 1 only
(B) 2 and 3
(C) 1, 2, 3, 4
(D) 4 and 5
23. If a small amount of short-lived dye was injected into the left ventricle, which heart chambers would have dye 30 seconds later?
(A) All chambers of the heart
(B) Only the ventricles
(C) Only the left atrium and the left ventricle
(D) Only the left ventricle
24. Some strains of viruses can change normal mammalian cells into cancer cells in vitro. This alteration of the mammalian cell is usually associated with the
(A) formation of a pilus between the mammalian cell and the virus
(B) incorporation of the viral genome into the mammalian cell’s nuclear DNA
(C) conversion of the host’s genome into the viral DNA
(D) release of spores into the mammalian cell
25. All of the following correctly describe meiosis EXCEPT
(A) Meiosis produces four haploid gametes
(B) Homologous chromosomes join during synapsis
(C) Sister chromatids separate during meiosis I
(D) Crossing over increases genetic variation in gametes
26. All of the following are examples of events that can prevent interspecies breeding EXCEPT
(A) the potential mates experience geographic isolation
(B) the potential mates experience behavioral isolation
(C) the potential mates have different courtship rituals
(D) the potential mates have similar breeding seasons
27. Which of the following is NOT a characteristic of asexual reproduction in animals?
(A) Progeny cells have the same number of chromosomes as the parent cell.
(B) Progeny cells are identical to the parent cell.
(C) The parent cell produces diploid cells.
(D) The progeny cells fuse to form a zygote.
28. Transpiration is a result of special properties of water. The special properties of water include all of the following EXCEPT
(A) cohesion
(B) adhesion
(C) capillary action
(D) hydrophobicity
Questions 29–32 refer to the following passage.
An experiment was performed to assess the growth of two species of plants when they were grown in different pHs, given different volumes of water, and watered at different times of day over 6 weeks. Two plants were grown of each species and the average heights are shown in the table.
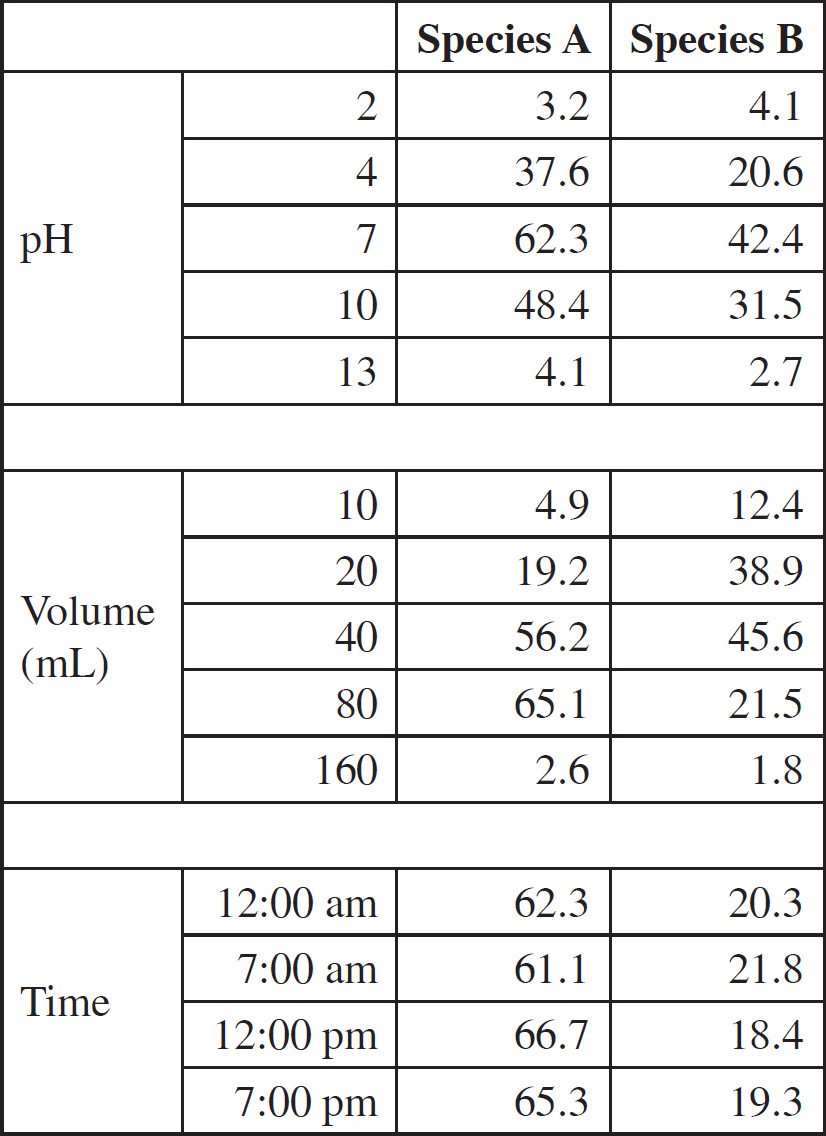
29. For which conditions do the species have different preferences?
(A) pH
(B) volume
(C) volume and watering time
(D) pH and volume and watering time
30. What are the preferred growth conditions for Species B?
(A) pH 7, 40 mL, any time of day
(B) pH 10, 40 mL, 7:00 A.M.
(C) pH 7, 80 mL, any time of day
(D) pH 10, 80 mL, 12:00 P.M.
31. Which pH and volume were likely used for the watering time experiment?
(A) pH 4 and 40 mL
(B) pH 10 and 40 mL
(C) pH 4 and 80 mL
(D) pH 10 and 80 mL
32. Which of the following would most improve the statistical significance of the results?
(A) Let the plants grow for a longer period of time
(B) Add more conditions to test, such as amount of light and amount of soil
(C) Test the same plants with more pHs and more volumes and times of day
(D) Increase the number of plants in each group
33. A plant grows in the opposite direction of the gravitational force. This is an example of
(A) positive thignotropism
(B) negative phototropism
(C) positive phototropism
(D) negative gravitropism
34. In most ecosystems, net primary productivity is important because it represents the
(A) energy available to producers
(B) total solar energy converted to chemical energy by producers
(C) biomass of all producers
(D) energy available to heterotrophs
35. Hawkmoths are insects that are similar in appearance and behavior to hummingbirds. Which of the following is LEAST valid?
(A) These organisms are examples of convergent evolution.
(B) These organisms were subjected to similar environmental conditions.
(C) These organisms are genetically related to each other.
(D) These organisms have analogous structures.
36. Which of the following describes a mutualistic relationship?
(A) A tapeworm feeds off of its host’s nutrients causing the host to lose large amounts of weight.
(B) Certain plants grow on trees in order to gain access to sunlight, not affecting the tree.
(C) Remora fish eat parasites off of sharks. The sharks stay free of parasites, and the remora fish are protected from predators.
(D) Meerkats sound alarm calls to warn other meerkats of predators.
37. The pancreas is an organ that makes insulin and glucagon in its beta and alpha cells, respectively. Insulin is released when blood glucose is high and glucagon is released when blood glucose in low. Anti-beta cell antibodies will cause which of the following to occur?
(A) Glucagon secretion will stop, and blood glucose levels will not decrease.
(B) Glucagon secretion will stop, and blood glucose levels will decrease.
(C) Glucagon secretion will stop, and digestive enzymes will be secreted.
(D) Insulin secretion will stop, and blood glucose levels will not decrease.
Questions 38–40 refer to the following passage.
The rainfall and biomass of several trophic levels in an ecosystem were measured over several years. The results are shown in the graph below.
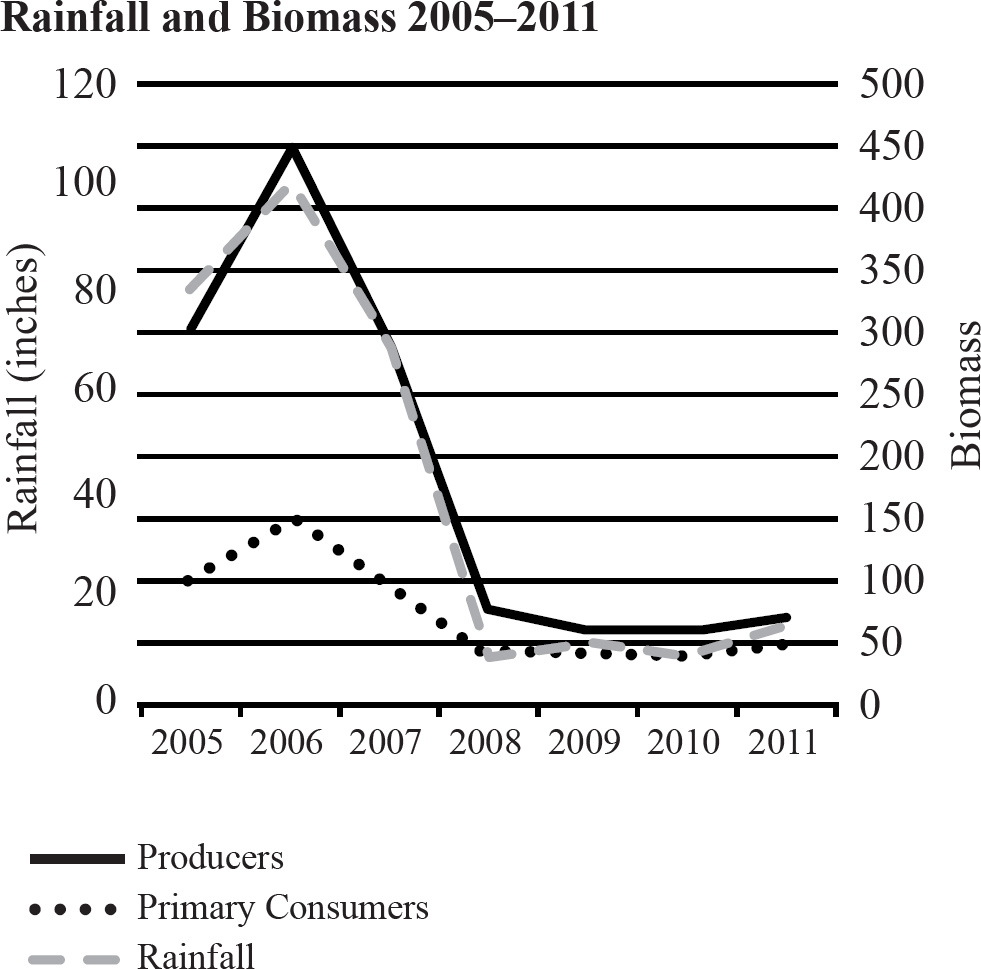
38. Which of the following concepts is best demonstrated by this experiment?
(A) Populations with higher genetic variation can withstand droughts better.
(B) Meteorological impacts will affect the evolution of populations.
(C) Environmental changes can affect all the levels of the ecosystem.
(D) Unoccupied biological niches are dangerous because they attract invasive species.
39. If it rained 120 inches, what would you project the primary consumer biomass to be?
(A) 150–200
(B) 60
(C) 45
(D) 20
40. Which of the following graphs best depicts the projected biomass of secondary consumers if they were measured?
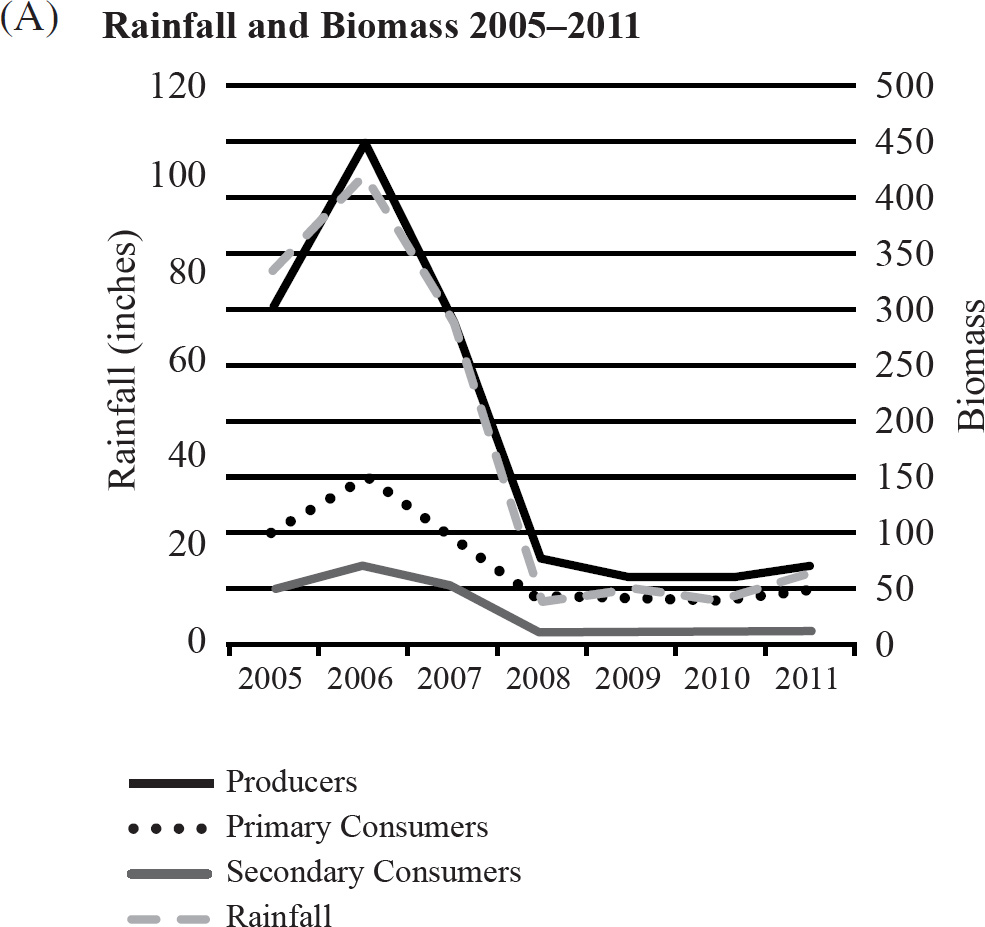
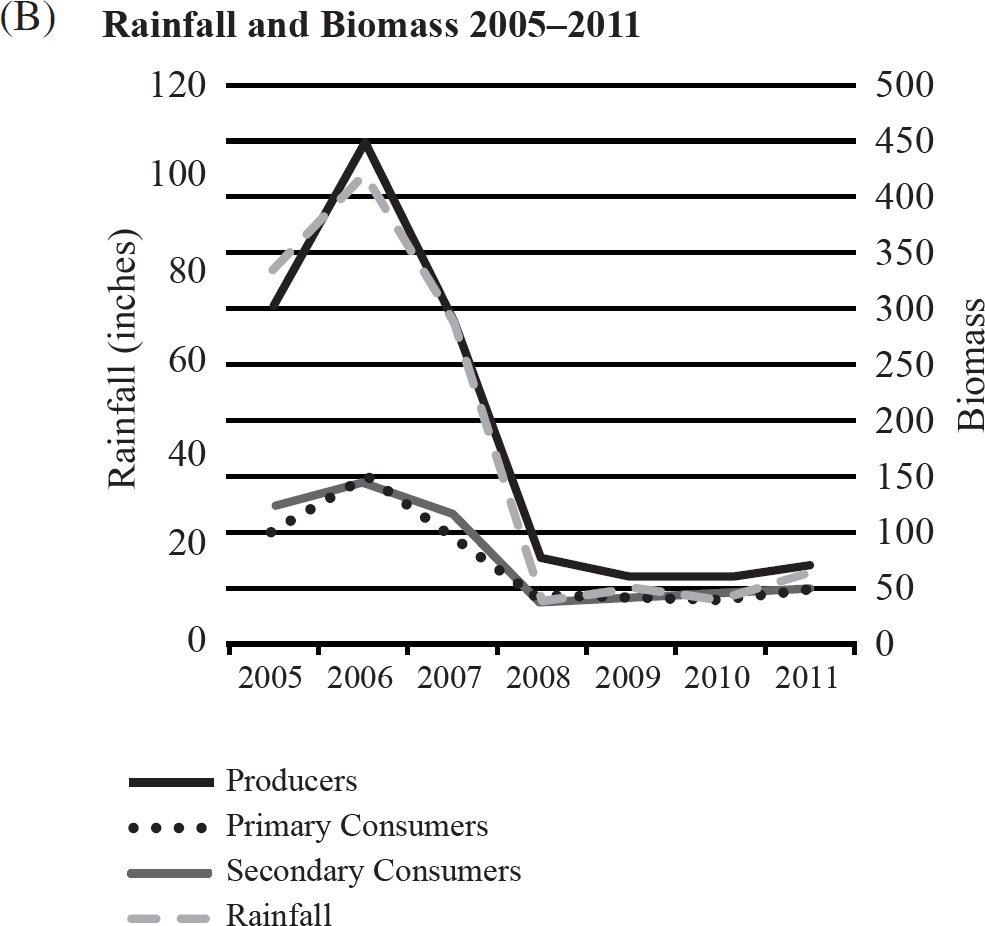
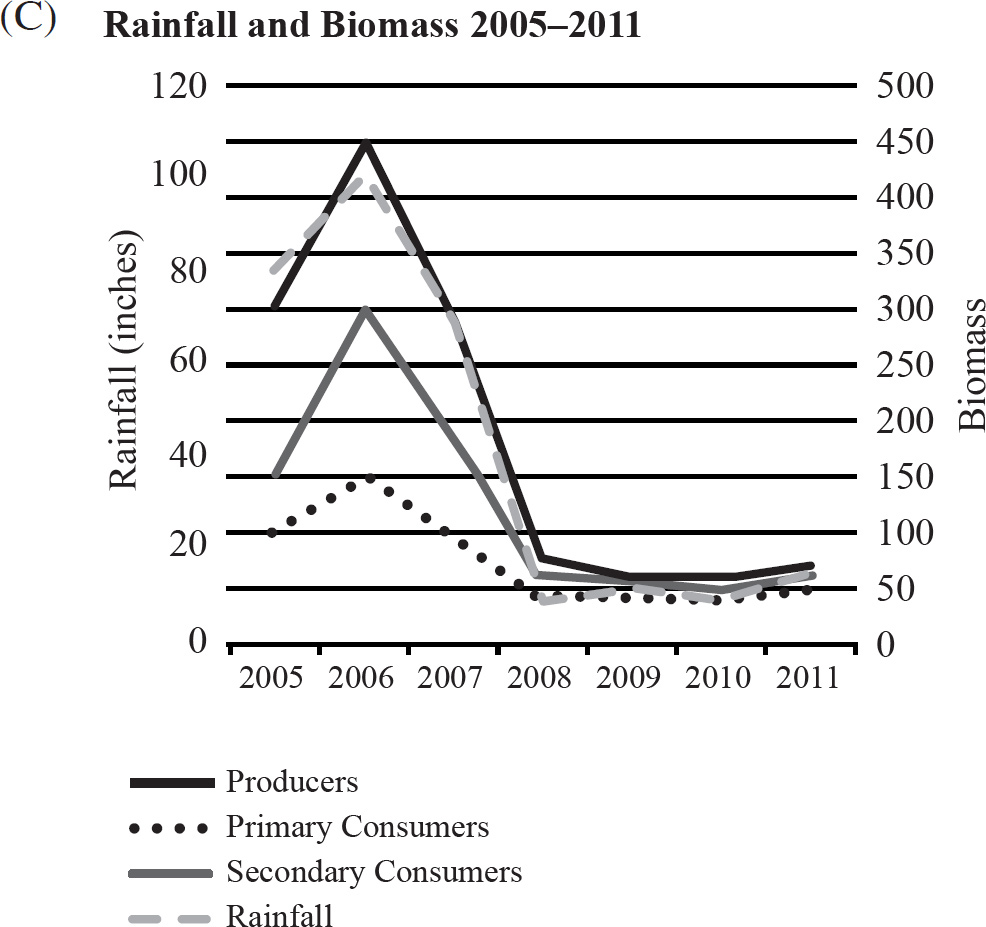
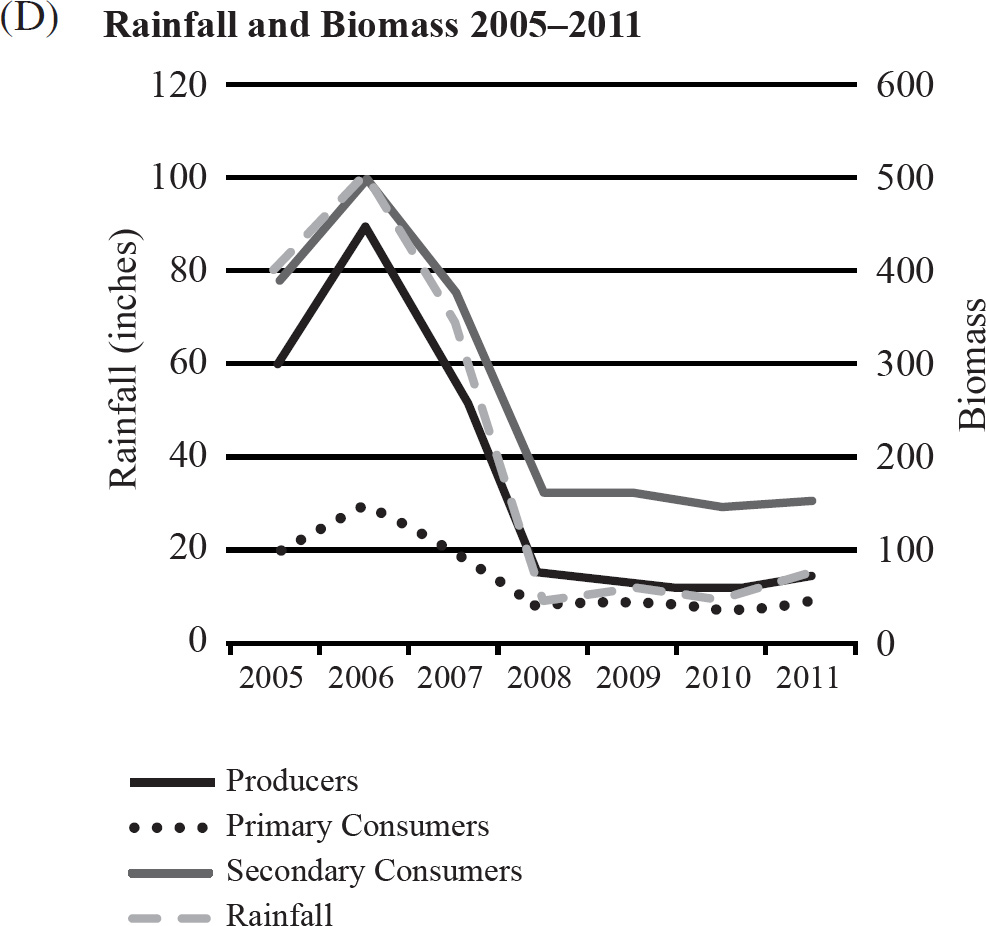
41. The calypso orchid, Calypso bulbosa, grows in close association with mycorrhizae fungi. The fungi penetrate the roots of the flower and take advantage of the plant’s food resources. The fungi concentrate rare minerals, such as phosphates, in the roots and make them readily accessible to the orchid. This situation is an example of
(A) parasitism
(B) commensalism
(C) mutualism
(D) endosymbiosis
42. Which of the following are characteristics of both bacteria and fungi?
(A) Cell wall, DNA, and plasma membrane
(B) Nucleus, organelles, and unicellularity
(C) Plasma membrane, multicellularity, and Golgi apparatus
(D) Cell wall, unicellularity, and mitochondria
43. The synthesis of new proteins necessary for lactose utilization by the bacterium E. coli using the lac operon is regulated
(A) by the synthesis of additional ribosomes
(B) at the transcription stage
(C) at the translation stage
(D) by differential replication of the DNA that codes for lactose-utilizing mechanisms
44. Trypsin is a digestive enzyme. It cleaves polypeptides after lysine and arginine amino acid residues. Which of the following statements about trypsin is NOT true?
(A) It is an organic compound made of proteins.
(B) It is a catalyst that alters the rate of a reaction.
(C) It is operative over a wide pH range.
(D) The rate of catalysis is affected by the concentration of substrate.
45. In DNA replication, which of the following does NOT occur?
(A) Helicase unwinds the double helix.
(B) DNA ligase links the Okazaki fragments.
(C) RNA polymerase is used to elongate both chains of the helix.
(D) DNA strands grow in the 5’ to 3’ direction.
46. A change in a neuron membrane potential from +50 millivolts to –70 millivolts is considered
(A) depolarization
(B) repolarization
(C) hyperpolarization
(D) an action potential
47. The energy given up by electrons as they move through the electron transport chain is used to
(A) break down glucose
(B) make glucose
(C) produce ATP
(D) make NADH
48. If a plant undergoing the light-dependent reactions of photosynthesis began to release 18O2 instead of normal oxygen, one could most reasonably conclude that the plant had been supplied with
(A) H2O containing radioactive oxygen
(B) CO2 containing radioactive oxygen
(C) C6H12O6 containing radioactive oxygen
(D) NO2 containing radioactive oxygen
49. Chemical substances released by organisms that elicit a physiological or behavioral response in other members of the same species are known as
(A) auxins
(B) hormones
(C) pheromones
(D) enzymes
50. Homologous structures are often cited as evidence for the process of natural selection. All of the following are examples of homologous structures EXCEPT
(A) the forearms of a cat and the wings of a bat
(B) the flippers of a whale and the arms of a man
(C) the pectoral fins of a porpoise and the flippers of a seal
(D) the forelegs of an insect and the forelimbs of a dog
51. Certain populations of finches have long been isolated on the Galapagos Islands off the western coast of South America. Compared with the larger stock population of mainland finches, these separate populations exhibit far greater variation over a wider range of species. The variation among these numerous finch species is the result of
(A) convergent evolution
(B) divergent evolution
(C) disruptive selection
(D) stabilizing selection
52. Which of the following contributes the MOST to genetic variability in a population?
(A) Sporulation
(B) Binary fission
(C) Vegetative propagation
(D) Mutation
Questions 53–55 refer to the following information and table.
A marine ecosystem was sampled in order to determine its food chain. The results of the study are shown below.
| Type of Organism | Number of Organisms |
| Shark | 2 |
| Small crustaceans | 400 |
| Mackerel | 20 |
| Phytoplankton | 1,000 |
| Herring | 100 |
53. Which of the following organisms in this population are secondary consumers?
(A) Sharks
(B) Phytoplankton
(C) Herrings
(D) Small crustaceans
54. Which of the following organisms has the largest biomass in this food chain?
(A) Phytoplanktons
(B) Mackerels
(C) Herrings
(D) Sharks
55. If the herring population is reduced by predation, which of the following would most likely be a secondary effect on the ecosystem?
(A) The mackerels will be the largest predator in the ecosystem.
(B) The small crustacean population will be greatly reduced.
(C) The phytoplankton population will be reduced over the next year.
(D) The small crustaceans will become extinct.
Questions 56–58 refer to the following information and diagram.
To understand the workings of neurons, an experiment was conducted to study the neural pathway of a reflex arc in frogs. A diagram of a reflex arc is given below.
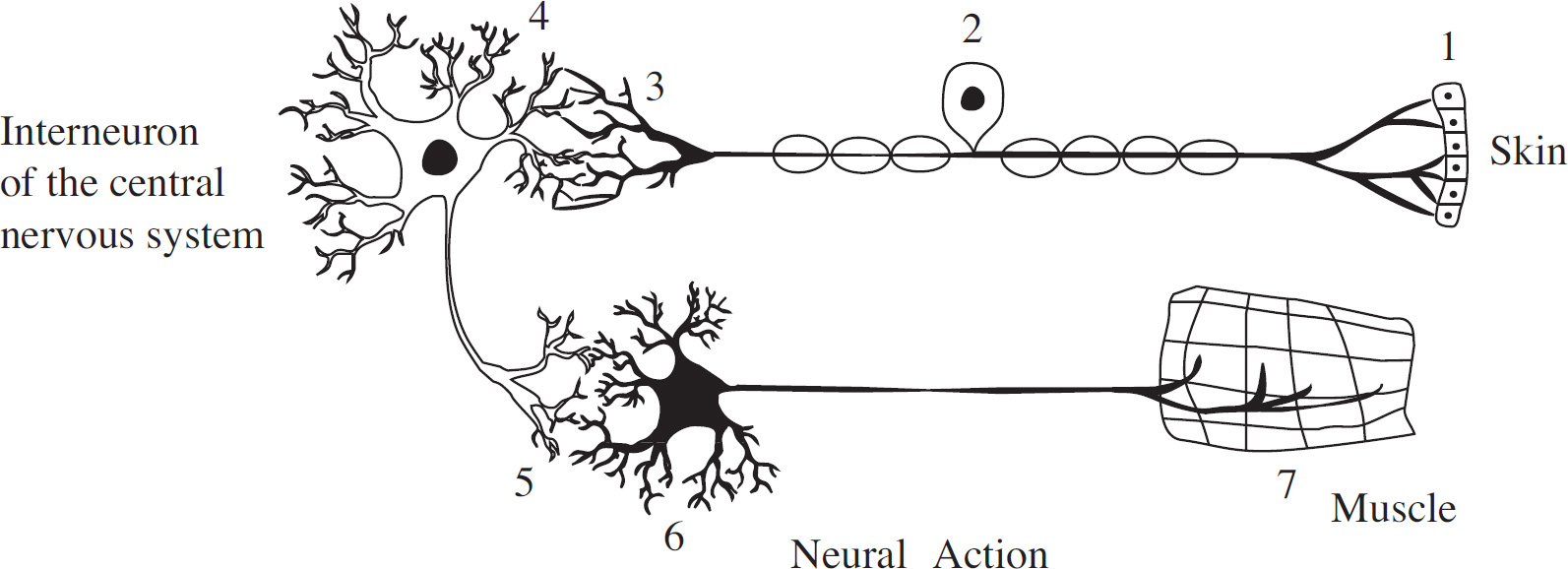
56. Which of the following represents the correct pathway taken by a nerve impulse as it travels from the spinal cord to effector cells?
(A) 1-2-3-4
(B) 6-5-4-3
(C) 2-3-4-5
(D) 4-5-6-7
57. The brain of the frog is destroyed. A piece of acid-soaked paper is applied to the frog’s skin. Every time the piece of paper is placed on its skin, one leg moves upward. Which of the following conclusions is best supported by the experiment?
(A) Reflex actions are not automatic.
(B) Some reflex actions can be inhibited.
(C) All behaviors in frogs are primarily reflex responses.
(D) This reflex action does not require the brain.
58. A nerve impulse requires the release of neurotransmitters at the axonal bulb of a presynaptic neuron. Which of the following best explains the purpose of neurotransmitters, such as acetylcholine?
(A) They speed up the nerve conduction in a neuron.
(B) They open the sodium channels in the axonal membrane.
(C) They excite or inhibit the postsynaptic neuron.
(D) They open the potassium channels in the axonal membrane.
Questions 59–61 refer to the figure and chart below.
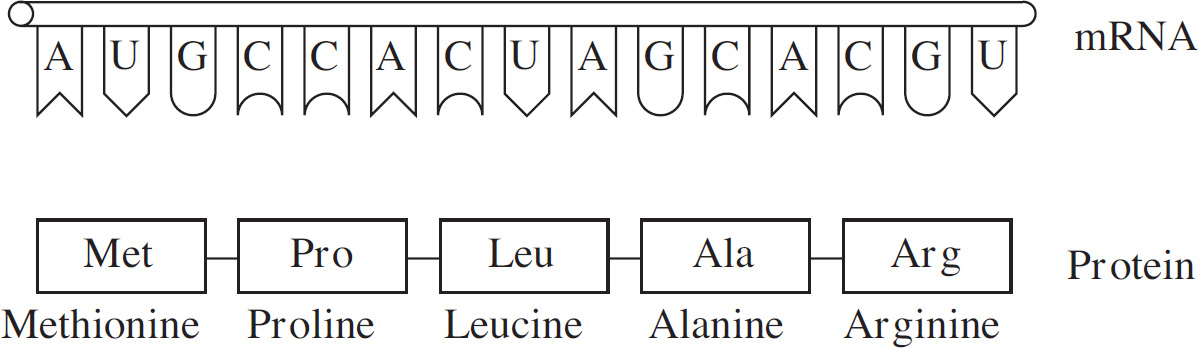
Formation of a Protein
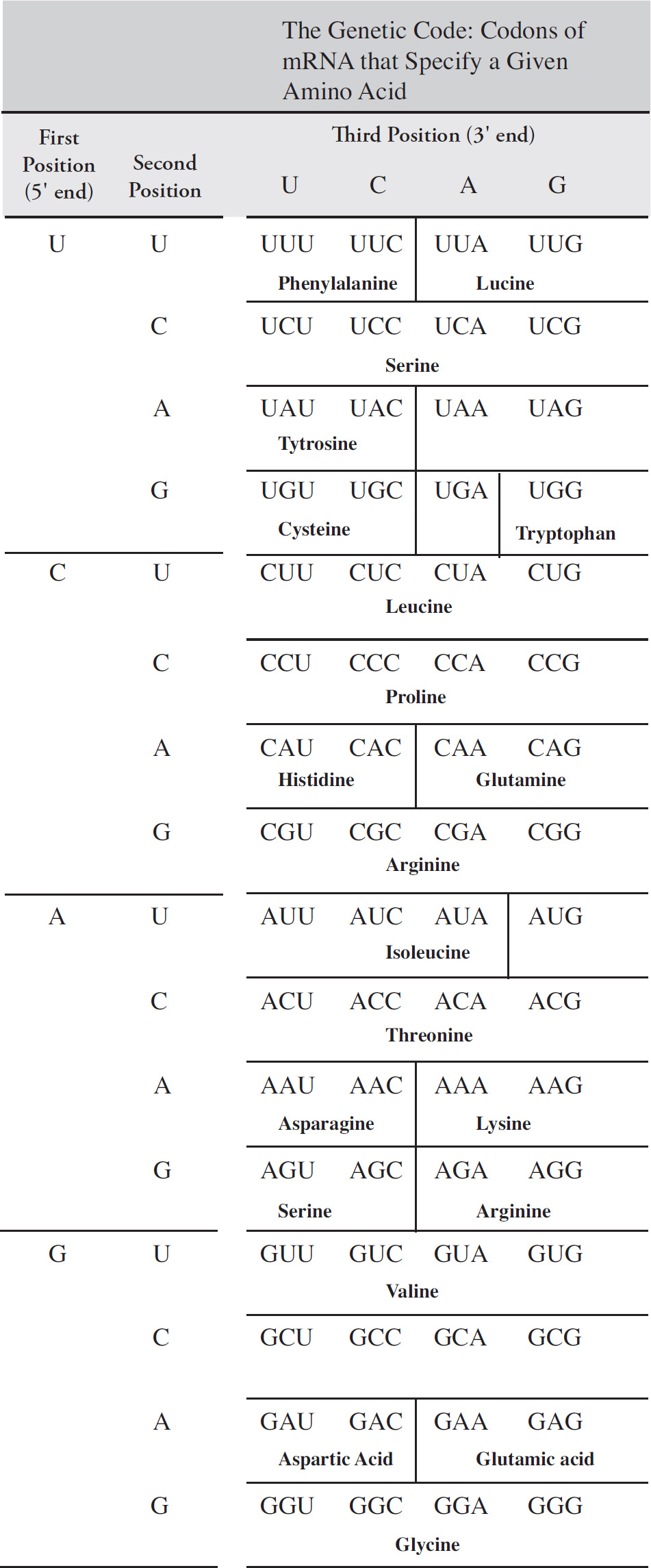
59. Which of the following DNA strands is the template strand that led to the amino acid sequence shown above?
(A) 3′-ATGCGACCAGCACGT-5′
(B) 3′-AUGCCACUAGCACGU-5′
(C) 3′-TACGGTGATCGTGCA-5′
(D) 3′-UACGGUGAUCGUGCA-5′
60. Immediately after the translation of methionine, a chemical is added which deletes all remaining uracil nucleotides in the mRNA, which of the following represents the resulting amino acid sequence?
(A) Serine–histidine–serine–threonine
(B) Methionine–proline–glutamine–histidine
(C) Methionine–proline–leucine–alanine–arginine
(D) Methionine–proline–alanine–arginine–arginine
61. The mRNA above was found to be much smaller than the original mRNA synthesized in the nucleus. This is due to the
(A) addition of a poly(A) tail to the mRNA molecule
(B) addition of a cap to the mRNA molecule
(C) excision of exons from the mRNA molecule
(D) excision of introns from the mRNA molecule
Questions 62 and 63 refer to the following information.
A scientist studies the storage and distribution of oxygen in humans and Weddell seals to examine the physiological adaptations that permit seals to descend to great depths and stay submerged for extended periods. The figure below depicts the oxygen storage in both organisms.
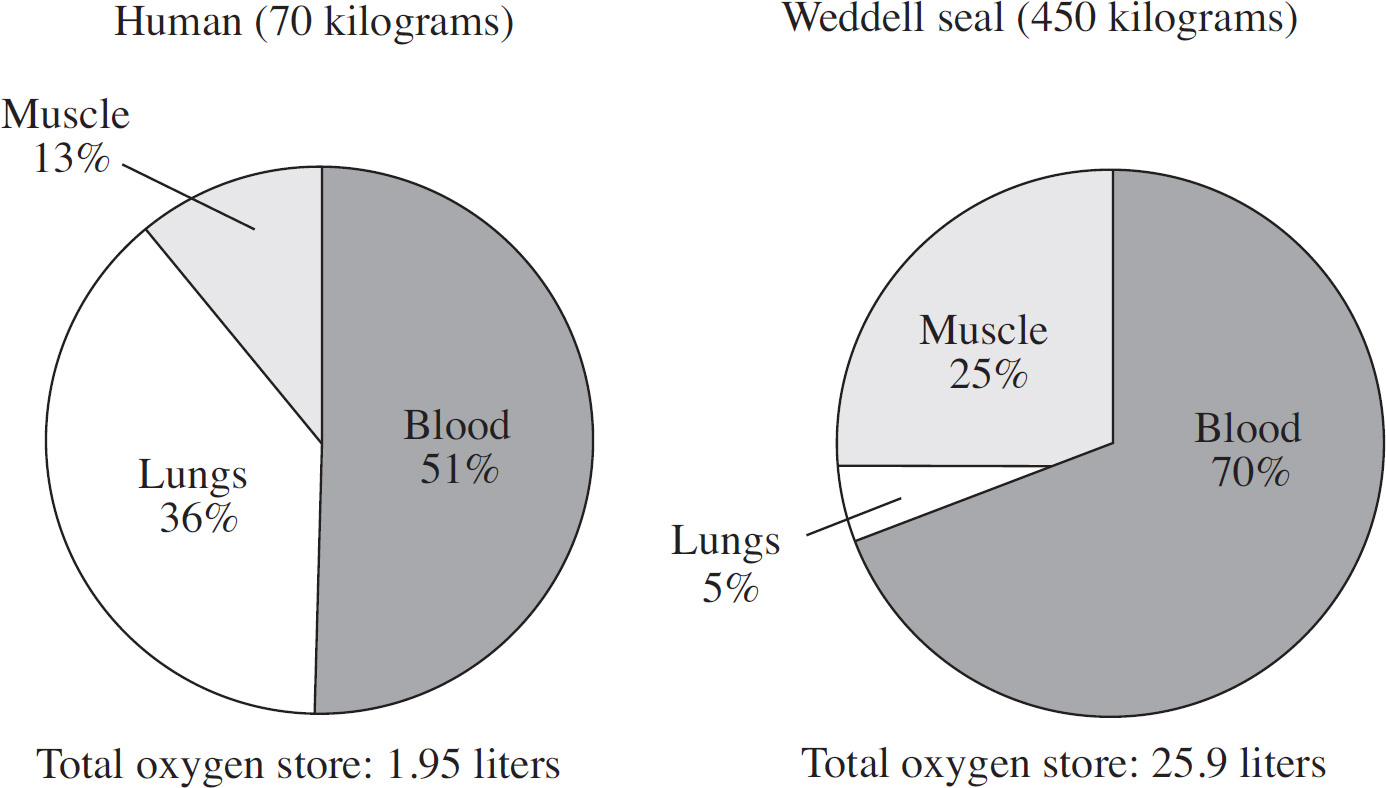
62. Compared with humans, approximately how many liters of oxygen does the Weddell seal store per kilogram of body weight?
(A) The same amount of oxygen
(B) Twice the amount of oxygen
(C) Three times the amount of oxygen
(D) Five times the amount of oxygen
63. During a dive, a Weddell seal’s blood flow to the abdominal organs is shut off, and oxygen-rich blood is diverted to the eyes, brain, and spinal cord. Which of the following is the most likely reason for this adaptation?
(A) To increase the number of red blood cells in the nervous system
(B) To increase the amount of oxygen reaching the skeletomuscular system
(C) To increase the amount of oxygen reaching the central nervous system
(D) To increase the oxygen concentration in the lungs
Directions: Part B consists of questions requiring numeric answers. Calculate the correct answer for each question.
64. An experiment was conducted to observe the light-absorbing properties of chlorophylls and carotenoids using a spectrophotometer. The pigments were first extracted and dissolved in a solution. They were then illuminated with pure light of different wavelengths to detect which wavelengths were absorbed by the solution. The results are presented in the absorption spectrum below.
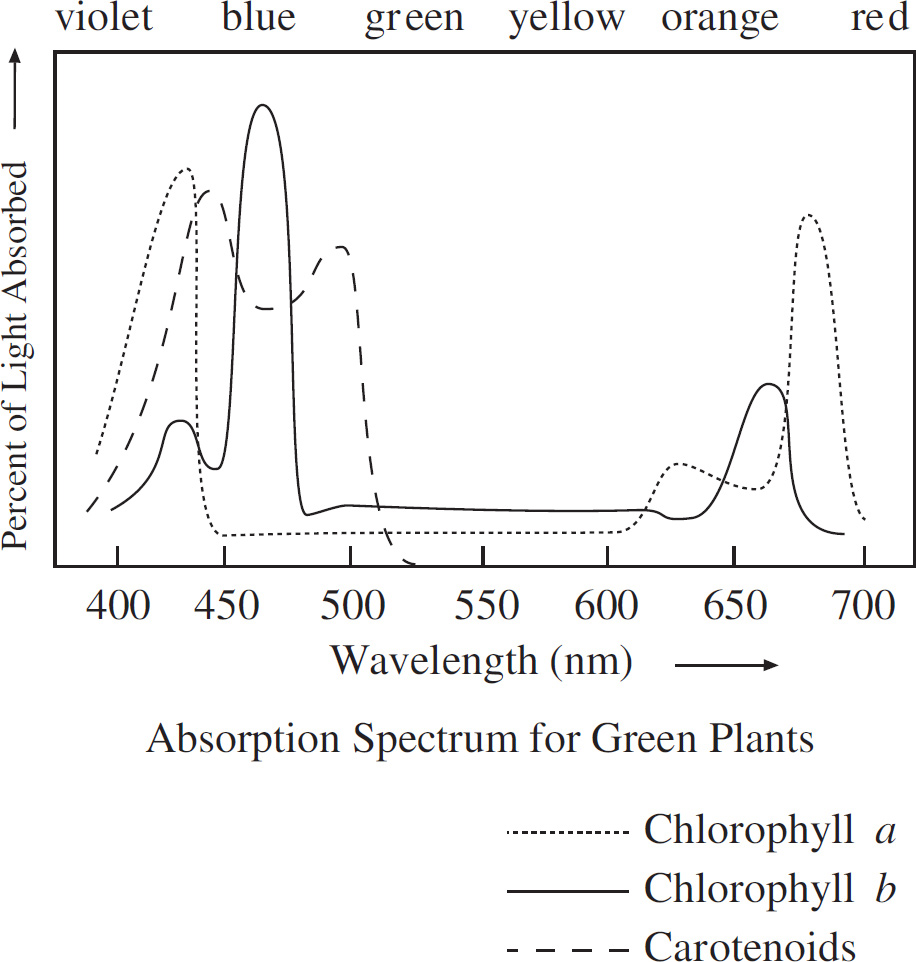
At approximately what wavelength does chlorophyll a maximally absorb light?

65. A woman with blood genotype IAi and a man with blood genotype IBi have two children, both type AB. What is the probability that a third child will be blood type AB?

66. The trophic level efficiency of large herbivores such as elks is frequently only about 5 percent. In tons, what volume of plants would be required to maintain 24,000 lbs of elk?

67. If the genotype frequencies of an insect population are AA = 0.49, Aa = 0.42, and aa = 0.09, what is the gene frequency of the recessive allele?

Question 68 refers to the diagram below.
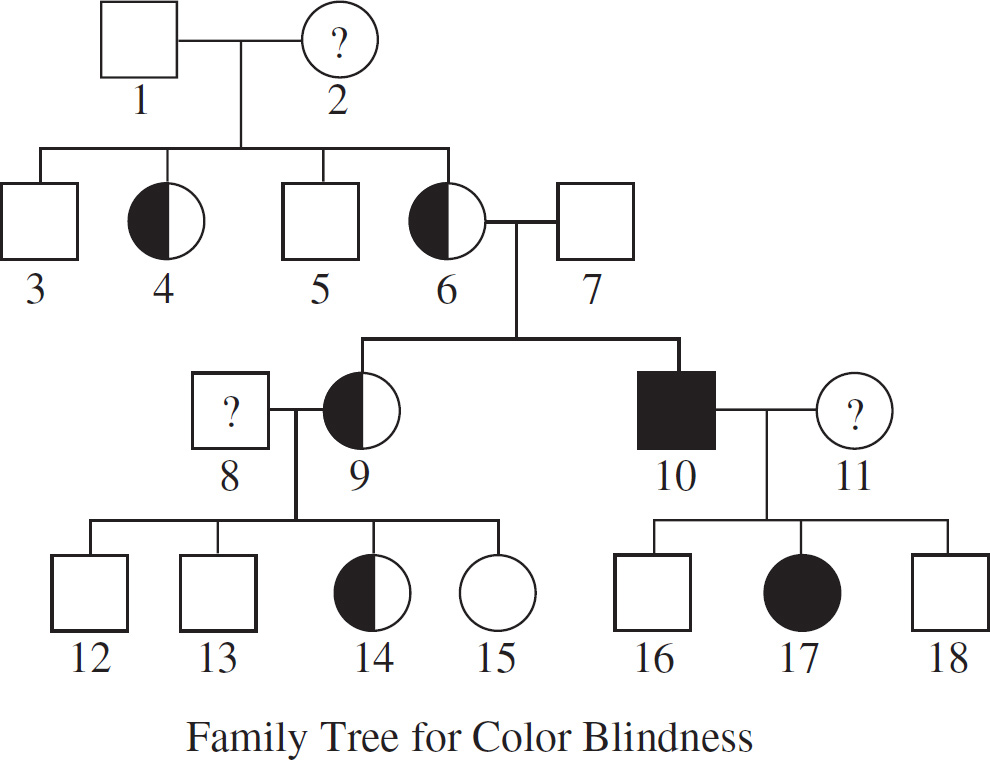
68. Based on the pedigree above, what is the probability that a male child born to individuals 6 and 7 will be color-blind?

69. The loss of water by evaporation from the leaf openings is known as transpiration. The transpiration rates of various plants are shown below.
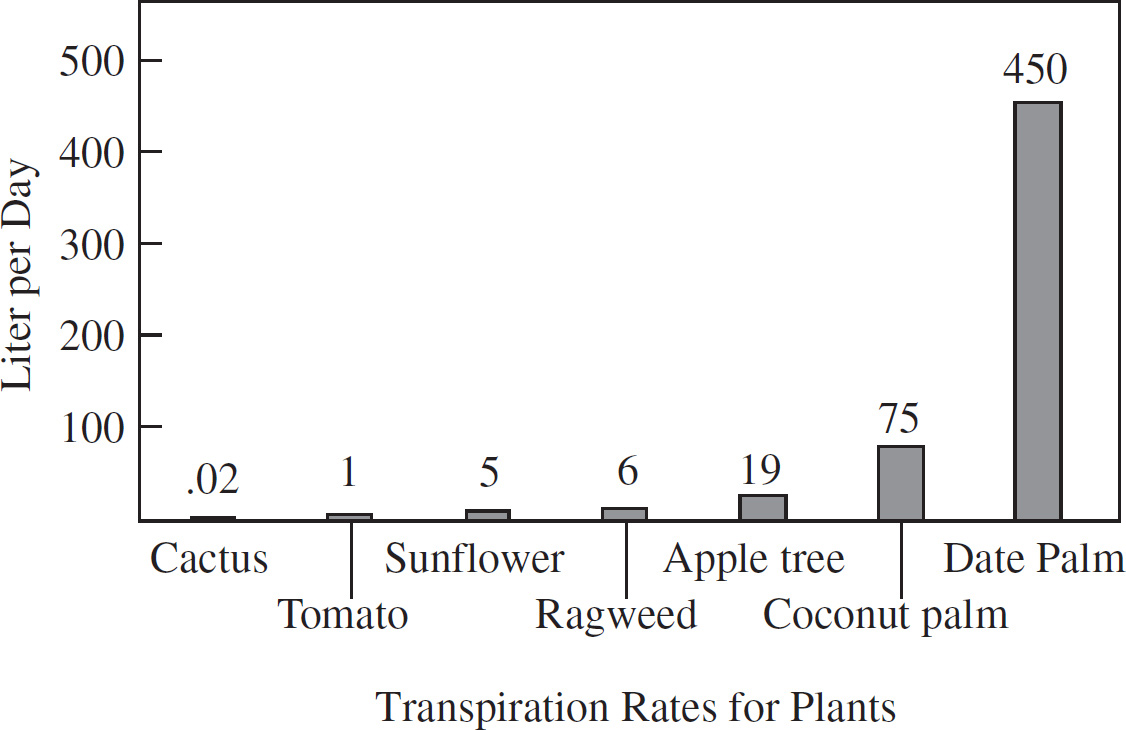
How many liters of water per week are lost by a coconut palm?

STOP
END OF SECTION I
IF YOU FINISH BEFORE TIME IS CALLED, YOU MAY CHECK YOUR WORK ON THIS SECTION. DO NOT GO ON TO SECTION II UNTIL YOU ARE TOLD TO DO SO.
BIOLOGY
SECTION II
8 Questions
Planning Time—10 minutes
Writing Time—80 minutes
Directions: Questions 1 and 2 are long free-response questions that should require about 22 minutes each to answer and are worth 10 points each. Questions 3 through 8 are short free-response questions that should require about 6 minutes each to answer. Questions 3 through 5 are worth 4 points each, and questions 6 through 8 are worth 3 points each.
Read each question carefully and completely. Write your response in the space provided following each question. Only material written in the space provided will be scored. Answers must be written out in paragraph form. Outlines, bulleted lists, or diagrams alone are not acceptable.
1. The cell membrane is an important structural feature of a nerve cell.
(a) Describe the role of the sodium potassium pump in maintaining the resting membrane potential.
(b) Discuss ion flow during an action potential.
(c) Predict the outcome on action potentials if a cell could not make voltage-gated sodium channels.
(d) Explain how myelin affects the speed of an action potential.
2. Sickle-cell anemia is a genetic disorder caused by the abnormal gene for hemoglobin S. A single substitution occurs in which glutamic acid is substituted for valine in the sixth position of the hemoglobin molecule. This change reduces hemoglobin’s ability to carry oxygen.
(a) Discuss the process by which mutation occurs in base substitution.
(b) Biologists used gel electrophoresis to initially identify the mutant gene. Explain how gel electrophoresis could be applied to the identification of the gene mutation. Discuss the use of restriction enzymes.
(c) Hemoglobin S is transmitted as a simple Mendelian allele. Describe the outcome if a female who does not carry the abnormal allele mates with a male homozygous for the disease. Include a Punnett square and phenotypic and genotypic ratios.
3. Cell size is limited by the surface area-to-volume ratio of the cell membrane.
(a) Discuss why cell size is limited by this ratio.
(b) Describe two adaptations that increase surface area in organisms.
(c) Describe the difference in how small polar and small nonpolar molecules cross cell membranes
4. Discuss the Krebs cycle and the electron transport chain, and chemiosmosis.
(a) Explain why these steps are considered aerobic processes.
(b) Discuss the location at which each stage occurs.
5. Describe three main differences between meiosis and mitosis.
6. Define homologous structures and give an example. Describe how they are different from analogous structures.
7. Describe the three ways that genetic information is transmitted laterally between bacteria.
8. Describe why viruses are typically not considered to be alive.
STOP
END OF EXAM
Practice Test 1: Answers and Explanations
PRACTICE TEST 1 ANSWER KEY
1. D
2. A
3. B
4. B
5. C
6. B
7. D
8. A
9. B
10. D
11. C
12. A
13. A
14. C
15. D
16. D
17. C
18. D
19. B
20. B
21. B
22. C
23. D
24. B
25. C
26. D
27. D
28. D
29. B
30. A
31. D
32. D
33. D
34. D
35. C
36. C
37. D
38. C
39. A
40. A
41. C
42. A
43. B
44. C
45. C
46. B
47. C
48. A
49. C
50. D
51. B
52. D
53. C
54. A
55. C
56. D
57. D
58. C
59. C
60. B
61. D
62. B
63. C
64. 425–440
65. 1/4 or 0.25 or 25%
66. 240
67. 0.3 or 30%
68. 1/2 or 0.5 or 50%
69. 525
PRACTICE TEST 1 EXPLANATIONS
Section I: Multiple Choice
1. D The resting potential depends on active transport (the Na+K+-ATPase pump) and the selective permeability of the axon membrane to K+ than to Na+, which leads to a differential distribution of ions across the axonal membrane.
2. A The Krebs cycle occurs in the mitochondrial matrix. Don’t forget to review the site of each stage of aerobic respiration. Glycolysis, the first step in aerobic respiration, occurs in the cytoplasm. The electron transport chain occurs along the inner mitochondrial membrane. Oxidative phosphorylation occurs as protons (H+ ions) move from the intermembrane space to the mitochondrial matrix.
3. B A heterotroph obtains its energy from organic molecules. An autotroph obtains energy from sunlight utilizing pigments such as chlorophyll and uses CO2 and water to make organic molecules. Therefore, (A) and (C) can be eliminated. Heterotrophs can obtain their energy from ingesting autotrophs, but they can also consume other heterotrophs. So you can eliminate (D), leaving (B) as your answer.
4. B In meiosis, the sister chromatids separate during the second metaphase of meiosis (Meiosis II), whereas the sister chromatids separate during metaphase of mitosis. Choice (A) is incorrect because in meiosis there are two rounds of cell division, whereas in mitosis there is only one round of cell division. Chromosomes are replicated during interphase in both meiosis and mitosis, so (C) is incorrect. Choice (D) can also be eliminated because spindle fibers form during prophase in both mitosis and meiosis.
5. C Amino acids are organic molecules that contain carbon, hydrogen, oxygen, and nitrogen, so eliminate (A), (B), and (D). Don’t forget to associate amino acids with nitrogen because of the amino group (NH2).
6. B Unlike eukaryotes, prokaryotes (which include bacteria) do not contain membrane-bound organelles. Bacteria contain circular double-stranded DNA, ribosomes, and a cell wall, so (A) and (D) are incorrect. Also eliminate (C) because bacterial cell membranes are made up of a bilipid layer with proteins interspersed.
7. D Populations can be described as the evolutionary unit because changes in the genetic makeup of populations can be measured over time. Eliminate (A), as genetic changes occur only at the individual level. Only under Hardy-Weinberg equilibrium does the gene pool remain fixed over time in a population. However, this statement does not explain why the population is the evolving unit, so (B) is incorrect. Choice (C) is true but does not address the question.
8. A Positive feedback occurs when a stimulus causes an increased response. Choices (B), (C), and (D) are examples of negative feedback.
9. B In order to determine the genotype of the parents, use the ratio of the offspring given in the question and work backward. The ratio of black-haired to white-haired guinea pigs is 3:1. In order to get a white-haired offspring, each parent must have been able to contribute a b allele. However, since the offspring were both black in color, they must have each been Bb.
10. D The mean weight of the offspring in the next generation will be heavier than the mean weight of the original population because all the lighter horses in the original population died off. The normal distribution for weight will therefore shift to the heavier end (to the right of the graph). You can therefore eliminate (C) because the mean weight should increase. The mean weight of the offspring could be heavier or lighter than their parents, so you can also eliminate (A) and (B).
11. C Apoptosis (programmed cell death), hox and homeotic genes (genes that control differentiation), and inductive effects (a tissue affecting the differentiation of another tissue) play a role in cell differentiation. Operons are sets of multiple genes regulated by a single regulatory unit in bacteria.
12. A The primitive atmosphere lacked oxygen (O2). It contained methane (CH4), ammonia (NH3), and hydrogen (H2).
13. A Villi and microvilli are fingerlike projections present in the small intestine which dramatically increase the surface area available for nutrient absorption.
14. C Accessory organs do not directly encounter the food/waste. Therefore, (C) is correct since it makes enzymes.
15. D Ribosomes are the site of protein synthesis. Therefore, the correct answer should start with ribosome. So eliminate (A) and (C). The polypeptide then moves through the rough ER to the Golgi apparatus, where it is modified and packaged into a vesicle. The vesicle then floats to the plasma membrane and is secreted. Choice (D) is your answer.
16. D This statement is false because an individual with two identical alleles is said to be homozygous, not heterozygous, with respect to that gene. Alleles are different forms of the same gene found on corresponding positions of homologous chromosomes, so (A) and (B) are incorrect. More than two alleles can exist for a gene, but a person can have only two alleles for each trait.
17. C To make multiple copies of a plasmid (a small circular DNA), it should be inserted into a bacterium. A plasmid would not replicate if it were inserted into a virus, so eliminate (A). If a plasmid were treated with a restriction enzyme, it would be cut into smaller fragments. This would not give us cloned versions of the plasmid, so (B) is incorrect. If the plasmid were run on a gel (using gel electrophoresis), this would only tell us the size of the plasmid; therefore, (D) is also incorrect.
18. D The least likely explanation for why mutations are low is that mutations produce variability in a gene pool. Any gene is bound to mutate. This produces a constant input of new genetic information in a gene pool. This answer choice doesn’t give us any additional information about the rate of mutations. Some mutations are subtle and cause only a slight decrease in reproductive output, so eliminate (A). Some mutations are harmful and decrease the productive success of the individual, so (B) is incorrect. Some mutations are deleterious and lead to total reproductive failure. The zygote fails to develop. Therefore, (C) can also be eliminated.
19. B The independent variable is the condition the scientist sets up. The dependent variable is the thing that is the measured outcome of the experiment. The scientist should set different temperatures and then measure the germination rate. The outdoor temperature is interesting, but it is too variable to be a good control for the groups with a set temperature.
20. B Legume plants are able to live in nitrogen-poor soil because they obtain nitrogen from nitrogen-fixing bacteria. These plants cannot make their own proteins without nitrogen from nitrogen-fixing bacteria, so eliminate (A), (C), and (D).
21. B Every cell in an organism has the same set of genes. Differences in the timing and expression levels of these genes lead to structural and functional differences.
22. C Blood in all chambers before the lungs is deoxygenated. This means the correct answer is chambers 1, 2, 3, and 4.
23. D The left ventricle is the last chamber of the heart that the blood travels through before being pumped out. Blood should never go backwards in the heart so the dye should only be found in the left ventricle.
24. B Normal cells can become cancerous when a virus invades the cell and takes over the replicative machinery. A pilus forms between two bacteria, so (A) is wrong. Also eliminate (C) because the host’s genome is not converted to the viral genome. Choice (D) is incorrect because spores are released by fungi, not viruses.
25. C Crossing-over and synapsis occur during meiosis, which produces haploid gametes. Separation of homologous chromosomes occurs during meiosis I, while separation of sister chromatids does not occur until meiosis II.
26. D If potential mates have similar breeding seasons they will most likely mate. Use common sense to eliminate the other answer choices. If the organisms don’t meet, they won’t reproduce; eliminate (A). Also eliminate (B) and (C): if potential mates do not share the same behaviors (such as courtship rituals), they may not mate.
27. D There is no union of gametes in mitosis. Choices (A) and (C) are incorrect: asexual reproduction involves the production of two new cells with the same number of chromosomes as the parent cell. If the parent cell is diploid, then the daughter cells will be diploid. The daughter cells are identical to the parent cell, so eliminate (B).
28. D Cohesion, adhesion, and capillary action are special properties of water that are necessary for transpiration to occur. Water is a hydrophilic, polar molecule.
29. B The two species differ in their preference for water volume. Species A prefers 80 mL, and Species B prefers 40 mL. The different watering times do not seem to play a role in growth as there is no obvious pattern and the times all had similar growth. Both plants prefer pH 7.
30. A The top growth conditions for Species B are pH 7, 40 mL, and any time of day. In those conditions, the species B plants grew the tallest.
31. D The plants in the watering time experiment reached the “optimal” height for Species A, but the Species B plants seemed a bit stunted. The pH must have been 10 rather than 4 to have the plants grow to those heights. This must have been done at 80 mL, which is preferred by Species A but not Species B.
32. D Choices (A), (B), and (C) would all give interesting information about the experiment, but the only one that would make the results more statistically significant is to increase the number plants in each group. Two plants is not enough to be sure about the results of the experiment.
33. D Tropism describes the reaction of plants to a stimulus. Gravitropism specifies a reaction to gravity, and negative specifies that the reaction is away from the stimulus.
34. D The net primary productivity is what the energy producers have left for storage after their own energy needs have been met. Energy stored = (total energy produced) – (energy used in cellular respiration). Choice (A) is incorrect; it is not the energy available to producers. You can also eliminate (B) because it is not the total chemical energy in producers—that is the gross primary productivity. Get rid of (C) as well because it is not the biomass (total living material) among producers.
35. C Choice (A) and (B) are incorrect: these organisms exhibit the same behavior because they were subjected to the same environmental conditions and similar habitats. This is an example of convergent evolution. However, they are not genetically similar, so (C) is the answer. (One is an insect, and the other a bird.) They are analogous, so (D) is also incorrect. They exhibit the same function but are structurally different.
36. C A mutualistic relationship is a relationship among two organisms in which both benefit. Choice (A) describes parasitism, and (B) describes commensalism. Choice (D) is an example of altruistic behavior.
37. D Beta cells secrete insulin. Destruction of beta cells in the pancreas will halt the production of insulin. Therefore, eliminate (A), (B), and (C). This will lead to an increase in blood glucose levels.
38. C The graph does show a drought, but it says nothing about genetic variation. It is true that the rainfall could affect evolution of the population, but the graph doesn’t address evolution. It only shows the decrease in biomass. Choice (C) is the best answer since it addresses how the environment effects ripple through the levels of the ecosystem. Invasive species do not need an unoccupied niche, and that is not shown in the graph.
39. A The biomass seems to correlate fairly well with the rainfall. Therefore, a rainfall higher than any recorded would give a biomass higher than any recorded. The axis on the right shows the biomass. With a rainfall of 120, the biomass should be greater than 150.
40. A The secondary consumers’ biomass should always be less than that of the primary consumers, so the only option is (A).
41. C This is an example of mutualism. Both organisms benefit. Choice (A), parasitism, is a type of symbiotic relationship in which one organism benefits and the other is harmed. Choice (B), commensalism, is when one organism benefits and the other is unaffected. Choice (D), endosymbiosis, is the idea that some organelles originated as symbiotic prokaryotes that live inside larger cells.
42. A They both contain genetic material (DNA), a plasma membrane, and a cell wall. Unlike fungi, bacteria lack a definite nucleus. Therefore, eliminate (B). Bacteria are unicellular, whereas fungi are both unicellular and multicellular. Therefore, eliminate (C) and (D).
43. B This question tests your understanding of what stage is responsible for the synthesis of new proteins for lactose utilization. The region of bacterial DNA that controls gene expression is the lac operon. Structural genes will be transcribed to produce enzymes, which produce an mRNA involved in digesting lactose.
44. C Trypsin is an enzyme. Enzymes are proteins and organic catalysts that speed up reactions without altering them. They are not consumed in the process. Therefore, you can eliminate (A) and (B). The rate of reaction can be affected by the concentration of the substrate up to a point, so (D) can be eliminated.
45. C DNA polymerase, not RNA polymerase, is the enzyme that causes the DNA strands to elongate. DNA helicase unwinds the double helix, so (A) is true and therefore incorrect. Choice (B), which states that DNA ligase seals the discontinuous Okazaki fragments, is also true. Eliminate it. In the presence of DNA polymerase, DNA strands always grow in the 5’ to 3’ direction as complementary bases attach. Therefore, (D) is also incorrect.
46. B A voltage change from +50 to –70 is called repolarization. You can eliminate (A) because a voltage change from –70 to +50 is called depolarization. Choice (C) is incorrect; a voltage change from –70 to –90 is called hyperpolarization. Choice (D) can be eliminated as well; an action potential is a traveling depolarized wave. It refers to the whole thing, from depolarization, repolarization, hyper-polarization, and back to a resting potential.
47. C Electrons passed down along the electron transport chain from one carrier to another lose energy and provide energy for making ATP. Glucose is decomposed during glycolysis, but this process is not associated with energy given up by electrons; eliminate (A). Glucose is made during photosynthesis, so eliminate (B). NADH is an energy-rich molecule, which accepts electrons during the Krebs cycle. Therefore, (D) is incorrect as well.
48. A The oxygen released during the light reaction comes from splitting water. (Review the reaction for photosynthesis.) Therefore, water must have originally contained the radioactive oxygen. Carbon dioxide is involved in the dark reaction and produces glucose, so eliminate (B). Glucose is the final product and would not be radioactive unless carbon dioxide was the radioactive material, so (C) is incorrect. Finally, eliminate (D), because nitrogen is not part of photosynthesis.
49. C Pheromones act as sex attractants, alarm signals, or territorial markers. Auxins are plant hormones that promote growth, so eliminate (A). Hormones are chemical messengers that produce a specific effect on target cells within the same organism, so eliminate (B). Enzymes are catalysts that speed up reactions, so eliminate (D).
50. D Homologous structures are organisms with the same structure but different functions. The forelegs of an insect and the forelimbs of a dog are not structurally similar. (One is an invertebrate, and the other a vertebrate.) They do not share a common ancestor. However, both structures are used for movement. All of the other examples are vertebrates that are structurally similar.
51. B Speciation occurred in the Galapagos finches as a result of the different environments on the islands. This is an example of divergent evolution. The finches were geographically isolated. Choice (A), convergent evolution, is the evolution of similar structures in distantly related organisms. Choice (C), disruptive selection, is selection that favors both extremes at the expense of the intermediates in a population. Choice (D), stabilizing selection, is selection that favors the intermediates at the expense of the extreme phenotypes in a population.
52. D Mutations produce genetic variability. All of the other answer choices are forms of asexual reproduction.
53. C Secondary consumers feed on primary consumers. If you set up a pyramid of numbers, you’ll see that the herrings belong to the third trophic level.
54. A The biomass is the total bulk of a particular living organism. The phytoplankton population has both the largest biomass and the most energy.
55. C If the herring population decreases, this will lead to an increase in the number of crustaceans and a decrease in the phytoplankton population. Reorder the organisms according to their trophic levels and determine which populations will increase and decrease accordingly.
56. D This question tests your ability to trace the neural pathway of a motor (effector) neuron. The nerve conduction will travel from the spinal cord (where interneurons are located) to the muscle.
57. D Because the brain is destroyed, it is not associated with the movement of the leg. Choice (A) is incorrect, as reflex actions are automatic. Choices (B) and (C) can also be eliminated; both of these statements are true but are not supported by the experiment.
58. C Neurotransmitters are released from the axonal bulb of one neuron and diffuse across a synapse to activate a second neuron. The second neuron is called a postsynaptic neuron. A neurotransmitter can either excite or inhibit the postsynaptic neuron. The myelin sheath speeds up the conduction in a neuron, so (A) is wrong. Also eliminate (B) and (D), as both sodium and potassium channels open during an action potential. Neurotransmitters are not involved in actions related to the axon membrane. They do not force potassium ions to move against a concentration gradient.
59. C The DNA template strand is complementary to the mRNA strand. Using the mRNA strand, work backward to establish the sequence of the DNA strand. Don’t forget that DNA strands do not contain uracil, so eliminate (B) and (D).
60. B Use the amino acid chart to determine the sequence after uracil is deleted. The deletion of uracil creates a frameshift.
61. D The mRNA is modified before it leaves the nucleus. It becomes smaller when introns (intervening sequences) are removed. A poly(A) tail and a cap are added to the mRNA and would therefore increase the length of the mRNA, so you can eliminate (A) and (B). Choice (C) is also incorrect, as exons are the coding sequences that are kept by the mRNA.
62. B The Weddell seal stores twice as much oxygen as humans. Calculate the liters per kilograms weight for both the seal and man using the information at the bottom of the chart. The Weddell seal stores 0.058 liters/kilograms (25.9 liters/450 kilograms) compared to 0.028 liters/kilograms (1.95 liters/70 kilograms) in humans.
63. C The most plausible answer is that blood is redirected toward the central nervous system, which permits the seal to navigate for long durations. Choice (A) is incorrect; the seal does not need to increase the number of red blood cells in the nervous system. Choice (B) can also be eliminated, as the seal does not need to increase the amount of oxygen to the skeletal system. Eliminate (D) because the diversion of blood does not increase the concentration of oxygen in the lungs.
64. 425–450
If you look at the absorption spectrum, you’ll see that chlorophyll a has two peaks, one at 425 nm and one at 680 nm. Chlorophyll a maximally absorbs light at approximately 425 nm.
65.  or 0.25
or 0.25
Make a Punnett square to determine the probability that the couple has a child with blood type AB. The probability is  whether it’s the first child or the third child.
whether it’s the first child or the third child.
66. 240 24,000 lbs of elk is equivalent to 12 tons since there are 2,000 lbs per ton. With a 5 percent efficiency of transfer from their food source, that 12 tons represents 5 percent of the weight of the plants they would need to consume. 5/100 = 12/tons needed, making the answer 240.
67. 0.3 The frequency of the homozygous dominant genotype (AA) is 0.49. To find the dominant allele frequency, we can use the formula provided by the Hardy-Weinberg theory, p2 + 2pq + q2 = 1, where p represents the dominant allele and q represents the recessive allele. Because we know that p2 represents the frequency of the homozygous dominant genotype, we can find the frequency of the dominant allele (p) by taking the square root of the frequency of the genotype. The square root of 0.49 is 0.7. Note that by using the formula p + q = 1, we can also determine the frequency of the recessive allele (q). It is 0.3 (0.7 + q = 1).
68.  Draw a Punnett square for the couple to determine the probability of color blindness for the boys. Individuals 6 and 7 (XcX and XY) will produce two males: one XcY and one XY. The probability of color blindness is therefore
Draw a Punnett square for the couple to determine the probability of color blindness for the boys. Individuals 6 and 7 (XcX and XY) will produce two males: one XcY and one XY. The probability of color blindness is therefore  .
.
69. 525 Make sure you read the question carefully. You are asked to calculate the number of liters per week, not per day. The chart tells us that a coconut palm loses 75 liters a day, which would mean 525 liters a week (7 × 75 = 525).
Section II: Free Response
On the following pages you’ll find two types of aids for grading your essays: checklists and sample responses. Checklists like these are used to grade your essays on the actual test. They’re actually quite simple to use.
For each item you mentioned, give yourself the appropriate number of points (1 point, 2 points, and so on). Remember that you can only get a maximum of 10 points for each essay. As you evaluate your work, don’t be kind to yourself just because you like your own essay. If the checklist mentions an explanation of structure and function and you failed to give both, then do not give yourself a point. The testing board is very particular about this. You need to mention precisely the things they’ve listed, in the way they’ve listed them, in order to gain points.
What if something you’ve mentioned doesn’t appear on the list? Provided you know it’s valid, give yourself a point. If it was something you pulled out of your hat at the last minute, odds are it’s not directly applicable to the question. However, because you might come up with details even more specific than those contained in the checklist, it’s not unlikely that the example or structure you’ve cited is perfectly valid. If so, go ahead and give yourself the point. Remember that the testing board hires college professors and high school teachers to read your essays, so they’ll undoubtedly recognize any legitimate information you slip into the essay.
The second part of the answer key involves short paragraphs. These are not templates; the testing board does not expect you to write this way. They are simply additional tools to help illustrate the things you need to squeeze into your essay in order to rack up the points. They explain in some detail how the various parts of the checklist relate to one another and may give you an idea about how best to integrate them into your own essays come test time.
If you find that you didn’t do too well on the essay portion, be sure to carefully read Chapter 14 on the free-response questions. You can use the chapters themselves as checklists. If you find it too difficult to grade your own essays, see if your teacher or a classmate will help you out. Good luck!
Long-Form Free-Response Checklist 1
a. Role of sodium potassium pump —4 points maximum
Active transport
3 sodium out of the cell
2 potassium into the cell
potassium leak channels
negative inside the cell
−70mV
created gradients
b. Ion flow—4 points maximum
Resting stage (voltage charge is –70 millivolts)
Trigger stimulus to reach threshold
Depolarization (Na+ moves into the cell, voltage-gate channels)
Repolarization (K+ ions move out of the cell, voltage-gated channels)
Back at resting stage
c. Predicted Outcome—1 point maximum
Unable to depolarize when threhold is reached
d. Myelin—1 point maximum
Faster, because the action potential can jump along the neuron
Sample Long-Form Free Response 1
The sodium potassium pump is responsible for maintaining the resting membrane potential. It actively pumps three Na+ ions out of the cell for every two K+ ions brought into the cell. Since this is active transport, these ions are pumped against the concentration gradient. The outside of the cell has more sodium, and the inside has more potassium. The inside is more negative, since more positives leave the cell than enter the cell. The potassium can also leak out a bit by potassium leak channels.
When a nerve cell is undisturbed, the membrane is said to be in a resting stage. The membrane is polarized, and the voltage charge is –70 millivolts. During a nerve impulse, there is a change in the membrane permeability, and the voltage charge becomes slightly positive. At a certain voltage, voltage-gated sodium channels are opened. Na+ ions rush into the cell, and the inside becomes more positively charged. The membrane is now said to be depolarized. Na+ move into the cell down its concentration gradient. Next, the Na+ channels close, voltage-gated K+ channels open, and K+ ions move out of the cell. The cell is now said to be repolarized as it gets more negative. Then, the original ion concentration is reestablished by the sodium-potassium pump.
If there were not any voltage-gated sodium channels, the action potential couldn’t “fire” when threshold is reached, and the cell would not have a rapid depolarization event.
Myelin is like an insulator that covers the outside of some axons. It allows action potentials to move more quickly because the wave of positive charge can jump along the axon.
Long-Form Free-Response Checklist 2
a. Type of Mutation—3 points maximum
(1 point each)
Base substitution (involves one nucleotide being replaced by another)
Change in DNA/codon of mRNA causes the wrong tRNA to be recruited
The wrong amino acid gets added to the polypeptide
Change in polypeptide, in turn, alters hemoglobin protein
Distorted red blood cell cannot efficiently carry oxygen
b. Gel Electrophoresis—5 points maximum
How it works—3 points maximum
(1 point each)
| Apparatus | DNA is put into wells of an agarose gel with a buffer |
| Electricity | Electrical potential (electrical charge moves fragments) |
| Charge | Negatively charged fragments move toward the positive pole |
| Size/Molecular weight | Smaller fragments move faster across the gel |
How it can be applied to identify the mutant gene—2 points maximum
(1 point each)
Compare two DNA samples with normal and mutant hemoglobin genes.
Use restriction enzymes to cut the DNA into several fragments.
The normal and the mutant hemoglobin will be cut into different fragments.
Normal hemoglobin can be used as a marker.
When the two samples are run on a gel, they will separate into different banding patterns because they will have been cut differently by the enzyme.
c. Punnett Square—2 points maximum
Sickle-cell disease is an autosomal recessive disorder. Gender does not matter.
A normal female could be heterozygous or homozygous dominant.
A male with the disease would be homozygous.
Sample Long-Form Free Response 2
Sickle-cell anemia is a disease in which the red blood cells have an abnormal shape because of a base substitution. Base substitution involves an error in DNA replication in which one nucleotide is replaced by another nucleotide. This causes the codon of an mRNA to contain an incorrect base. The codon, therefore, matches up with the anticodon of a different tRNA. This tRNA carries a different amino acid. The change in the amino acid alters the polypeptide. The polypeptide, in turn, alters the hemoglobin protein. In this case, the distorted red blood cell cannot efficiently carry oxygen.
Biologists must have determined the nature of the hemoglobin mutation by comparing the normal hemoglobin gene to the abnormal hemoglobin gene, using the technique gel electrophoresis. Gel electrophoresis identifies the difference between two molecules by examining the different rates each molecule moves across the gel. Substances move across a gel according to their molecular weight. For example, smaller fragments move faster than larger fragments. The biologists must have placed fragments of the two genes on an agarose gel and used the normal hemoglobin gene as the marker—the source of comparison to the abnormal hemoglobin gene. (A restriction enzyme was used to cut the two DNA sequences into several fragments prior to loading the gel.) The two DNA sequences should have been identical except for the fragment that contained the abnormal gene.
If a normal noncarrier female mates with a male who is homozygous for the disease, these are the results using a Punnett square.
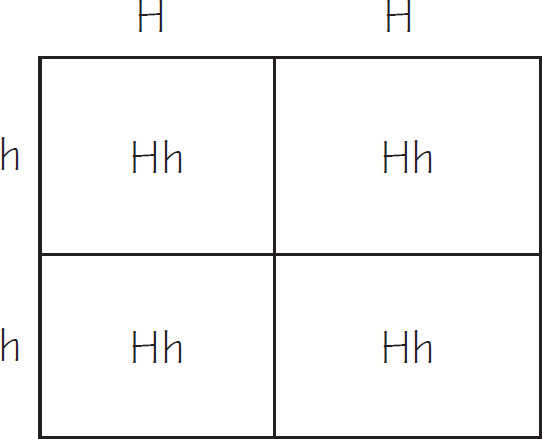
All of the offspring would be heterozygotes. For sickle-cell anemia, these offspring would be carriers.
Short-Form Free-Response Checklist 3
a. Cell size ratio—1 point maximum
(1 point each)
A higher ratio of surface area-to-volume allows for greater space for solutes to move in and out of cells.
Cells must maintain homeostasis and, in order to do this, must eliminate wastes, ingest nutrients, and maintain osmotic and ion balances.
As a cell grows larger, this ratio diminishes, limiting cell size.
b. Adaptations—4 points maximum
(1 point each)
Alveoli
Convoluted membranes in chloroplasts and mitochondria
Root hairs
Villi and microvilli
Endoplasmic reticulum
c. Transport across the membrane—1 point maximum
Small polar molecules cross by facilitated diffusion and require membrane channels.
Small nonpolar molecules can freely diffuse across the lipid bilayer.
Sample Short-Form Free Response 3
Cells need a certain surface area-to-volume ratio to exchange materials with the environment in order to maintain homeostasis. A high surface area-to-volume ratio allows cells to regulate ion concentrations as well as take in nutrients and eliminate wastes. As a cell becomes larger, the ratio of surface area to volume decreases, and if a cell grows too large, it cannot carry out these functions and therefore will not survive. Organisms have adapted a variety of ways to increase surface area. One example is alveoli, small air sacs in the lungs that maximize surface area in order to allow for maximum respiration. Another example of increasing surface area is villi in the small intestine, whose expansion results in a large surface area for the absorption of nutrients.
Depending on polarity, molecules cross the cell membrane in different ways. Small nonpolar molecules, like carbon dioxide, can freely diffuse through the cell membrane. However, due to the hydrophobic nature of the interior of the cell membrane, small polar molecules need protein channels to allow selective diffusion into or out of the cell. One example of a polar molecule is water, which crosses the membrane through channels called aquaporins.
Short-Form Free-Response Checklist 4
a. Krebs cycle and the electron transport chain, and chemiosmosis as aerobic processes—2 points maximum
They require oxygen.
They cannot occur under anaerobic conditions.
b. Site of each step—3 points maximum
| Stage | Site |
| Krebs cycle | Mitochondrial matrix |
| Electron transport chain | Along the inner mitochondrial membrane |
| Chemiosmosis | As hydrogens move from the intermembrane space to the mitochondrial matrix |
Sample Short-Form Free Response 4
a. The Krebs cycle, electron transport chain, and chemiosmosis are all part of aerobic respiration. During aerobic respiration, glucose is completely converted to CO2, ATP, and water. These steps are considered aerobic processes because they cannot occur under anaerobic conditions; they require oxygen. Glycolysis, on the other hand, can occur under both aerobic and anaerobic conditions.
b. These three stages of aerobic respiration occur in different parts of the mitochondria. The Krebs cycle occurs in the mitochondrial matrix. Two acetyl-CoA enter the Krebs cycle and produce NADH, FADH2, ATP, and CO2. The products of the Krebs cycle (NADH and FADH2) are sent to the electron transport chain. The electron transport chain occurs along the inner mitochondrial membrane. The final stage of aerobic respiration—chemiosmosis—occurs as hydrogens move across from the intermembrane space to the mitochondrial matrix.
Short-Form Free-Response Checklist 5
(1 point each)
Produces somatic cells versus produces gametes
One round of division versus two rounds of division
Meiosis only: recombination between homologous chromosomes
Sample Short-Form Free Response 5
The difference between meiosis and mitosis arises because of the different goals of these cellular processes. First, mitosis is designed to reproduce copies of somatic cells, while meiosis is designed to create gametes with greater variability. Second, in order to create the appropriate level of genetic information, meiosis requires two rounds of division, whereas recreating somatic cells only requires one round. Third, in order to increase genetic variability, recombination can occur as part of meiosis between homologous chromosomes. Such mixing is not part of mitosis since the goal is to simply recreate the original cell.
Short-Form Free-Response Checklist 6
(1 point each)
Definition of homologous structures
Relevant example (many possible)
Compare to analogous structures
Sample Short-Form Free Response 6
Homologous structures are those with similar structures because they arose from the same ancestral source. An example of homologous structures would be the arm of a human and the front leg of a cat. Their functions are different in their current form, but the ancestral origin is shared. They are different from analogous structures because they occur in species that have have a common ancestor. Analogous structures evolved separately in two lineages and don’t come from a common ancestor.
Short-Form Free-Response Checklist 7
(1 point each)
Transduction
Transformation
(2 points each)
Conjugation
Sample Short-Form Free Response 7
There are three ways in which bacteria transmit genetic information laterally to introduce new phenotypes. These are transduction, transformation, and conjugation. Transduction is the transmission of genetic material from one bacteria to another via a lysogenic virus. Transformation is the uptake of naked DNA from the environment. Conjugation is the cell-to-cell transfer of genetic material in the form of plasmids across a pilus formed between two cells.
Short-Form Free-Response Checklist 8
(1 point each)
No respiration
Not able to create own energy
Not able to reproduce independently
Not able to live outside host cells
Sample Short-Form Free Response 8
Viruses are typically not considered to be alive because they are not able to engage in many of the functions associated with life. Though viruses must engage in processes like translation that require energy, they do not engage in any process to create that energy. Along that same line, there is no respiration of any kind being done by viruses. Further, viruses have no active existence outside of their host cells; no viral functions happen outside of the context of a cell. Due to that restriction, viruses are also not able to reproduce themselves. Being entirely reliant on a host cellular environment for reproduction is another reason to consider them as not alive.
Part III
About the AP Biology Exam
• The Structure of the AP Biology Exam
• How the AP Biology Exam Is Scored
• Overview of Content Topics
• How AP Exams Are Used
• Other Resources
• Designing Your Study Plan
THE STRUCTURE OF THE AP BIOLOGY EXAM
The AP Biology Exam is three hours long and is divided into two sections: Section I (multiple-choice questions and grid-in quantitative question) and Section II (free-response questions).
Section I consists of 69 questions. These are broken down into Part A (63 multiple-choice questions) and Part B (6 grid-in questions). You will have 90 minutes to complete this section.
Section II involves free-response questions. You’ll be presented with two long-form free-response questions and six short-form free-response questions touching upon key issues in biology. You’ll be given a 10-minute reading period followed by 80 minutes to answer all eight questions.
If you’re thinking that this sounds like a heap of work to try to finish in three hours, you’re absolutely right. How can you possibly tackle so much science in so little time? Fortunately, there’s absolutely no need to. As you’ll soon see, we’re going to ask you to leave a small chunk of the test blank. Which part? The parts you don’t like. This selective approach to the test, which we call “pacing,” is probably the most important part of our overall strategy. But before we talk strategy, let’s look at the topics that are covered by the AP Biology Exam.
HOW THE AP BIOLOGY EXAM IS SCORED
AP scores are calculated from your scores on the multiple-choice and free-response sections. The final score is reported on a scale from 1 to 5. The following table explains what that final score means:
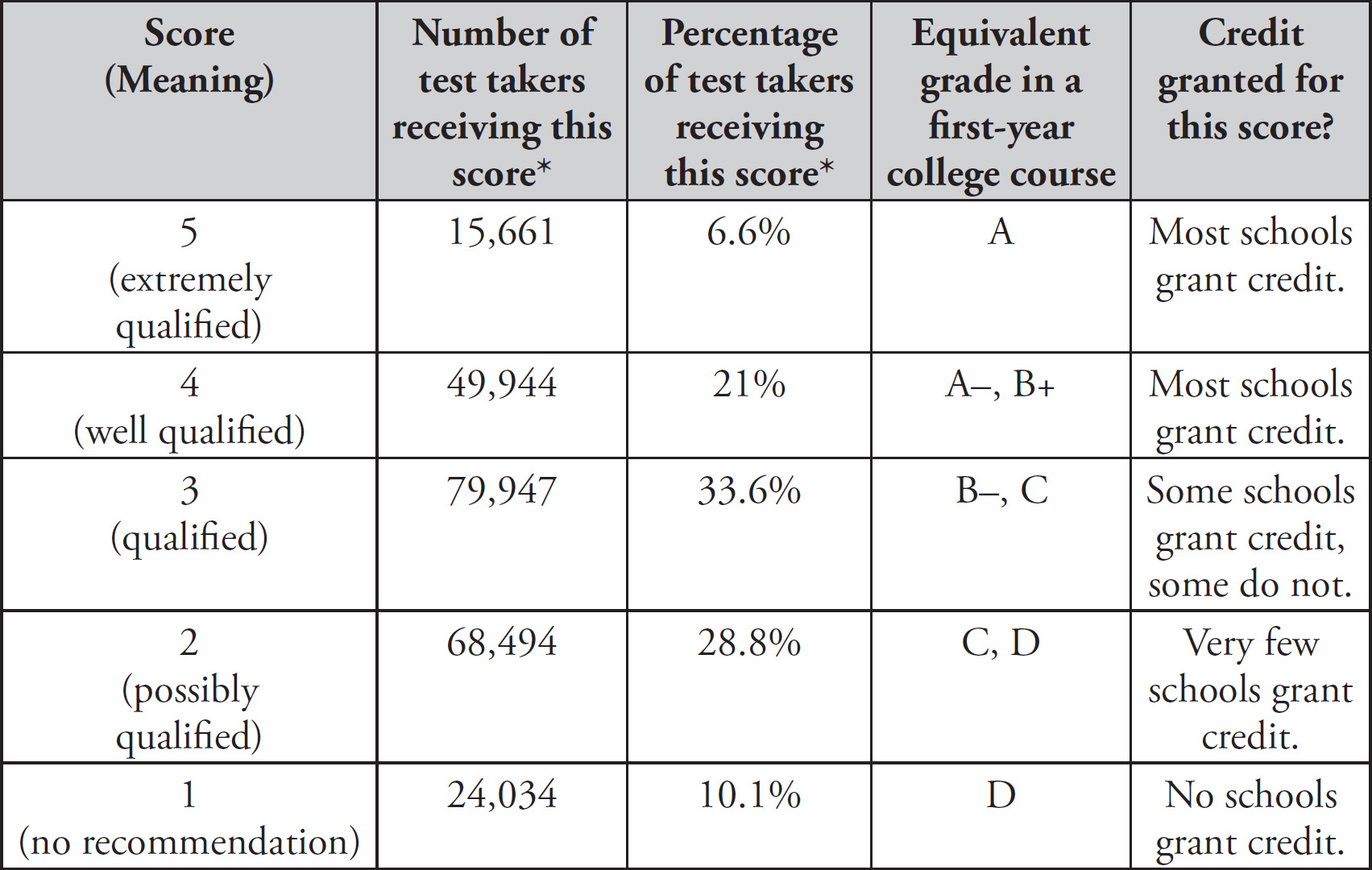
*The data above is from the College Board website and based on the May 2016 test administration.
Remember that colleges’ rules may vary when it comes to granting credit for AP courses. You should contact the individual admissions departments to find out what score you need on the exam to ensure you’ll be given credit.
For the AP Biology Exam, your scores on the multiple-choice section and free-response section are each worth 50 percent of your final score. On the multiple-choice section, your total score is based on the number of questions answered correctly, and you do not lose any points for incorrect answers. Unanswered questions do not receive points. The free-response section is graded on separate point system: long-form responses (Questions 1 and 2) are graded on a 10-point scale, while half of your short-form responses are graded on a 3-point scale and half are scored on a 4-point scale. Your scores are tallied to determine your total free-response score. The free-response score is then combined with your multiple-choice score and weighted to figure out where your score falls within the standard AP scoring scale (1 to 5).
OVERVIEW OF CONTENT TOPICS
The AP Biology Exam covers these four Big Ideas.
These are the official Big Ideas:
• Big Idea 1: The process of evolution drives the diversity and unity of life.
• Big Idea 2: Biological systems utilize free energy and molecular building blocks to grow, to reproduce, and to maintain dynamic homeostasis.
• Big Idea 3: Living systems store, retrieve, transmit, and respond to information essential to life processes.
• Big Idea 4: Biological systems interact, and these systems and their interactions possess complex properties.
These are the big ideas broken down into simple concepts. As you read this book, think about how these themes fit with various areas of Biology:
• Building blocks/hierarchy
• Responses to stimuli
• Organization
• Structure/function
• Communication
• Relationships
• Disruptions and Consequences
• Critical Thinking
To fully understand the four big ideas, a solid grasp of the following topics is required. These topics include the following:
• Chemistry of Life
 Important properties of water
Important properties of water
 pH
pH
 Carbohydrates
Carbohydrates
 Proteins
Proteins
 Lipids
Lipids
 Nucleic acids
Nucleic acids
 Origins of life
Origins of life
• Cells
 Prokaryotic and eukaryotic cells
Prokaryotic and eukaryotic cells
 Organelles
Organelles
 Membranes and transport
Membranes and transport
 Cell junctions
Cell junctions
 Cell communication
Cell communication
• Cellular Energetics
 Change in free energy
Change in free energy
 Enzymes
Enzymes
 Coupled reactions and ATP
Coupled reactions and ATP
 Photosynthesis
Photosynthesis
 Cellular respiration (glycolysis, Krebs, oxidative phosphorylation)
Cellular respiration (glycolysis, Krebs, oxidative phosphorylation)
 Fermentation
Fermentation
• Cell Reproduction
 Cell cycle
Cell cycle
 Mitosis
Mitosis
 Meiosis
Meiosis
• Molecular Biology
 DNA and genome structure
DNA and genome structure
 Transcription
Transcription
 Translation
Translation
 Gene regulation
Gene regulation
 Mutation
Mutation
 Biotechnology
Biotechnology
• Heredity
 Mendelian genetics
Mendelian genetics
 Inheritance patterns
Inheritance patterns
• Evolutionary Biology
 Natural selection
Natural selection
 Evidence of evolution
Evidence of evolution
 Phylogenetic trees
Phylogenetic trees
 Impact of genetic variation
Impact of genetic variation
 Speciation
Speciation
 Hardy-Weinberg equilibrium
Hardy-Weinberg equilibrium
• Structure and Function of Living Things
 Embryo development and body plan
Embryo development and body plan
 Immune system
Immune system
 Viruses and bacteria
Viruses and bacteria
 Nervous system
Nervous system
 Endocrine system
Endocrine system
• Ecology
 Behavior and communication
Behavior and communication
 Food webs and energy pyramids
Food webs and energy pyramids
 Succession
Succession
 Communities and ecosystems
Communities and ecosystems
 Global issues
Global issues
• Quantitative Skills
 Types of data
Types of data
 Use of mathematical formulas
Use of mathematical formulas
 Descriptive statistics
Descriptive statistics
 Graphing
Graphing
 Hypothesis testing
Hypothesis testing
This might seem like an awful lot of information, but for each topic, there are just a few key facts you’ll need to know. Your biology textbooks may go into far greater detail about some of these topics than we do. That’s because they’re trying to teach you “correct science,” whereas we’re aiming to improve your scores. Our science is perfectly sound; it’s just cut down to size. We’ve focused on crucial details and given you only what’s important. Moreover, as you’ll soon see, our treatment of these topics is far easier to handle.
The AP Biology exam not only tests your content knowledge, but it also tests how you apply that knowledge during scientific inquiry. Simply put, the test’s authors are testing if you can design and/or think critically about experiments and the hypotheses, evidence, math, data, conclusions, and theories therein. There are seven broad science practices that are tested.
• Science Practice 1: The student can use representations and models to communicate scientific phenomena and solve scientific problems.
• Science Practice 2: The student can use mathematics appropriately.
• Science Practice 3: The student can engage in scientific questioning to extend thinking or to guide investigations within the context of the AP course.
• Scientific Practice 4: The student can plan and implement data collection strategies appropriate to a particular scientific question.
• Scientific Practice 5: The student can perform data analysis and evaluation of evidence.
• Science Practice 6: The student can work with scientific explanations and theories.
• Science Practice 7: The student is able to connect and relate knowledge across various scales, concepts and representations in and across domains.
HOW AP EXAMS ARE USED
Different colleges use AP Exams in different ways, so it is important that you go to a particular college’s website to determine how it uses AP Exams. The three items below represent the main ways in which AP Exam scores can be used:
• College Credit. Some colleges will give you college credit if you score well on an AP Exam. These credits count toward your graduation requirements, meaning that you can take fewer courses while in college. Given the cost of college, this could be quite a benefit, indeed.
• Satisfy Requirements. Some colleges will allow you to “place out” of certain requirements if you do well on an AP Exam, even if they do not give you actual college credits. For example, you might not need to take an introductory-level course, or perhaps you might not need to take a class in a certain discipline at all.
• Admissions Plus. Even if your AP Exam will not result in college credit or allow you to place out of certain courses, most colleges will respect your decision to push yourself by taking an AP Course or an AP Exam outside of a course. A high score on an AP Exam shows mastery of more difficult content than is taught in many high school courses, and colleges may take that into account during the admissions process.
OTHER RESOURCES
There are many resources available to help you improve your score on the AP Biology Exam, not the least of which are your teachers. If you are taking an AP class, you may be able to get extra attention from your teacher, such as obtaining feedback on your essays. If you are not in an AP course, reach out to a teacher who teaches AP Biology and ask if the teacher will review your essays or otherwise help you with content.
Another wonderful resource is AP Central, the official site of the AP Exams. The scope of the information at this site is quite broad and includes:
• a course description, which includes details on what content is covered and sample questions
• sample questions from the AP Biology Exam
• free-response question prompts and multiple-choice questions from previous years
The AP Students home page address is: http://apstudent.collegeboard.org/home
For up-to-date information about the ongoing changes to the AP Biology Exam Course, please visit: https://apstudent.collegeboard.org/apcourse/ap-biology.
Finally, The Princeton Review offers tutoring and small group instruction. Our expert instructors can help you refine your strategic approach and add to your content knowledge. For more information, call 1-800-2REVIEW.
DESIGNING YOUR STUDY PLAN
As part of the Introduction, you identified some areas of potential improvement. Let’s now delve further into your performance on Practice Test 1 with the goal of developing a study plan appropriate to your needs and time commitment.
Read the answers and explanations associated with the multiple-choice questions (starting at this page). After you have done so, respond to the following questions:
• Review the bulleted list of topics on this page and this page. Next to each topic, indicate your rank of the topic as follows: 1 means “I need a lot of work on this,” 2 means “I need to beef up my knowledge,” and 3 means “I know this topic well.”
• How many days/weeks/months away is your exam?
• What time of day is your best, most focused study time?
• How much time per day/week/month will you devote to preparing for your exam?
• When will you do this preparation? (Be as specific as possible: Mondays and Wednesdays from 3:00 to 4:00 P.M., for example)
• Based on the answers above, will you focus on strategy (Part IV), content (Part V), or both?
• What are your overall goals in using this book?
Use those answers to create a study plan. Start with the topics that need the most work and map out when you will study and what you will study.
Remember, your schedule may evolve along the way. If a certain time/location is not working for you, then try mixing it up. If you are struggling with a topic, perhaps try tackling it with a teacher, tutor, or a classmate.
Part IV
Test-Taking Strategies for the AP Biology Exam
• Preview
1 How to Approach Multiple-Choice Questions
2 How to Approach Free-Response Questions
3 Using Time Effectively to Maximize Points
• Reflect
PREVIEW
Review your responses to the three questions on this page and then respond to the following questions:
• How many multiple-choice questions did you miss even though you knew the answer?
• On how many multiple-choice questions did you guess blindly?
• How many multiple-choice questions did you miss after eliminating some answers and guessing based on the remaining answers?
• Did you find any of the free-response questions easier/harder than the others—and, if so, why?
HOW TO USE THE CHAPTERS IN THIS PART
For the following Strategy chapters, think about what you are doing now before you read the chapters. As you read and engage in the directed practice, be sure to appreciate the ways you can change your approach. At the end of Part IV, you will have the opportunity to reflect on how you will change your approach.
Chapter 1
How to Approach Multiple-Choice Questions
SECTION I
As we mentioned earlier, the multiple-choice section consists of the following two parts:
• Part A—63 multiple-choice questions dealing with an experiment or a set of data or with general knowledge of biology
• Part B—6 grid-in questions
Part A
Part A of Section I consists of 63 run-of-the-mill multiple-choice questions. These questions test your grasp of the fundamentals of biology and your ability to apply biological concepts to help solve problems. Here’s an example:
If a coding segment of DNA reads 5′-ATG-CCA-GCT-3′, the mRNA strand that results from the transcription of this segment will be
(A) 3′-TAC-GGT-CGA-5′
(B) 3′-UCG-ACC-GUA-5′
(C) 3′-UAA-GGU-CGA-5′
(D) 3′-TAC-GGT-CTA-5′
Don’t worry about the answer to this question for now. By the end of this book, it will be a piece of cake. The majority of the questions in this section are presented in this format. A few questions may include a figure, a diagram, or a chart.
The second part of the first portion also consists of multiple-choice questions, but here you’re asked to think logically about different biological experiments or data. Here’s a typical example:
Questions 60 and 61 refer to the following diagram and information.
To understand the workings of neurons, an experiment was conducted to study the neural pathway of a reflex arc in frogs. A diagram of a reflex arc is given below.
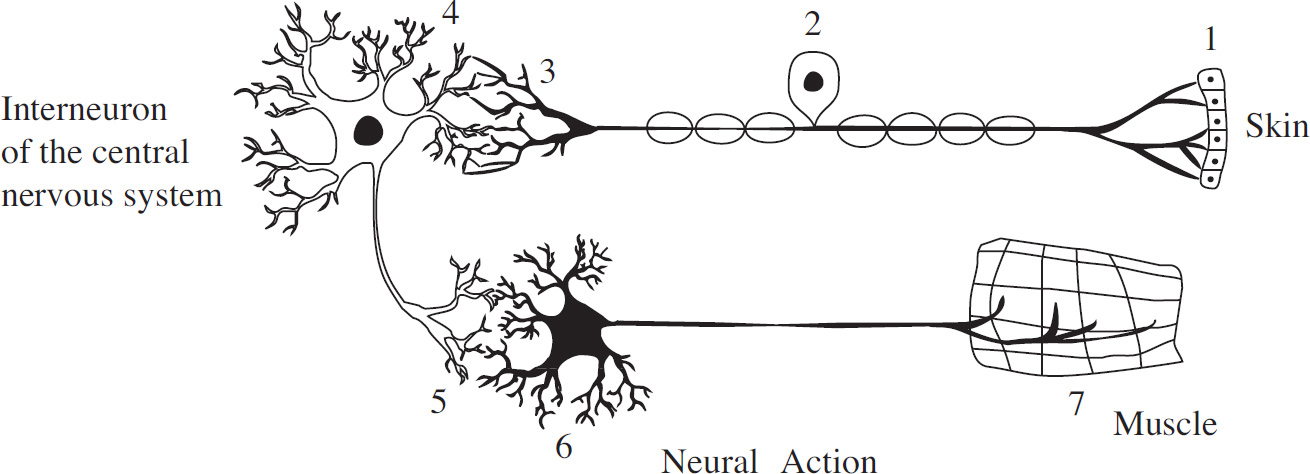
60. Which of the following represents the correct pathway taken by a nerve impulse as it travels from the spinal cord to effector cells?
(A) 1-2-3-4
(B) 6-5-4-3
(C) 2-3-4-5
(D) 4-5-6-7
61. The brain of the frog is destroyed. A piece of acid-soaked paper is applied to the frog’s skin. Every time the piece of paper is placed on its skin, one leg moves upward. Which of the following conclusions is best supported by the experiment?
(A) Reflex actions are not automatic.
(B) Some reflex actions can be inhibited.
(C) All behaviors in frogs are primarily reflex responses.
(D) This reflex action does not require the central nervous system.
You’ll notice that these particular questions refer to an experiment. Many of the questions in this portion test your ability to integrate information, interpret data, and draw conclusions from the results.
Part B
Part B of Section I consists of six grid-in questions which require using information presented in the question to calculate an answer. That numeric response is then filled in on a grid and bubbled accordingly. Answers can be in the form of integers, decimals, or fractions. A four-function calculator can be used on these questions. Here’s a typical example:
Popular Topics For Quantitative Questions
• Surface area-to-volume ratio
• Hardy-Weinberg
• Water potential
• Energy pyramids/biomass
• Chi-squared analysis
• Gene linkage
• Inheritance/probability
If the genotype frequencies of an insect population are AA = 0.49, Aa = 0.42, and aa = 0.09, what is the gene frequency of the dominant allele?

We mentioned earlier that our approach is strategy-based. As you’re about to see, many of these strategies are based on common sense—for example, using mnemonics like “ROY G. BIV.” (Remember that one? It’s the mnemonic for red, orange, yellow, green, blue, indigo, violet—the colors of the spectrum of visible light.) Others do not make as much sense. In fact, we’re going to ask you to throw out much of what you’ve been taught when it comes to taking standardized tests.
PACE YOURSELF
When you take a test in school, how many questions do you answer? Naturally, you try to answer all of them. You do this for two reasons: (1) Your teacher told you to, and (2) if you left a question blank, your teacher would mark it wrong. However, that’s not the case when it comes to the AP Biology Exam. In fact, finishing the test is the worst thing you can do. Before we explain why, let’s talk about timing.
One of the main reasons that taking the AP Biology Exam can be so stressful is the time constraint we discussed above—75 seconds per multiple-choice question, 20 minutes per long essay, and 6 minutes per short essay. If you had all day, you would probably ace the test. We can’t give you all day, but we can do the next best thing: we can give you more time for each question. How? By having you slow down and answer fewer questions.
Slowing down and doing well on the questions you do answer is the best way to improve your score on the AP Biology Exam. Rushing through questions in order to finish, on the other hand, will always hurt your score. When you rush, you’re far more likely to make careless errors, misread, and fall into traps. Keep in mind that blank answers are not counted against you.
THE THREE-PASS SYSTEM
The AP Biology Exam covers a broad range of topics. There’s no way, even with our extensive review, that you will know everything about every topic in biology. So what should you do?
Do the Easiest Questions First
The best way to rack up points is to focus on the questions you find the easiest. If you know the answer, nail it and move on. Other questions, however, will be a little more complicated. As you read each question, decide if you think it’s easy, medium, or hard. During a first pass, do all the easy questions. If you come across a problem that seems time-consuming or completely incomprehensible, skip it. Remember:
Questions that you find easy are worth just as many points as the ones you find difficult, so your time is better spent focusing on the ones in the former group.
Save the medium questions for the second pass. These questions are either time-consuming or require that you analyze all the answer choices (i.e., the correct answer doesn’t pop off the page). If you come across a question that makes no sense from the outset, save it for the last pass. You’re far less likely to fall into a trap or settle on a silly answer.
Watch Out for Those Bubbles!
Since you’re skipping problems, you need to keep careful track of the bubbles on your answer sheet. One way to accomplish this is by answering all the questions on a page and then transferring your choices to the answer sheet. If you prefer to enter them one by one, make sure you double-check the number beside the ovals before filling them in. We’d hate to see you lose points because you forgot to skip a bubble!
So then, what about the questions you don’t skip?
PROCESS OF ELIMINATION (POE)
On most tests, you need to know your material backward and forward to get the right answer. In other words, if you don’t know the answer beforehand, you probably won’t answer the question correctly. This is particularly true of fill-in-the-blank and essay questions. We’re taught to think that the only way to get a question right is by knowing the answer. However, that’s not the case on Section I of the AP Biology Exam. You can get a perfect score on this portion of the test without knowing a single right answer, provided you know all the wrong answers!
What are we talking about? This is perhaps the single most important technique in terms of the multiple-choice section of the exam. Let’s take a look at the example below.
The structures that act as the sites of gas exchange in a woody stem are the
(A) lungs
(B) lenticels
(C) ganglia
(D) lentil beans
Now if this were a fill-in-the-blank-style question, you might be in a heap of trouble. But let’s take a look at what we’ve got. You see “woody stem” in the question, which leads you to conclude that we’re talking about plants. Right away, you know the answer is not (A) or (C) because plants don’t have lungs or ganglia. Now we’ve narrowed it down to (B) and (D). Notice that (B) and (D) look very similar. Obviously, one of them is a trap. At this point, if you don’t know what “lentil beans” are, you have to guess. However, even if we don’t know precisely what they are, it’s safe to say that most of us know that lentil beans have nothing to do with plant respiration. Therefore, the correct answer is (B), lenticels.
Although our example is a little goofy and doesn’t look exactly like the questions you’ll be seeing on the test, it illustrates an important point:
Process of Elimination (POE) is the best way to approach the multiple-choice questions.
Even if you don’t know the answer right off the bat, you’ll surely know that two or three of the answer choices are not correct. What then?
Aggressive Guessing
The testing board, the service that develops and administers the exam, tells you that random guessing will not affect your score. This is true. There is no guessing penalty on the AP Biology Exam. For each correct answer you’ll receive one point and you will not lose any points for each incorrect answer.
Although you won’t lose any points for wrong answers, you should guess aggressively by getting rid of the incorrect answer choices. The moment you’ve eliminated a couple of answer choices, your odds of getting the question right, even if you guess, are far greater. If you can eliminate as many as two answer choices, your odds improve enough that it’s in your best interest to guess.
WORD ASSOCIATIONS
Another way to rack up the points on the AP Biology Exam is by using word associations in tandem with your POE skills. Make sure that you memorize the words in the Key Terms lists throughout this book. Know them backward and forward. As you learn them, make sure you group them by association, since the testing board is bound to ask about them on the AP Biology Exam. What do we mean by “word associations?” Let’s take the example of mitosis and meiosis.
You’ll soon see from our review that there are several terms associated with mitosis and meiosis. Synapsis, crossing-over, and tetrads, for example, are words associated with meiosis but not mitosis. We’ll explain what these words mean later in this book. For now, just take a look.
Which of the following typifies cytokinesis during mitosis?
(A) Crossing-over
(B) Formation of tetrads
(C) Synapsis
(D) Division of the cytoplasm
This might seem like a difficult problem. But let’s think about the associations we just discussed. The question asks us about mitosis. However, (A), (B), and (C) all mention events that we’ve associated with meiosis. Therefore, they are out. Without even racking your brain, you’ve managed to find the correct answer: (D). Not bad!
Once again, don’t worry about the science for now. We’ll review it later. What is important to recognize is that by combining the associations we’ll offer throughout this book and your aggressive POE techniques, you’ll be able to rack up points on problems that might have seemed difficult at first.
MNEMONICS—OR THE BIOLOGY NAME GAME
One of the big keys to simplifying biology is the organization of terms into a handful of easily remembered packages. The best way to accomplish this is by using mnemonics. Biology is all about names: the names of chemical structures, processes, theories, and so on. How are you going to keep them all straight? A mnemonic, as you may already know, is a convenient device for remembering something.
For example, one important issue in biology is taxonomy, that is, the classification of life-forms, or organisms. Organisms are classified in a descending system of similarity, leading from domains (the broadest level) to species (the most specific level). The complete order runs: domain, kingdom, phylum, class, order, family, genus, and species. Don’t freak out yet. Look how easy it becomes with a mnemonic:
King Philip of Germany decided to walk to America. What do you think happened?
| Dumb | → | Domain |
| King | → | Kingdom |
| Philip | → | Phylum |
| Came | → | Class |
| Over | → | Order |
| From | → | Family |
| Germany | → | Genus |
| Soaked | → | Species |
Learn the mnemonic and you’ll never forget the science!
Mnemonics can be as goofy as you like, as long as they help you remember. Throughout this book, we’ll give you mnemonics for many of the complicated terms we’ll be seeing. Use ours, if you like them, or feel free to invent your own. Be creative! Remember: the important thing is that you remember the information, not how you remember it.
IDENTIFYING EXCEPT QUESTIONS
About 10 percent of the multiple-choice questions in Section I are EXCEPT/NOT/LEAST questions. With this type of question, you must remember that you’re looking for the wrong (or the least correct) answer. The best way to go about these is by using POE.
More often than not, the correct answer is a true statement, but is wrong in the context of the question. However, the other three tend to be pretty straightforward. Cross off the three that apply, and you’re left with the one that does not. Here’s a sample question.
All of the following are true statements about gametes EXCEPT
(A) they are haploid cells
(B) they are produced only in the reproductive structures
(C) they bring about genetic variation among offspring
(D) they develop from polar bodies
If you don’t remember anything about gametes and gametogenesis, or the production of gametes, this might be a particularly difficult problem. We’ll see these again later on, but for now, remember that gametes are the sex cells of sexually reproducing organisms. As such, we know that they are haploid and are produced in the sexual organs. We also know that they come together to create offspring.
From this very basic review, we know immediately that (A) and (B) are not our answers. Both of these are accurate statements, so we eliminate them. That leaves us with (C) and (D). If you have no idea what (D) means, focus on (C). In sexual reproduction, each parent contributes one gamete, or half the genetic complement of the offspring. This definitely helps vary the genetic makeup of the offspring. Choice (C) is a true statement, so it can be eliminated. The correct answer is (D).
Don’t sweat it if you don’t recall the biology. We’ll be reviewing it in detail soon enough. For now, remember that the best way to answer these types of questions is to spot all the right statements and cross them off. You’ll wind up with the wrong statement, which happens to be the correct answer.
SOLVING GRID-IN QUESTIONS
Part B of the multiple-choice section requires you to answer six grid-in questions. Unlike the questions in Part A, you will not be able to use Process of Elimination. However, there are strategies you can use to increase your chances of efficiently getting these questions correct. First, identify what the question is actually asking for. Are you being asked to find a value on a graph or chart? Perhaps they are having you calculate the frequency of an allele using Hardy-Weinberg equilibrium. If you have no idea how to find out what they are asking you, take a quick educated guess and move on. For instance, if they are asking for a frequency, try guessing a decimal between 0 and 1, such as 0.25. However, if you do know how to find the answer, then write down the question number and any work (notes, calculations, etc.) that you would need to answer the question neatly. Before filling in your answer, make sure that you haven’t made any mistakes on interpreting data or making calculations. Be sure not to forget to bubble in your answers.
Although these grid-in questions require calculations, fear not! The College Board provides you with a handy formula sheet, which you can find at the end of this book. You can expect to be given this sheet as part of the exam, so you do not need to memorize these equations. However, it will be important for you to practice using them and familiarizing yourself with the formula sheet prior to the exam. Keep this page handy as you review and practice problems so that you will be accustomed to using it on the day of the exam.
Chapter 2
How to Approach Free-Response Questions
THE ART OF THE FREE-RESPONSE ESSAY
On Section II of the exam, you are given two long-form free-response questions and six short-form free-response questions to answer in 80 minutes. You will also have a 10-minute reading period, which means you have a total of 90 minutes to complete Section II. The best way to rack up points on this section is to give the readers what they’re looking for. Fortunately, we know precisely what that is.
The readers, or graders, have a checklist of key terms and concepts that they use to assign points. We like to call these “hot button” terms. Simply put, for each hot button term that you include in a response, you will receive a predetermined number of points. For example, if the question deals with the function of enzymes, the graders are instructed to give two points for a mention of the “lock-and-key theory of enzyme specificity.”
Naturally, you can’t just compose a “laundry list” of scientific terms. Otherwise, it wouldn’t be an essay. What you can do, however, is organize your free-response essays around a handful of these key points. The most effective and efficient way to do this is by using the 10-minute reading period to brainstorm and come up with the scientific terms. Then outline your response before you begin to write, using your hot buttons as your guide.
READ THE QUESTIONS CAREFULLY
The testing board gives you 10 minutes to read the questions and organize your thoughts before you begin writing. The first thing you should do is take less than a minute to skim all of the questions and put them into your own personal order of difficulty from easiest to toughest. Once you’ve decided the order in which you will answer the questions (easiest first, hardest last), you can begin to formulate your responses. Your first step should be a more detailed assessment of each question.
The most important advice we can give you is to read each question at least twice. As you read the question, focus on key words, especially “direction words.” Almost every essay question begins with a direction word. These words are often in bold, so they are easy to spot. Some examples of direction words are discuss, define, explain, describe, compare, and contrast. If a question asks you to discuss a particular topic, you should give a viewpoint and support it with examples. If a question asks you to compare two things, you should discuss how the two things are similar. On the other hand, if the question asks you to contrast two things, you need to show how these things are different.
Many students lose points on their essays because they either misread the question or fail to do what’s asked of them.
BRAINSTORM
Your next objective during the 10-minute reading period should be to organize your thoughts.
Once you’ve read the questions, you need to brainstorm. Jot down as many key terms and concepts as you can. Don’t forget, the test readers assign points on the basis of these key concepts. For each one that you mention and/or explain, you get a point.
How many do you need? You won’t need all of them, that’s for sure. However, you will need enough to get you the maximum number of points for that question. Most of these can be pulled directly from your reading in this book. At the end of each chapter is a Key Terms list which is full of terms that you should incorporate into your free responses. In addition to this list, you can feel free to create your own thorough lists of terms for your own use.
Don’t spend more than about 2 minutes per question brainstorming. Once your 10-minute reading period is up, you should be ready to start writing your essays.
OUTLINE YOUR RESPONSE
Have you ever written yourself into a corner? You’re halfway through your essay when you suddenly realize that you have no idea whatsoever where you’re going with your train of thought. To avoid this (and the panic that accompanies it), take a few minutes to draft an outline.
Your outline should incorporate as many of the hot buttons as you need in order to maximize your score. In other words, if a question asks for two examples, choose only those two with which you are most comfortable. In your outline, make notes about the crucial points to mention with regard to each topic or key word. Once your outline is complete, you’re ready to move on to writing the actual essay.
All of this preparation may seem time-consuming. However, it should take no more than two minutes per long question and one minute per short question. What’s more, it will greatly simplify the whole essay-writing process. So while you lose a little time at the outset, you’ll more than make up for it when it comes time to actually write your answer.
DEVELOP YOUR IDEAS IN EACH RESPONSE
Now you can use your outline to write your essay. Most students can come up with key terms or phrases that concern a particular biology topic. What separates a high-scoring student from a low-scoring student is how the student develops his or her thoughts in each essay. Besides giving the hot buttons, you’ll need to elaborate on your thoughts and ideas. For example, don’t just throw out a list of terms that pertain to meiosis and mitosis (such as synapsis, crossing-over, and gametes). Go one step further. Make sure you mention the significance of meiosis (it produces genetic variability). This extra piece of information will earn you an extra point.
Generally, you’ll need to write about two to four paragraphs, depending on the number of parts contained in each essay question. In addition, be sure to give the appropriate number of examples for each essay. If the question asks for three examples, give only three examples. If you present more than is required, the test readers won’t even read them or count them toward your score. The bottom line is this: stick to the question.
ANSWER EACH PART OF THE QUESTION SEPARATELY
The more parts there are to a free-response question, the more important it is to pace yourself. For each question, you’re better off writing a little bit for each part than you are spending all your time on any one part of a question. Why? Even if you were to write the perfect answer to one part of a question, there’s a limit to the number of points the test readers can assign to that part.
As you address each part of the question, make sure you break your response into paragraphs to make the test reader’s job easier. When test readers have an easy time reading your essay, they’re more likely to award points: your writing comes across as clearer and more organized. Readers have also requested that students label each part of their essay answers, so write “Part (a),” “Part (b),” and so on, accordingly in your response.
Finally, don’t spend too much time writing a fancy introduction. You won’t get brownie points for beautifully written openings like, “It was the best of experiments, it was the worst of experiments.” Just leap right into the essay. And don’t worry too much about grammar or spelling errors. Your grammar can hurt only if it’s so bad that it seriously impairs your ability to communicate.
INCORPORATE ELEMENTS OF AN EXPERIMENTAL DESIGN
Since one of the free-response questions will be experimentally based, you’ll need to know how to design an experiment. Most of these questions require that you present an appropriately labeled diagram or graph. Otherwise, you’ll only get partial credit for your work.
There are two things you must remember when designing experiments on the AP Biology Exam: (1) always label your figures, and (2) include controls in all experiments. Let’s take a closer look at these two points.
Know How to Label Diagrams and Figures
Let’s briefly discuss the important elements in setting up a graph. The favorite type of graph on the AP Biology Exam is the coordinate graph. The coordinate graph has a horizontal axis (x-axis) and a vertical axis (y-axis).
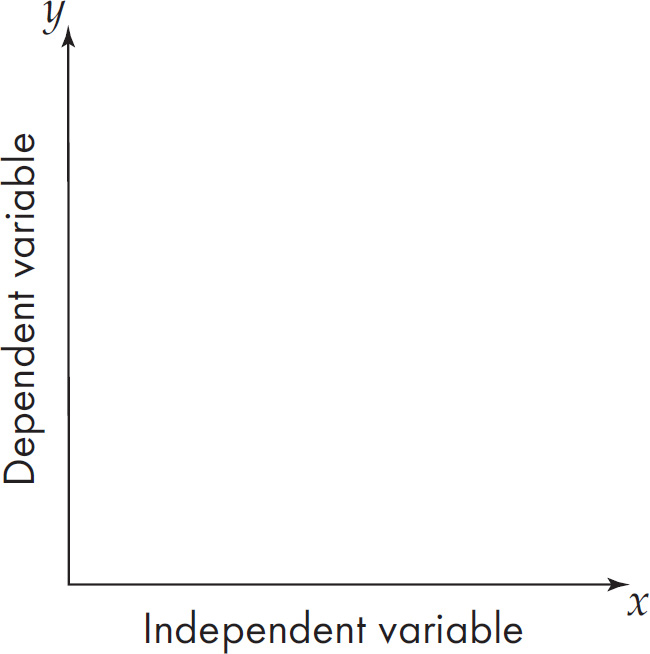
The x-axis usually contains the independent variable—the thing that’s being manipulated or changed. The y-axis contains the dependent variable—the thing that’s affected when the independent variable is changed.
Now let’s look at what happens when you put some points on the graph. Every point on the graph represents both an independent variable and a dependent variable.
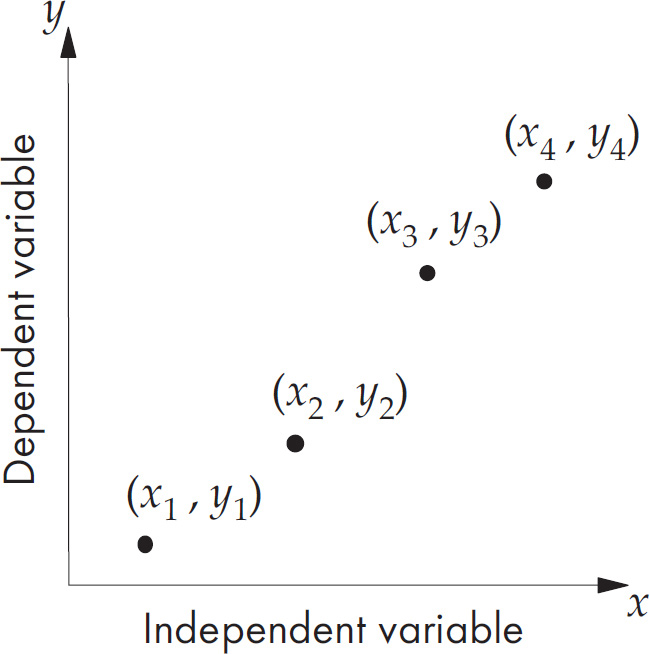
Once you draw both axes and label the axes as x and y, you can plot the points on the graph. Let’s look at the following question.
Enzymes are biological catalysts.
(a) Relate the chemical structure of an enzyme to its catalytic activity and specificity.
(b) Design an experiment that investigates the influence of temperature, substrate concentration, or pH on the activity of an enzyme.
(c) Describe what information concerning enzyme structure could be inferred from the experiment you have designed.
For now, let’s discuss only (b), which asks us to design an experiment. Let’s set up a graph that shows the results of an experiment examining the relationship between pH and enzyme activity. Notice that we’ve chosen only one factor here, pH. We could have chosen any of the three. Why did we choose pH and not temperature or substrate concentration? Well, perhaps it’s the one we know the most about.
What is the independent variable? It is pH. In other words, pH is being manipulated in the experiment. We’ll therefore label the x-axis with pH values from 0 through 14.
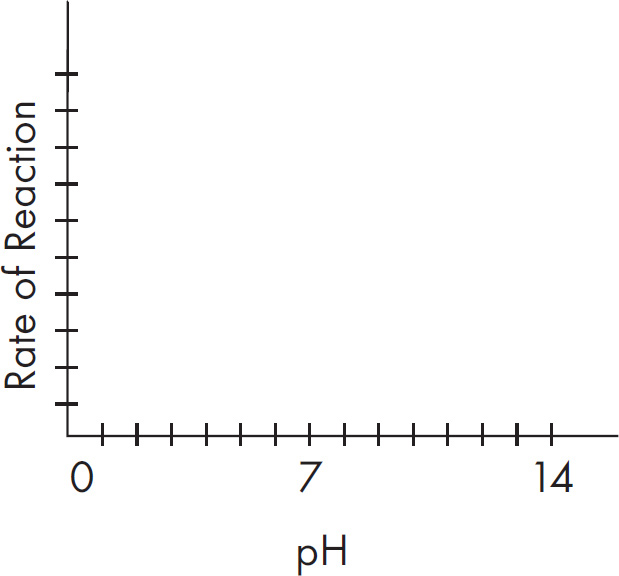
What is the dependent variable? It’s the enzyme activity—the thing that’s affected by pH. Let’s label the y-axis “Rate of Reaction.” Now we’re ready to plot the values on the graph. Based on our knowledge of enzymes, we know that for most enzymes the functional range of pH is narrow, with optimal performance occurring at or around a pH of 7.
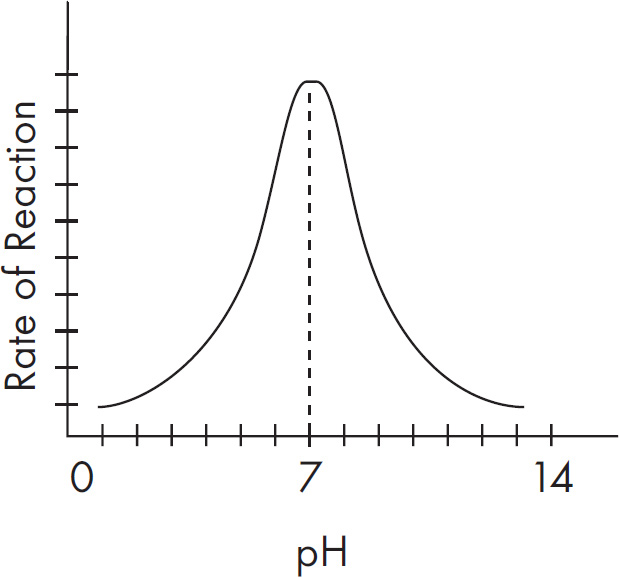
Now you should interpret your graph. If the pH level decreases from a neutral pH of 7, the reaction rate of the enzyme will decrease. If the pH level increases, the rate of reaction will also decrease. Don’t forget to include a simple explanation of your graph.
Additional quantitative and graphing skills will be discussed at length in a later chapter of this text. There you will learn how to include statistics and hypothesis testing in your free-response questions and how best to present data using graphs.
INCLUDE CONTROLS IN YOUR EXPERIMENTS
Almost every experiment will have at least one group that represents the normal/unchanged/neutral version of the independent variable. This is called the control. A control is simply a standard of comparison. What does a control do? It enables the biologist to verify that the outcome of the study is due to changes in the independent variable and nothing else.
Let’s say the principal of your school thinks that students who eat breakfast do better on the AP Biology Exam than those who don’t eat breakfast. He gives a group of 10 students from your class free breakfast every day for a year. When the school year is over, he administers the AP Biology Exam and they all score brilliantly! Did they do well because they ate breakfast every day? We don’t know for sure. Maybe those kids were the smartest kids in the class and would have scored well anyway.
In this case, the best way to be sure that eating breakfast made a difference is to have a control group. In other words, he would need to also follow for a year a group of students of equal intelligence to the first group, BUT who are known to never eat breakfast. At the end of that year, he could have them take the AP Biology Exam. If they do just as well as the group of similar intelligence that ate breakfast, then we can probably conclude that eating breakfast wasn’t necessarily leading to higher AP scores. The group of students that didn’t eat breakfast is called the control group because those students were not “exposed” to the variable of interest—in this case, breakfast. Control groups are generally any group that you include just for the sake of comparison.
Chapter 3
Using Time Effectively to Maximize Points
BECOMING A BETTER TEST TAKER
Very few students stop to think about how to improve their test-taking skills. Most assume that if they study hard, they will test well and that if they do not study, they will do poorly. Most students continue to believe this even after experience teaches them otherwise. Have you ever studied really hard for an exam and then blown it on test day? Have you ever aced an exam for which you thought you weren’t well prepared? Most students have had one, if not both, of these experiences. The lesson should be clear: factors other than your level of preparation influence your final test score. This chapter will provide you with some insights that will help you perform better on the AP Biology Exam and on other exams as well.
PACING AND TIMING
A big part of scoring well on an exam is working at a consistent pace. The worst mistake made by inexperienced or less savvy test takers is that they come to a question that stumps them and rather than just skip it, they panic and stall. Time seems to stand still when you’re working on a question you cannot answer, and it is not unusual for students to waste five minutes on a single question (especially a question involving a graph or the word EXCEPT) because they are too stubborn to cut their losses. It is important to be aware of how much time you have spent on a given question and section. There are several ways to improve your pacing and timing for the test:
• Know your average pace. While you prepare for your test, try to gauge how long you take on five, ten, or twenty questions. Knowing how long you spend on average per question will help you identify how many questions you can answer effectively and how best to pace yourself for the test.
• Have a watch or clock nearby. You are permitted to have a watch or clock nearby to help you keep track of time. However, it’s important to remember that constantly checking the clock is in itself a waste of time and can be distracting. Devise a plan. Try checking the clock every fifteen or thirty questions to see if you are keeping the correct pace or whether you need to speed up. This will ensure that you’re cognizant of the time but will not permit you to fall into the trap of dwelling on it.
• Know when to move on. Since all questions are scored equally, investing appreciable amounts of time on a single question is inefficient and can potentially deprive you of the chance to answer easier ones later on. You should eliminate answer choices if you are able to, but don’t feel bad about picking a random answer and moving on if you cannot find the correct answer. Remember, tests are like marathons; you do best when you work through them at a steady pace. You can always come back to a question you don’t know. When you do, very often you will find that your previous mental block is gone and you will wonder why the question perplexed you the first time around (as you gleefully move on to the next question). Even if you still don’t know the answer, you will not have wasted valuable time you could have spent on easier questions.
• Be selective. You don’t have to do any of the questions in a given section in order. If you are stumped by an essay or multiple-choice question, skip it or choose a different one. In the next section, you will see that you may not have to answer every question correctly to achieve your desired score. Select the questions or essays that you can answer and work on them first. This will make you more efficient and give you the greatest chance of getting the most questions correct.
• Use Process of Elimination (POE) on multiple-choice questions. Many times, one or more answer choices can be eliminated. Every answer choice that can be eliminated increases the odds that you will answer the question correctly.
Remember, when all the questions on a test are of equal value, no one question is that important, and your overall goal for pacing is to get the most questions correct. Finally, you should set a realistic goal for your final score. In the next section, we will break down how to achieve your desired score and how to pace yourself to do so.
GETTING THE SCORE YOU WANT
Depending on the score you need, it may be in your best interest not to try to work through every question. Check with the schools to which you are applying to find out what scores they will accept for credit.
AP Exams in all subjects no longer include a “guessing penalty” of a quarter of a point for every incorrect answer. Instead, students are assessed only on the total number of correct answers. A lot of AP materials, even those you receive in your AP class, may not include this information. It’s really important to remember that if you are running out of time, you should fill in all the bubbles before the time for the multiple-choice section is up. Even if you don’t plan to spend a lot of time on every question or even if you have no idea what the correct answer is, you need to fill something in.
TEST ANXIETY
Everybody experiences anxiety before and during an exam. To a certain extent, test anxiety can be helpful. Some people find that they perform more quickly and efficiently under stress. If you’ve ever pulled an all-nighter to write a paper and ended up doing good work, you know the feeling.
However, too much stress is definitely a bad thing. Hyperventilating during the test, for example, almost always leads to a lower score. If you find that you stress out during exams, here are a few preemptive actions you can take:
• Take a reality check. Evaluate your situation before the test begins. If you have studied hard, remind yourself that you are well prepared. Remember that many others taking the test are not as well prepared, and (in your classes, at least) you are being graded against them, so you have an advantage. If you didn’t study, accept the fact that you will probably not ace the test. Make sure you get to every question you know something about. Don’t stress out or fixate on how much you don’t know. Your job is to score as high as you can by maximizing the benefits of what you do know. In either scenario, it’s best to think of a test as if it were a game. How can you get the most points in the time allotted to you? Always answer questions you can answer easily and quickly before tackling those that will take more time.
• Try to relax. Slow, deep breathing works for almost everyone. Close your eyes, take a few slow, deep breaths, and concentrate on nothing but your inhalation and exhalation for a few seconds. This is a basic form of meditation that should help you to clear your mind of stress and, as a result, concentrate better on the test. If you have ever taken yoga classes, you probably know some other good relaxation techniques. Use them when you can (obviously, anything that requires leaving your seat and, say, assuming a handstand position won’t be allowed by any but the most free-spirited proctors).
• Eliminate as many surprises as you can. Make sure you know where the test will be given, when it starts, what type of questions are going to be asked, and how long the test will take. You don’t want to be worrying about any of these things on test day or, even worse, after the test has already begun.
The best way to avoid stress is to study both the test material and the test itself. Congratulations! By reading this book, you are taking a major step toward a stress-free AP Biology Exam.
REFLECT
Respond to the following questions:
• How long will you spend on multiple-choice questions?
• How will you change your approach to multiple-choice questions?
• What is your multiple-choice guessing strategy?
• How much time will you spend on the free-response questions?
• What will you do before you begin writing your free-response answers?
• Will you seek further help outside of this book (such as a teacher, tutor, or AP Students) on how to approach the questions that you will see on the AP Biology Exam?
Part V
Content Review for the AP Biology Exam
4 Chemistry of Life
5 Cells
6 Cellular Energetics
7 Molecular Biology
8 Cell Reproduction
9 Heredity
10 Evolutionary Biology
11 Animal Structure and Function
12 Behavior and Ecology
13 Quantitative Skills and Biostatistics
14 Sample Free-Response Questions
15 Laboratory
16 Chapter Drill Answers and Explanations
Chapter 4
Chemistry of Life
ELEMENTS
Although organisms exist in many diverse forms, they all have one thing in common: they are all made up of matter. Matter is made up of elements. Elements, by definition, are substances that cannot be broken down into simpler substances by chemical means.
THE ESSENTIAL ELEMENTS OF LIFE
Although there are 92 natural elements, 96% of the mass of all organisms is made up of just four of them: oxygen (O), carbon (C), hydrogen (H), and nitrogen (N). Other elements such as calcium (Ca), phosphorus (P), potassium (K), sulfur (S), sodium (Na), Chlorine (Cl), and magnesium (Mg) are also present, but in smaller quantities. These elements make up most of the remaining four percent of an organism’s weight. Some elements are known as trace elements because they are only required by an organism in very small quantities. Trace elements include iron (Fe), iodine (I), and copper (Cu).
SUBATOMIC PARTICLES
If you break down an element into smaller pieces, you’ll eventually come to the atom—the smallest unit of an element that retains its characteristic properties. Atoms are the building blocks of the physical world.
Within atoms, there are even smaller subatomic particles called protons, neutrons, and electrons. Let’s take a look at a typical atom.
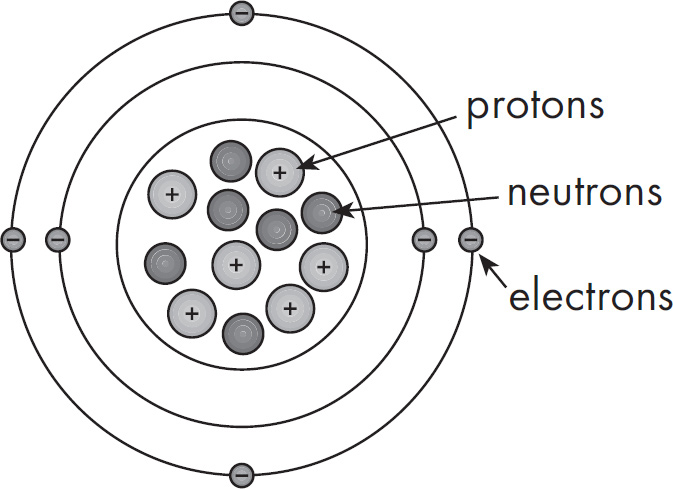
Protons and neutrons are particles that are packed together in the core of an atom called the nucleus. You’ll notice that protons are positively charged (+) particles, whereas neutrons are uncharged particles.
Electrons, on the other hand, are negatively charged (–) particles that spin around the nucleus. Electrons are pretty small compared to protons and neutrons. In fact, for our purposes, electrons are considered massless. Most atoms have the same number of protons and electrons, making them electrically neutral. Some atoms have the same number of protons but differ in the number of neutrons in the nucleus. These atoms are called isotopes.
COMPOUNDS
When two or more different types of atoms are combined in a fixed ratio, they form a chemical compound. You’ll sometimes find that a compound has different properties from those of its elements. For instance, hydrogen and oxygen exist in nature as gases. Yet when they combine to make water, they often pass into a liquid state. When hydrogen atoms get together with oxygen atoms to form water, we’ve got a chemical reaction.
2H2 (g) + O2(g) → 2H2O (l)
The atoms of a compound are held together by chemical bonds, which may be ionic bonds, covalent bonds, or hydrogen bonds.
An ionic bond is formed between two atoms when one or more electrons are transferred from one atom to the other. In this reaction, one atom loses electrons and becomes positively charged, and the other atom gains electrons and becomes negatively charged. The charged forms of the atoms are called ions. An ionic bond results from the attraction between two oppositely charged ions. For example, when Na reacts with Cl, the charged ions Na+ and Cl– are formed.
A covalent bond is formed when electrons are shared between atoms. If the electrons are shared equally between the atoms, the bond is called nonpolar covalent. If the electrons are shared unequally, the bond is called polar covalent. When one pair of electrons is shared between two atoms, the result is a single covalent bond. When two pairs of electrons are shared, the result is a double covalent bond. When three pairs of electrons are shared, the result is a triple covalent bond.
WATER: THE VERSATILE MOLECULE
One of the most important substances in nature is water. Water is considered a unique molecule because it plays an important role in chemical reactions.
Let’s take a look at one of the properties of water. Water has two hydrogen atoms joined to an oxygen atom.
In water molecules, the hydrogen atoms have a partial positive charge and the oxygen atom has a partial negative charge. Molecules that have partially positive and partially negative charges are said to be polar. Water is therefore a polar molecule. The positively-charged ends of the water molecules strongly attract the negatively-charged ends of other polar compounds (including water). Likewise, the negatively-charged ends strongly attract the positively-charged ends of polar compounds. These forces are most readily apparent in the tendency of water molecules to stick together, as in the formation of water beads or raindrops.
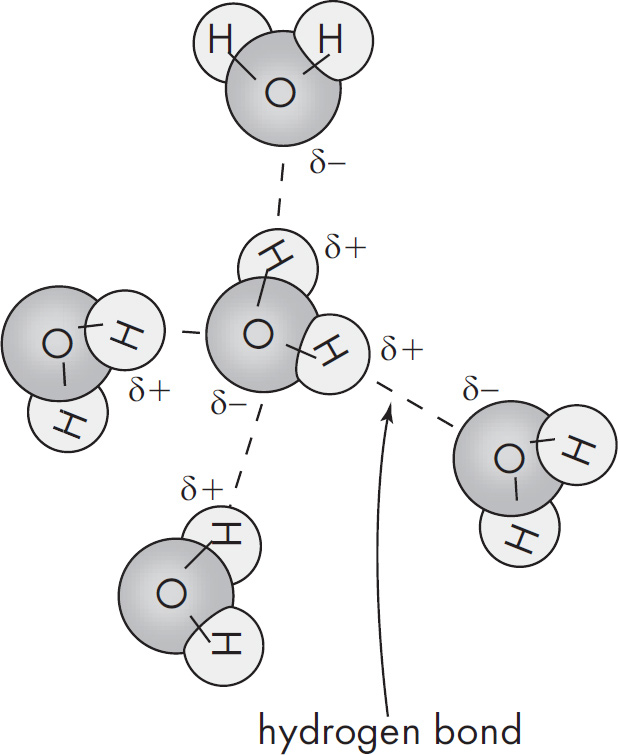
These types of intermolecular attractions are called hydrogen bonds. Hydrogen bonds are weak chemical bonds that form when a hydrogen atom that is covalently bonded to one electronegative atom is also attracted to another electronegative atom. Water molecules are held together by hydrogen bonds.
Although hydrogen bonds are individually weak, they are strong when present in large numbers. Because water reacts well with other polar substances, it makes a great solvent; it can dissolve many kinds of substances. The hydrogen bonds that hold water molecules together contribute to a number of special properties, including cohesion, adhesion, heat capacity, and expansion on freezing.
Cohesion and Adhesion
As mentioned above, water molecules have a strong tendency to stick together. That is, water exhibits cohesive forces. These forces are extremely important to life. For instance, during transpiration water molecules evaporate from a leaf, “pulling” on neighboring water molecules. These, in turn, draw up the molecules immediately behind them, and so on, all the way down the plant vessels. The resulting chain of water molecules enables water to move up the stem. Water also has a high surface tension because of the cohesiveness of its molecules.
The strong cohesion of water molecules leads to high surface tension, which is the tension on the surface of water from water molecules huddling together tightly to minimize contact with the air.
Water molecules also like to stick to other substances—that is, they’re adhesive. Have you ever tried to separate two glass slides stuck together by a film of water? They’re difficult to separate because of the water sticking to the glass surfaces.
These two forces taken together—cohesion and adhesion—account for the ability of water to rise up the roots, trunks, and branches of trees. Since this phenomenon occurs in thin vessels, it’s called capillary action.
Heat Capacity
Another remarkable property of water is its high heat capacity. What’s heat capacity? Your textbook will give you a definition something like this: “heat capacity is the quantity of heat required to change the temperature of a substance by 1 degree.” What does that mean? In plain English, heat capacity refers to the ability of a substance to resist temperature changes. For example, when you heat up an iron kettle, it gets hot pretty quickly. Why? Because it has a low specific heat. It doesn’t take much heat to increase the temperature of the kettle. Water, on the other hand, has a high heat capacity. You have to add a lot of heat to get an increase in temperature. Water’s ability to resist temperature changes is one of the things that helps keep the temperature in our oceans fairly stable. It’s also why organisms that are mainly made up of water, like us, are able to keep a constant body temperature.
Expansion on Freezing
Perhaps one of the most important properties of water is yet another result of hydrogen bonding. When four water molecules are bound in a solid lattice of ice, the hydrogen bonds actually cause solid water to expand on freezing. In most materials, when the molecules lose kinetic energy and cool from liquid to solid, the molecules get more dense, moving closer together. However, in liquid water, the molecules are slightly more dense than in solid water. Because water expands on freezing, becoming slightly less dense than liquid water, ice can crack lead pipes in the winter, or a soda can will pop if it is left in your cold car. The important consequence of this property of water is that ice floats on the top of lakes or streams, allowing animal life to live underneath the ice. If ice was denser than water, it would sink to the bottom, freezing the body of water solid. All aquatic life would be unable to survive.
So let’s review the unique properties of water:
• Water is polar and can dissolve other polar substances.
• Water has cohesive and adhesive properties.
• Water has a high heat capacity.
• Water expands when it freezes (or, ice floats).
THE ACID TEST
We just said that water is important because most reactions occur in watery solutions. Well, there’s one more thing to remember: reactions are also influenced by whether the solution in which they occur is acidic, basic, or neutral.
What makes a solution acidic or basic? A solution is acidic if it contains a lot of hydrogen ions (H+). That is, if you dissolve an acid in water, it will release a lot of hydrogen ions. When you think about acids, you usually think of substances with a sour taste, like lemons. For example, if you squeeze a little lemon juice into a glass of water, the solution will become acidic. That’s because lemons contain citric acid.
Bases, on the other hand, do not release hydrogen ions when added to water. They release a lot of hydroxide ions (OH–). These solutions are said to be alkaline (the fancy name for a basic solution). Bases usually have a slippery consistency. Common soap, for example, is composed largely of bases.
The acidity or alkalinity of a solution can be measured using a pH scale. The pH scale is numbered from 1 to 14. The midpoint, 7, is considered neutral pH. The concentration of hydrogen ions in a solution will indicate whether it is acidic, basic, or neutral. If a solution contains a lot of hydrogen ions, then it will be acidic and have a low pH. Here’s the trend:
An increase in H+ ions causes a decrease in the pH.
pH = –log [H+]
Therefore, as the concentration of H+ ions increases by a factor of 10, the pH becomes one number smaller. For example, stomach acid has a pH of 2, and if we use the equation, we discover that the concentration of H+ ions in stomach acid is 10–2 M. This is pretty high when you consider the other extreme is lye, which has a pH of 14, a concentration of H+ ions around 10–14 M! Use your calculator to double check that these numbers are correct.
You’ll notice from the scale below that stronger acids have lower pHs. If a solution has a low concentration of hydrogen ions, it will have a high pH.
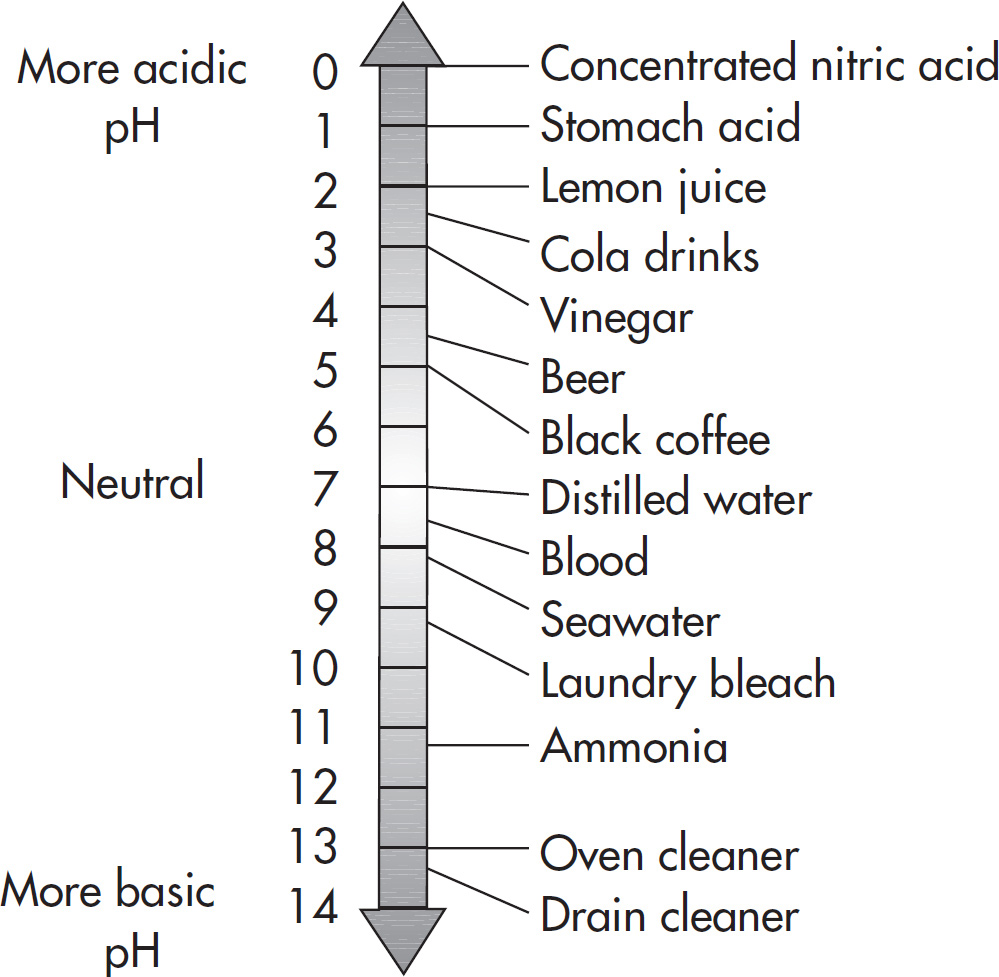
One more thing to remember: The pH scale is not a linear scale—it’s logarithmic. That is, a change of one pH number actually represents a tenfold change in hydrogen ion concentration. For example, a pH of 3 is actually ten times more acidic than a pH of 4. This is also true in the reverse direction: A pH of 4 represents a tenfold decrease in acidity compared to a pH of 3.
The equation for pH is listed on the AP Biology Equations and Formulas sheet. However, you will not be expected to perform calculations using this equation. Instead, you should understand how the equation works and when pH calculations are useful.
ORGANIC MOLECULES
Now that we’ve discussed chemical compounds in general, let’s talk about a special group of compounds. Most of the chemical compounds in living organisms contain a skeleton of carbon atoms. These molecules are known as organic compounds. By contrast, molecules that do not contain carbon atoms are called inorganic compounds. For example, salt (NaCl) is an inorganic compound.
Carbon is important for life because it is a versatile atom, meaning that it not only has the ability to bind with other carbons but also with a number of other elements including nitrogen, oxygen, and hydrogen. The resulting molecules are key in carrying out the activities necessary for life.
To recap:
• Organic compounds contain carbon atoms AND hydrogen atoms (and sometimes other things too).
• Inorganic compounds don’t contain both carbon atoms and hydrogen atoms.
Now let’s focus on the following four classes of organic compounds central to life on Earth:
• carbohydrates
• proteins
• lipids
• nucleic acids
Most macromolecules are chains of building blocks, called polymers. The individual building blocks of a polymer are called monomers.
Carbohydrates
Organic compounds that contain carbon, hydrogen, and oxygen are called carbohydrates. They usually contain these three elements in a ratio of appoximately 1:2:1, respectively. We can represent the proportion of elements within carbohydrate molecules by the formula CnH2nOn, which can be simplified as (CH2O)n.
Most carbohydrates are categorized as either monosaccharides, disaccharides, or polysaccharides. The term saccharides is a fancy word for “sugar.” The prefixes mono-, di-, and poly- refer to the number of sugars in the molecule. Mono- means “one,” di- means “two,” and poly- means “many.” A monosaccharide is therefore a carbohydrate made up of a single sugar molecule.
Monosaccharides: The Simplest Sugars
Monosaccharides, the simplest sugars, serve as an energy source for cells. The two most common sugars are (1) glucose and (2) fructose.
Both of these monosaccharides are six-carbon sugars with the chemical formula C6H12O6. Glucose, the most abundant monosaccharide, is the most popular sugar around. Glucose is an important part of the food we eat and it is the favorite food made by plants during photosynthesis. This glucose can be broken down to give us (and plants) energy. Fructose, the other monosaccharide you need to know for the test, is a common sugar in fruits.
Glucose and fructose can be depicted as either “straight” or “rings.” Both of them are pretty easy to spot; just look for the carbon molecules with a lot of OHs and Hs attached to them.
Here are the two different forms.
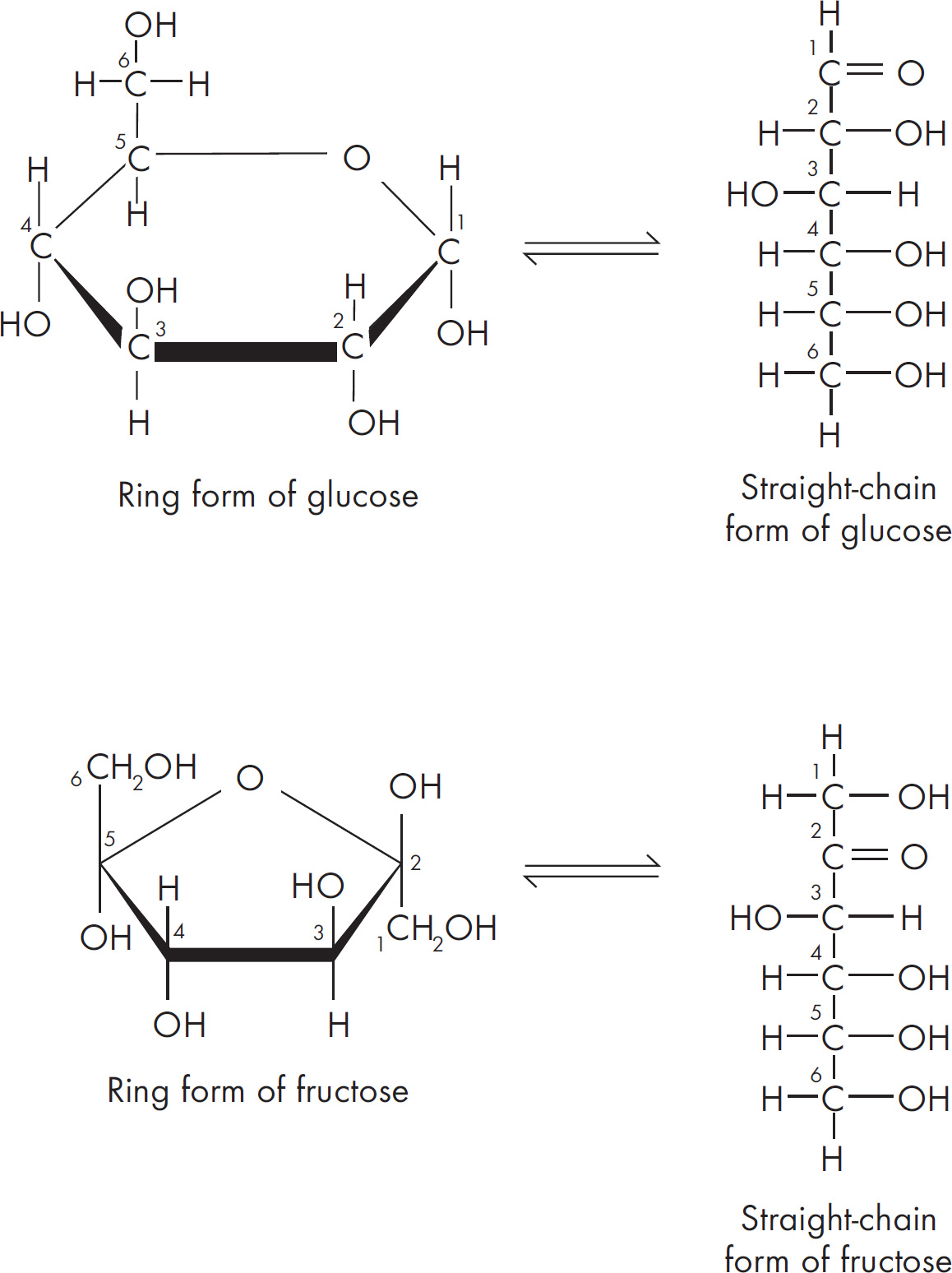
Disaccharides
What happens when two monosaccharides are brought together? The hydrogen (–H) from one sugar molecule combines with the hydroxyl group (–OH) of another sugar molecule. What do H and OH add up to? Water (H2O)!
Maltose is an example of a disaccharide.
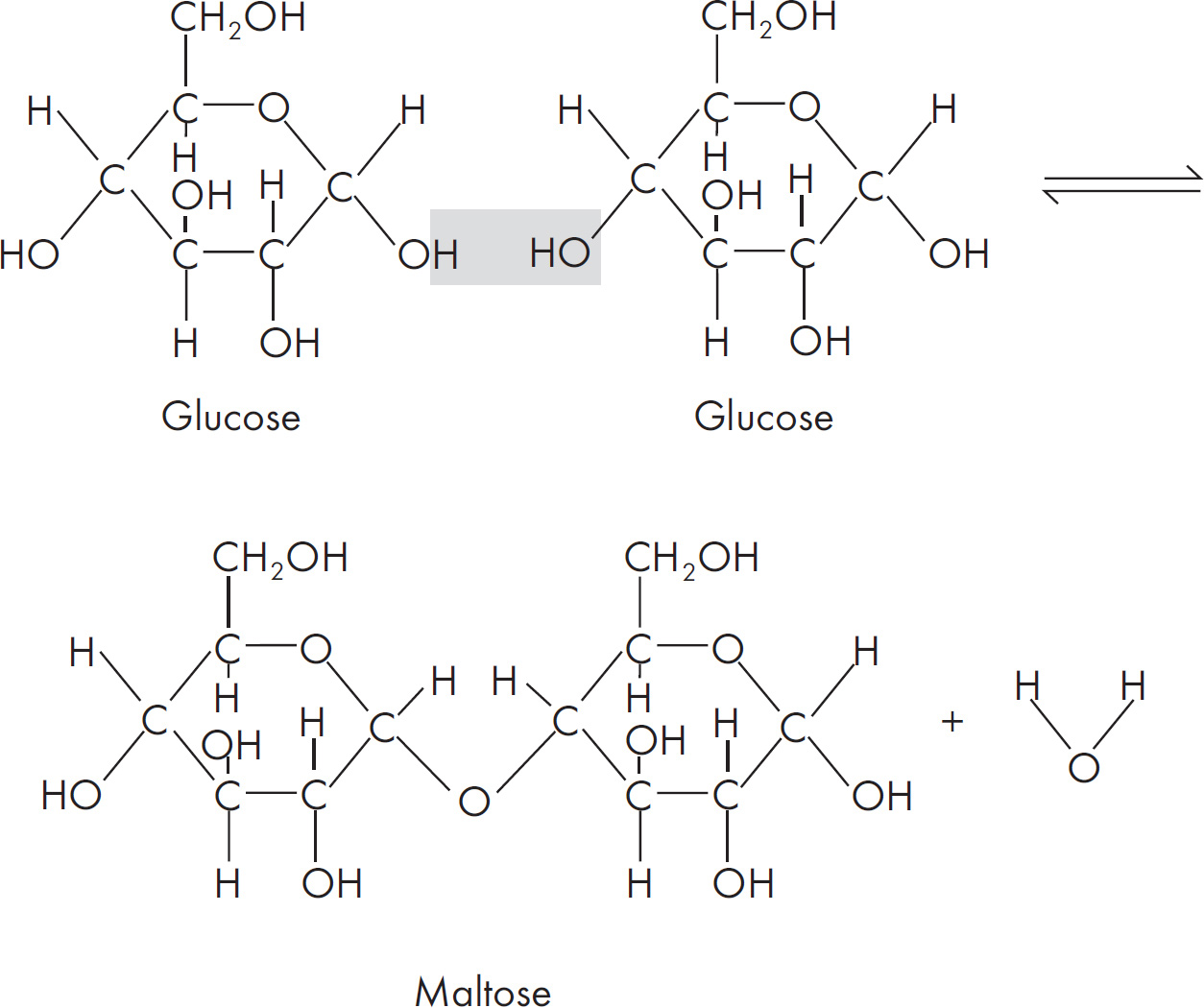
This process is called dehydration synthesis, or condensation, because during this process, a water molecule is lost. When two monosaccharides are joined, the bond is called a glycosidic linkage, and the resulting sugar is called a disaccharide. The resulting sugar is maltose if a disaccharide formed from two glucose molecules.
Now, what if you want to break up the disaccharide and form two monosaccharides again? Just add water. That’s called hydrolysis, which means “water” (hydro-) and “breaking” (-lysis). The water breaks the bond between the two monomers. Hydrolysis is a common reaction in biology.
Polysaccharides
Polysaccharides are made up of many repeated units of monosaccharides. The most common polysaccharides you’ll need to know for the test are starch, cellulose, and glycogen. Animals store glucose molecules in the form of glycogen in the liver and muscle cells. Plants “stockpile” α-glucose in the form of starch in structures called plastids. Cellulose, on the other hand, is made up of β-glucose and is a major part of the cell walls in plants. Its function is to lend structural support. Chitin, a polymer of β-glucose molecules, serves as a structural molecule in the walls of fungus and in the exoskeletons of arthropods.
Here’s an AP question that may come up on the test: why can’t humans digest cellulose? This is because the bonds joining the glucose subunits in cellulose are different than those in starch. Starch is composed of α-glucose subunits held together by 1,4 glycosidic linkages, while cellulose contains β-glucose subunits held together by 1,4 linkages. Humans cannot digest the β-linkages; however, humans can digest polysaccharides that have α-linkages, like starch and glycogen.
Proteins
Amino acids are organic molecules that serve as the building blocks of proteins. They contain carbon, hydrogen, oxygen, and nitrogen atoms. There are twenty different amino acids commonly found in proteins. Fortunately, you don’t have to memorize the twenty amino acids. But you do have to remember that every amino acid has four important parts: an amino group (–NH2), a carboxyl group (–COOH), a hydrogen, and an R group.
Here’s a typical amino acid.
Amino acids differ only in the R group, which is also called the side chain. The R group associated with an amino acid could be as simple as a hydrogen atom (as in the amino acid glycine) or as complex as a charged carbon skeleton (as in the amino acid arginine).
When it comes to spotting an amino acid, simply keep an eye out for the amino group (NH2), then look for the carboxyl molecule (COOH). The side chains for each amino acid vary greatly. You don’t have to memorize the structures of the side chains for individual amino acids, but you should know a few key characteristics they can have.
How do the side chains of the amino acids differ?
• Composition of elements (C, H, O, N, and S)
• Polarity (polar, nonpolar)
• Charge (neutral, positive, negative)
• Shape (long chain, short chain, ring-shape)
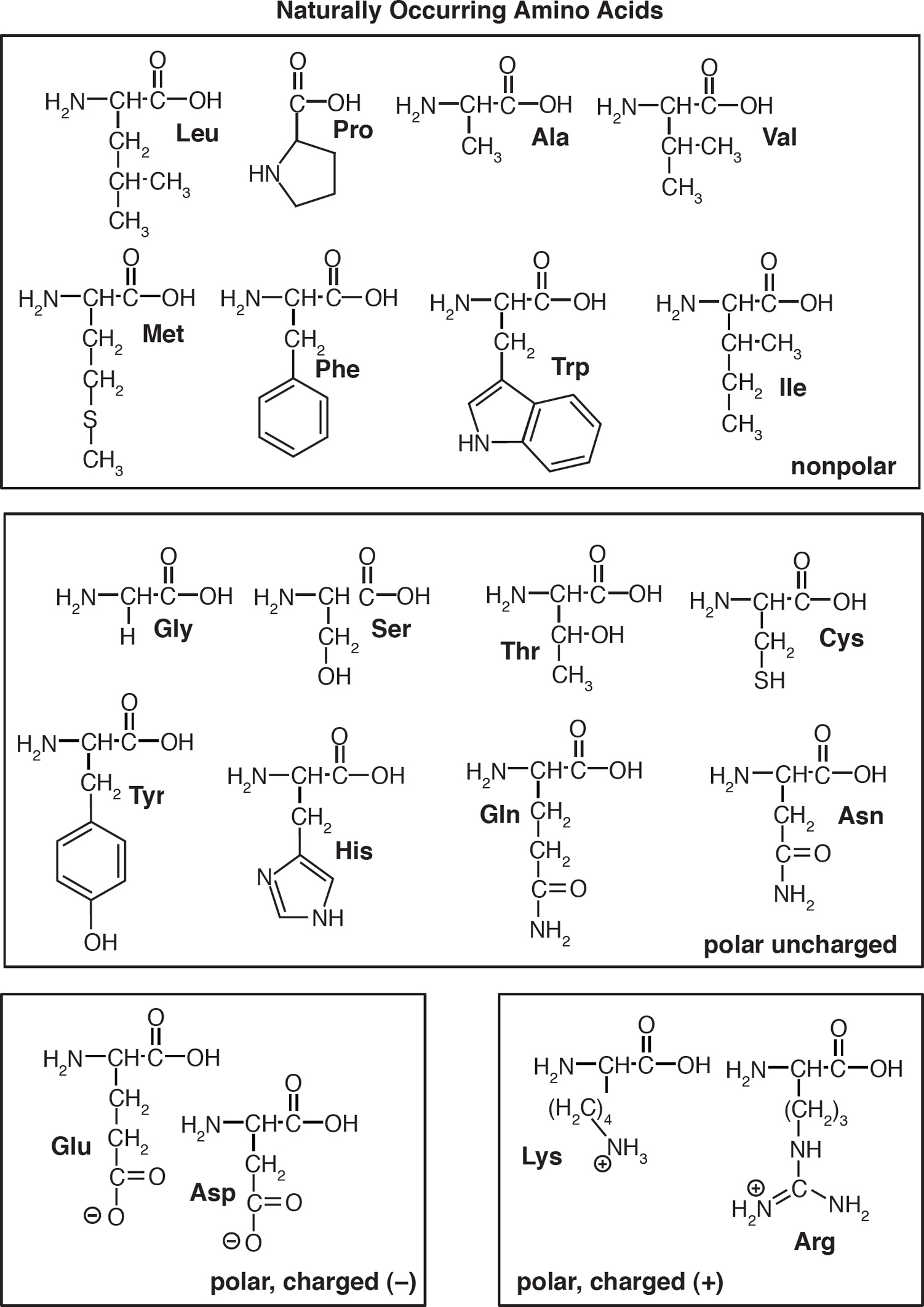
Polypeptides
When two amino acids join they form a dipeptide. The carboxyl group of one amino acid combines with the amino group of another amino acid. Here’s an example:
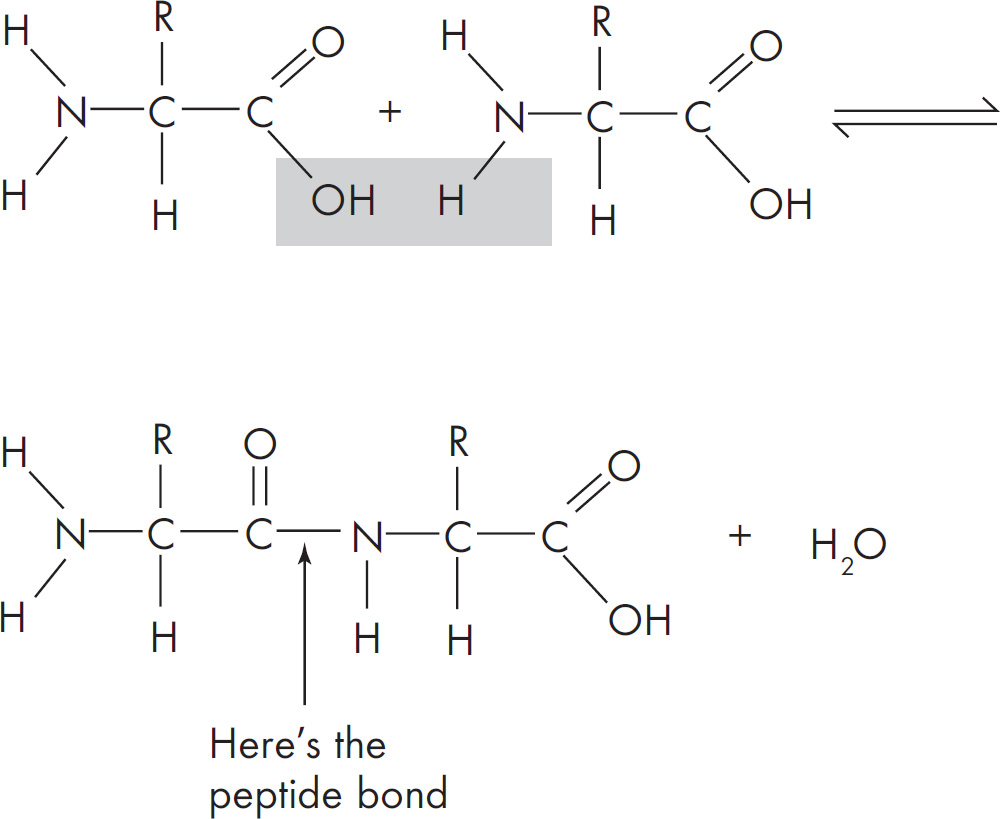
This is the same dehydration synthesis process we saw earlier. Why? Because a water molecule is removed to form a bond. By the way, the bond between two amino acids has a special name—a peptide bond. If a group of amino acids are joined together in a “string,” the resulting organic compound is called a polypeptide. Once a polypeptide chain twists and folds on itself, it forms a three-dimensional structure called a protein.
Higher Protein Structure
The polypeptide has to go through several changes before it can officially be called a protein. Proteins can have four levels of structure. The linear sequence of the amino acids is called the primary structure of a protein. Now the polypeptide begins to twist, forming either a coil (known as an alpha helix) or zigzagging pattern (known as beta-pleated sheets). These are examples of proteins’ secondary structures.
A polypeptide folds and twists because the different R-groups of the amino acids are interacting with each other. Remember, each R-group is unique and has a particular size, shape, charge, etc. that allows it to react to things around it. Depending on which amino acids are in a protein and the order that they are in, the protein can twist and fold in very different ways. This is why proteins can form so many different shapes.
The secondary structure is formed by amino acids that interact with other amino acids closeby in the primary structure. However, after the secondary structure reshapes the polypeptide, amino acids that were far away in the primary structure arrangement can now also interact with each other. This is called the tertiary structure. Sometimes, two cysteine amino acids can react with each other to form a covalent disulfide bond that stabilizes the tertiary structure.
Lastly, several different polypeptide chains sometimes interact with each other to form a quaternary structure. Only proteins that have folded correctly into their specific three-dimensional structure can perform their function properly. Mistakes in the amino acid chain can create nonfunctional proteins.
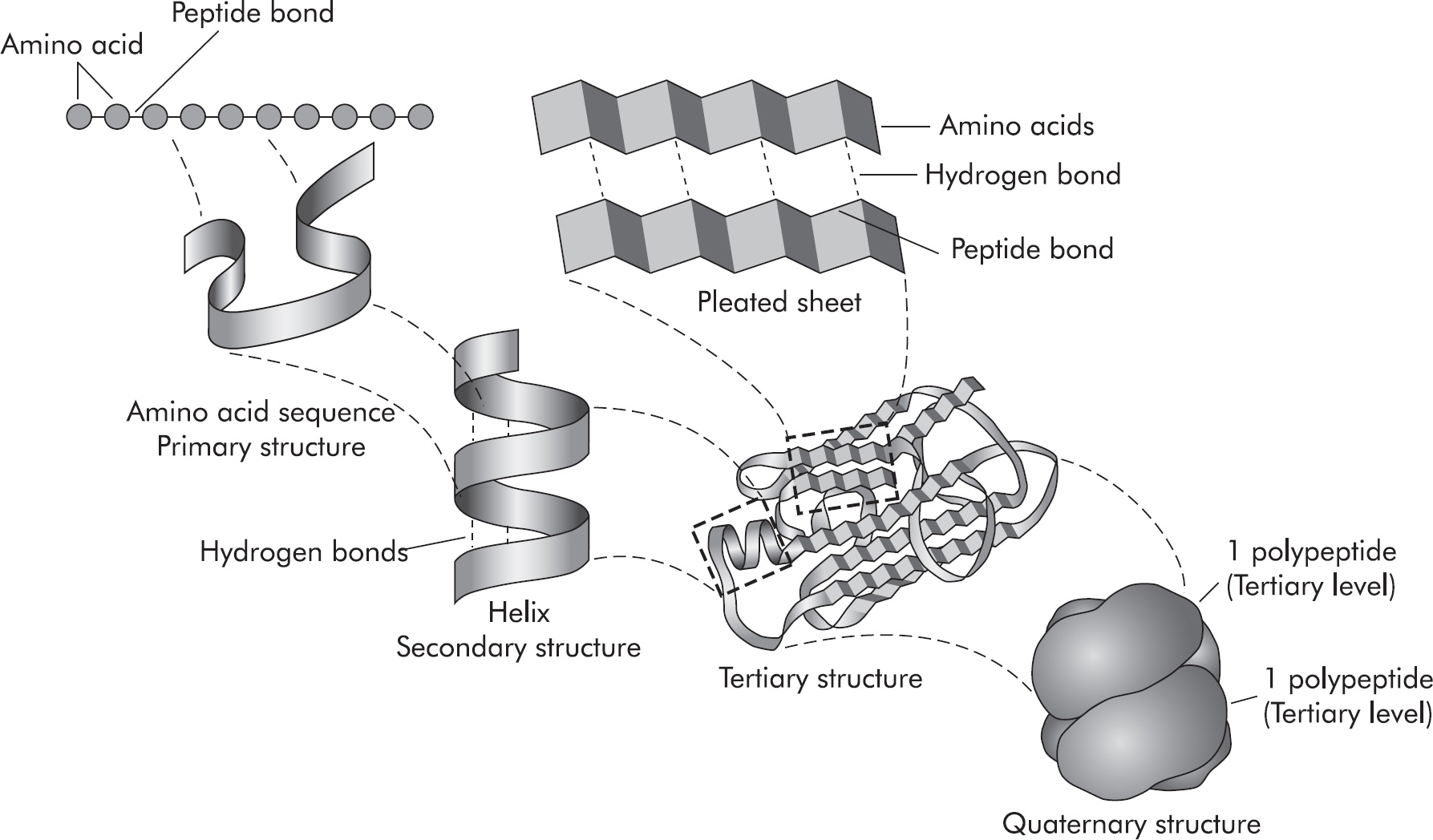
One more thing: In some cases, the folding of proteins involves other proteins known as chaperone proteins (or chaperonins). They help the protein fold properly and make the process more efficient.
Lipids
Like carbohydrates, lipids consist of carbon, hydrogen, and oxygen atoms, but not in the 1:2:1 ratio typical of carbohydrates. The most common examples of lipids are triglycerides, phospholipids, and steroids.
Triglycerides
Our fat stores in the body are filled with lipids called triglycerides. Each triglyceride is made of a glycerol molecule with three fatty acid chains attached to it.
Let’s take a look.
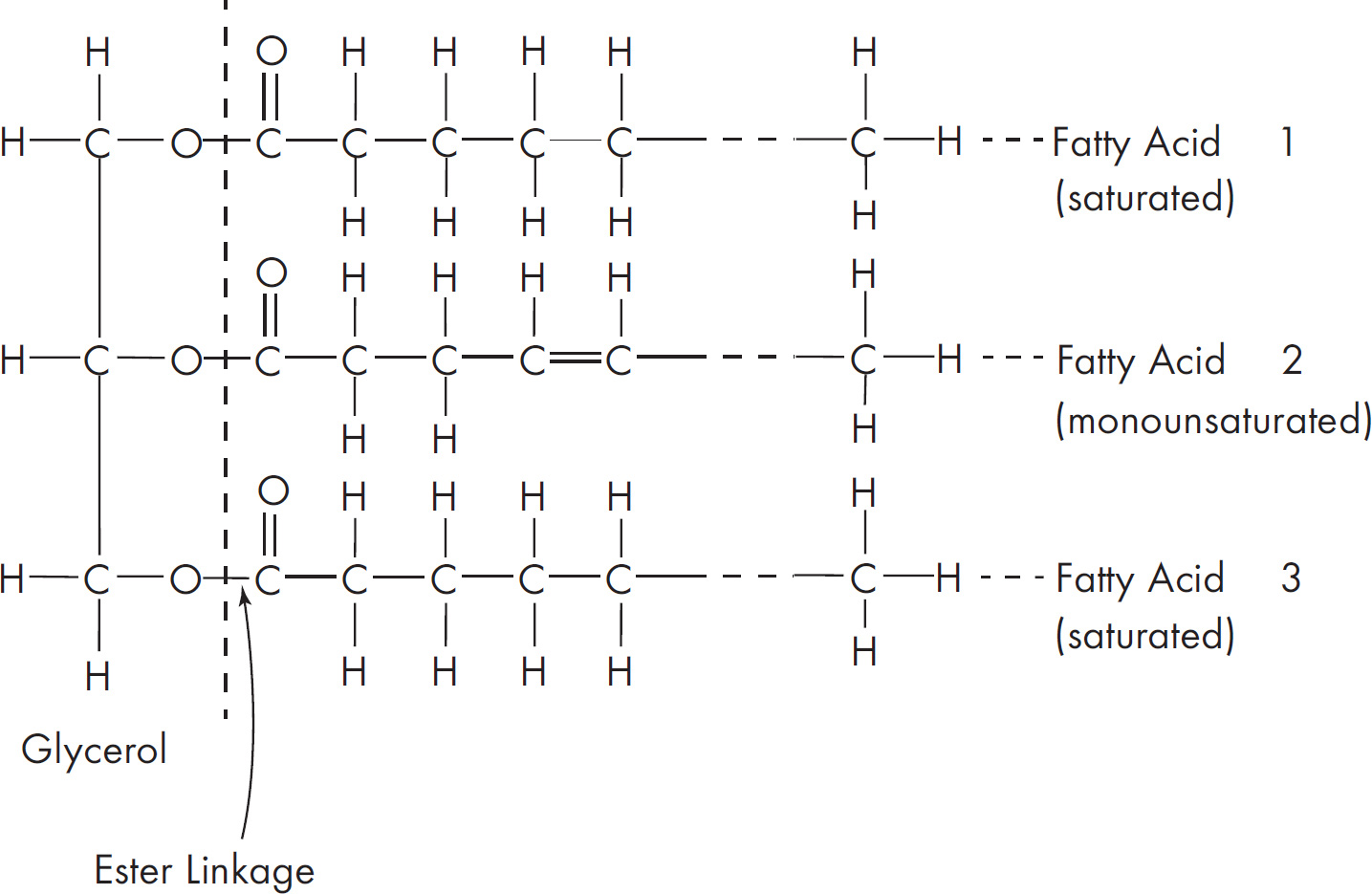
To make a triglyceride, each of the carboxyl groups (–COOH) of the three fatty acids must react with one of the three hydroxyl groups (–OH) of the glycerol molecule. This happens by the removal of a water molecule. So, the creation of a fat requires the removal of three molecules of water. Once again, what have we got? You probably already guessed it—dehydration synthesis! The linkage now formed between the glycerol molecule and the fatty acids is called an ester linkage. A fatty acid can be saturated, which means it has a single covalent bond between each pair of carbon atoms, or it can be unsaturated, which means adjacent carbons are joined by double bonds instead of single bonds. A polyunsaturated fatty acid has many double bonds within the fatty acid.
Lipids are important because they function as structural components of cell membranes, sources of insulation, and a means of energy storage.
Phospholipids
Another special class of lipids is known as phospholipids. Phospholipids contain two fatty acid “tails” and one negatively charged phosphate “head.” Take a look at a typical phospholipid.
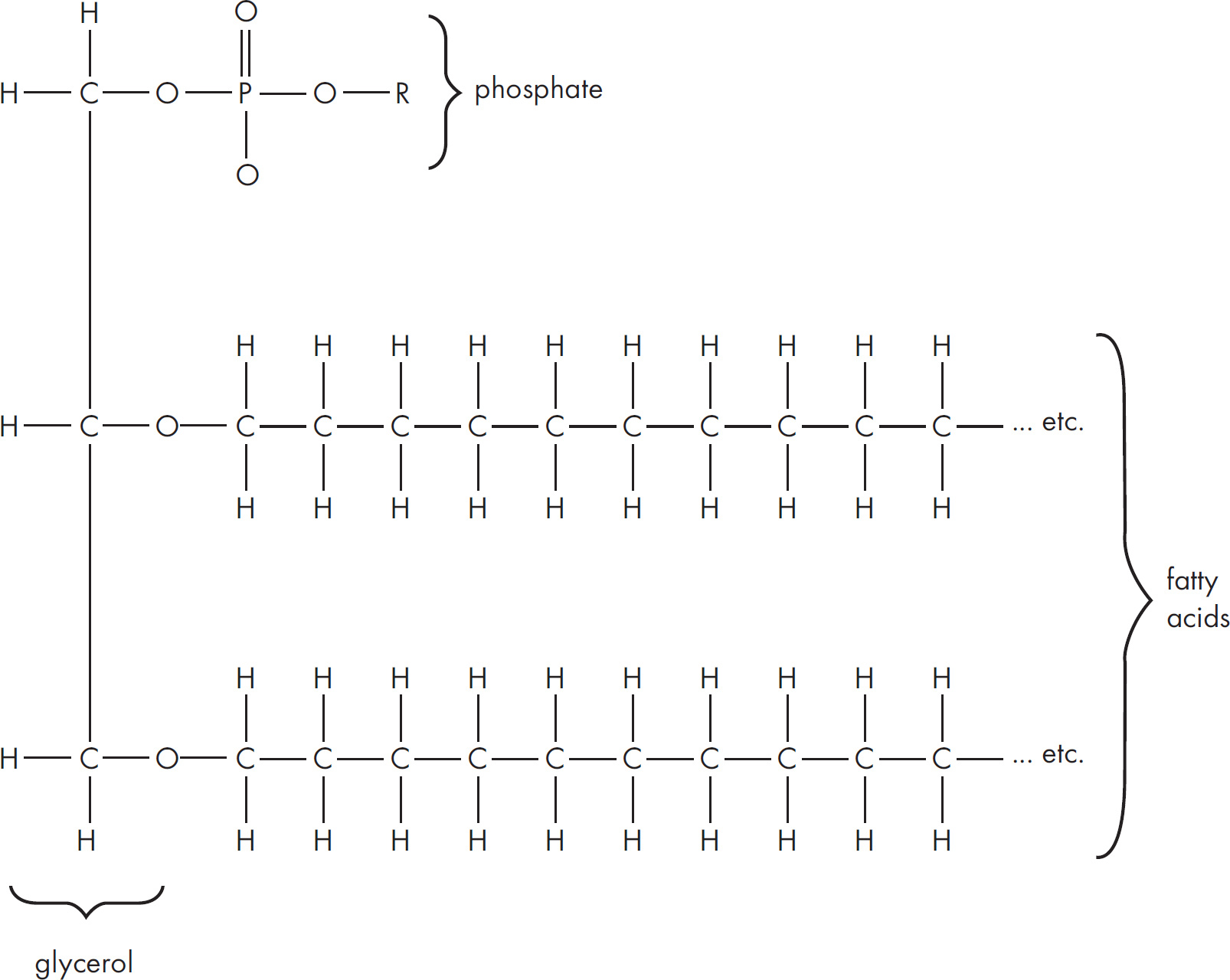
Phospholipids are extremely important, mainly because of some unique properties they possess, particularly with regard to water.
The two fatty acid tails are hydrophobic (“water-hating”). In other words, just like oil and vinegar, fatty acids and water don’t mix. The reason for this is that fatty acid tails are nonpolar, and nonpolar substances don’t mix well with polar ones, such as water.
On the other hand, the phosphate “head” of the lipid is hydrophilic (“water- loving”), meaning that it does mix well with water. Why? It carries a negative charge, and this charge draws it to the positively charged end of a water molecule. Since a phospholipid has both a hydrophilic region and a hydrophobic region, it is an amphipathic molecule. One side of a phospholipid loves to hang out with water, and the other side hates to.
Thus, the two fatty acid chains orient themselves away from water, while the phosphate portion orients itself toward the water. Keep these properties in mind. We’ll see later how this orientation of phospholipids in water relates to the structure and function of cell membranes.
Cholesterol
Cholesterol is another important type of lipid. Cholesterol is a a four-ringed molecule that is found in membranes. It affects membrane fluidity by preventing it from freezing or melting. Cholesterol is also important for making certain types of hormones and for making vitamin D.
Nucleic Acids
The fourth class of organic compounds are the nucleic acids. Like proteins, nucleic acids contain carbon, hydrogen, oxygen, and nitrogen, but nucleic acids also contain phosphorus. Nucleic acids are molecules that are made up of simple units called nucleotides. For the AP Biology Exam, you’ll need to know about two kinds of nucleic acids: deoxyribonucleic acid (DNA) and ribonucleic acid (RNA).
DNA is important because it contains the hereditary “blueprints” of all life. RNA is important because it’s essential for protein synthesis. We’ll discuss DNA and RNA in greater detail when we discuss heredity. The following table summarizes the important macromolecules discussed here.
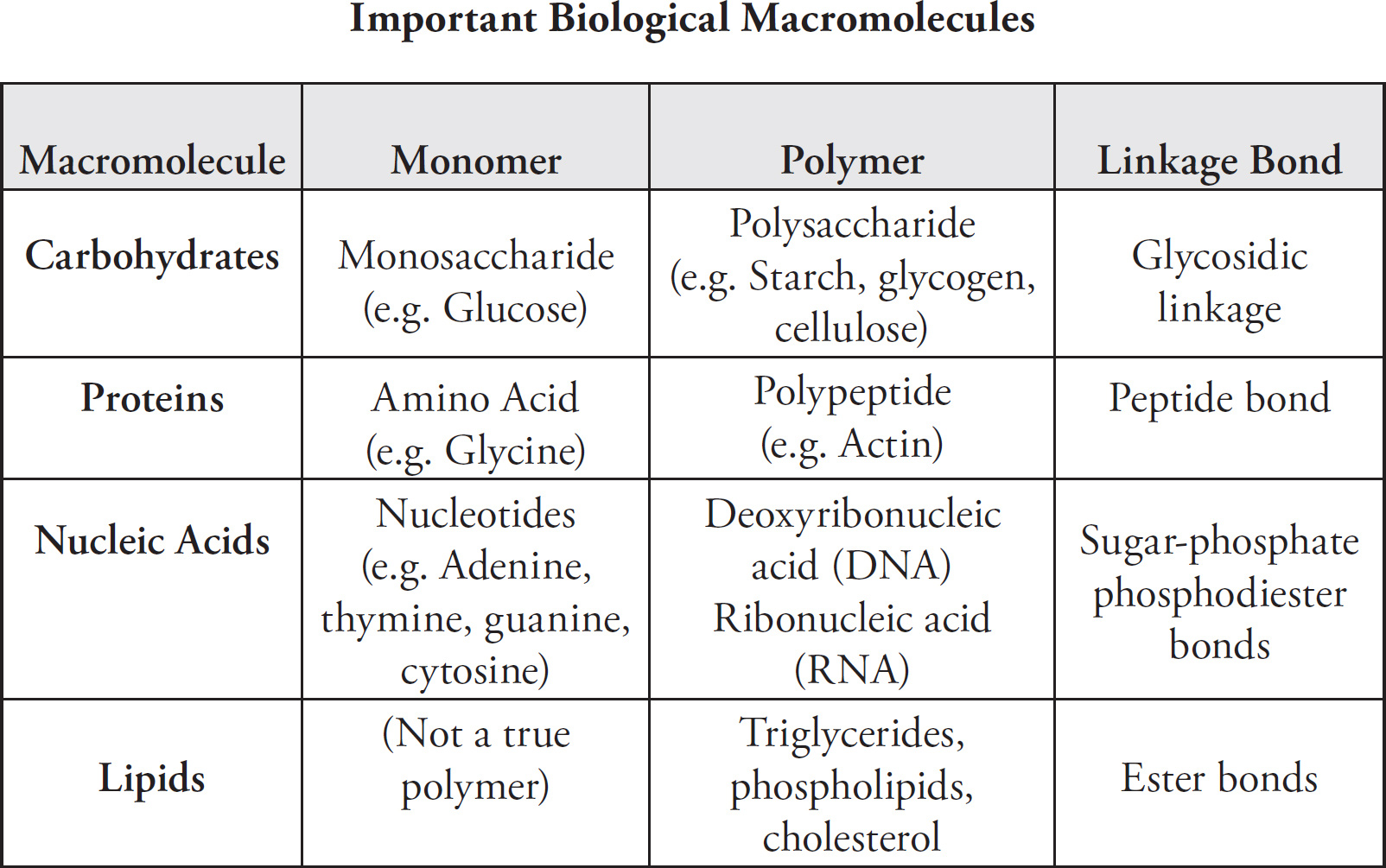
ORIGINS OF THE EARTH
We’ve just seen the organic compounds that are essential for life. But where did they come from in the first place? This is still a hotly debated topic among scientists. Most scientists believe that the earliest precursors of life arose from nonliving matter (basically gases) in the primitive oceans of the earth. But this theory didn’t take shape until the 1920s. Two scientists, Alexander Oparin and J. B. S. Haldane, proposed that the primitive atmosphere contained the following gases: methane (CH4), ammonia (NH3), hydrogen (H2), and water (H2O). Interestingly enough, there was almost no free oxygen (O2) in this early atmosphere. They believed that these gases collided, producing chemical reactions that eventually led to the organic molecules we know today.
This theory didn’t receive any substantial support until 1953. In that year, Stanley Miller and Harold Urey simulated the conditions of primitive Earth in a laboratory. They put the gases theorized to be abundant in the early atmosphere into a flask, struck them with electrical charges in order to mimic lightning, and organic compounds similar to amino acids appeared!
But how do we make the leap from simple organic molecules to more complex compounds and life as we know it? Since no one was around to witness the process, no one knows for sure how (or when) it occurred. It is likely that the original life-forms were simply molecules of RNA. This is called the RNA-world hypothesis. RNA can take many shapes because it is not restricted to be a double helix. It is possible that RNA molecules capable of replicating and thus passing along themselves (their genome) were the first life-forms. Complex organic compounds (such as proteins) must have formed via dehydration synthesis. Simple cells then used organic molecules as their source of food. Over time, simple cells evolved into complex cells.
Now let’s throw in a few new terms. Living organisms that rely on organic molecules for food are called heterotrophs (or consumers). For example, we’re heterotrophs, but the earliest heterotrophs were simple unicellular life-forms. Eventually, some life-forms found a way to make their own food; most commonly through photosynthesis. These organisms are called autotrophs (or producers). Early autotrophs (most likely cyanobacteria) are responsible for Earth’s oxygenated atmosphere.
KEY TERMS
elements
oxygen
carbon
hydrogen
nitrogen
trace elements
atom
protons
neutrons
electrons
nucleus
isotopes
radiometric dating
compound
chemical reaction
chemical bond
ionic bond
covalent bond
hydrogen bond
ions
nonpolar covalent
polar covalent
polar
cohesion
adhesion
heat capacity
expansion on freezing
capillary action
acidic
basic
neutral
alkaline
pH scale
organic compounds
inorganic compounds
polymer
monomer
carbohydrates
monosaccharides
disaccharides
polysaccharides
glucose
fructose
dehydration synthesis (or condensation)
glycosidic bond
hydrolysis
starch
cellulose
glycogen
plastids
amino acids
amino group
carboxyl group
R group
side chain
dipeptide
peptide bond
polypeptide
protein
primary structure
secondary structure
tertiary structure
quaternary structure
chaperone proteins
lipids
triglycerides
cholesterol
phospholipids
steroids
glycerol
ester linkage
saturated
unsaturated
polyunsaturated
hydrophobic
hydrophilic
amphipathic
nucleic acids
nucleotides
deoxyribonucleic acid (DNA)
ribonucleic acid (RNA)
Alexander Oparin and J. B. S. Haldane
Stanley Miller and Harold Urey
RNA-world hypothesis
heterotrophs (or consumers)
autotrophs (or producers)
Summary
 Compounds are composed of two or more types of elements in a fixed ratio.
Compounds are composed of two or more types of elements in a fixed ratio.
 Compounds are held together by ionic bonds and covalent bonds.
Compounds are held together by ionic bonds and covalent bonds.
 Hydrogen bonding is essential for the following emergent properties of water:
Hydrogen bonding is essential for the following emergent properties of water:
• cohesion
• adhesion
• high heat capacity
• expansion upon freezing
• polar structure
• ability to dissolve other polar (hydrophilic) substances but cannot dissolve nonpolar (hydrophobic) substances
 pH is important for biological reactions. Solutions may be acidic, basic or alkaline, or neutral.
pH is important for biological reactions. Solutions may be acidic, basic or alkaline, or neutral.
 Organic molecules contain carbon atoms. Many molecules in biology are composed of several monomers bound together into polymers. The creation of these polymers occurs through dehydration synthesis reactions. The breakdown of these polymers occurs through hydrolysis reactions.
Organic molecules contain carbon atoms. Many molecules in biology are composed of several monomers bound together into polymers. The creation of these polymers occurs through dehydration synthesis reactions. The breakdown of these polymers occurs through hydrolysis reactions.
 Important organic macromolecules in biology include
Important organic macromolecules in biology include
• carbohydrates: monosaccharides link by glyosidic bonds to form di- and polysaccharides. Examples are glucose, fructose, glycogen, cellulose, and starch.
• proteins: twenty different amino acids, each with a different R-group, link together via peptide bonds to form polypeptides. The chain further folds up into special shapes to make the final protein structure.
• lipids: chains called fatty acids (either saturated or unsaturated) commonly form as phospholipids and triglycerides. Cholesterol is another type of lipid. Lipids are hydrophobic.
• nucleic acids: chains of either DNA or RNA nucleotides. They carry the genetic recipes in the body.
 Heterotrophs are consumers and autotrophs are producers.
Heterotrophs are consumers and autotrophs are producers.
Chapter 4 Drill
Answers and explanations can be found in Chapter 16.
1. Water is a critical component of life due to its unique structural and chemical properties. Which of the following does NOT describe a way that the exceptional characteristics of water are used in nature to sustain life?
(A) The high heat capacity of water prevents lakes and streams from rapidly changing temperature and freezing completely solid in the winter.
(B) The high surface tension and cohesiveness of water facilitates capillary action in plants.
(C) The low polarity of water prevents dissolution of cells and compounds.
(D) The high intermolecular forces of water, such as hydrogen bonding, result in a boiling point which exceeds the tolerance of most life on the planet.
2. The intracellular pH of human cells is approximately 7.4. Yet, the pH within the lumen (inside) of the human stomach averages 1.5. Which of the following accurately describes the difference between the acidity of the cellular and gastric pH?
(A) Gastric juices contain approximately 5 times more H+ ions than the intracellular cytoplasm of cells and are more acidic.
(B) Gastric juices contain approximately 100,000-fold more H+ ions than the intracellular cytoplasm of cells and are more acidic.
(C) The intracellular cytoplasm of cells contain approximately 5 times more H+ ions than gastric juices and is more acidic.
(D) The intracellular cytoplasm of cells contains approximately 100,000-fold more H+ ons than gastric juices and is more acidic.
3. Amino acids are the basic molecular units which compose proteins. All life on the planet forms proteins by forming chains of amino acids. Which labeled component of the amino acid structure of phenylalanine shown below will vary from amino acid to amino acid?
Questions 4-6 refer to the following paragraph and diagram.
In 1953, Stanley Miller and Harold Urey performed an experiment at the University of Chicago to test the hypothesis that the conditions of the early Earth would have favored the formation of larger, more complex organic molecules from basic precursors. The experiment, as shown below, consisted of sealing basic organic chemicals (representing the atmosphere of the primitive Earth) in a flask, which was exposed to electric sparks (to simulate lightning) and water vapor.
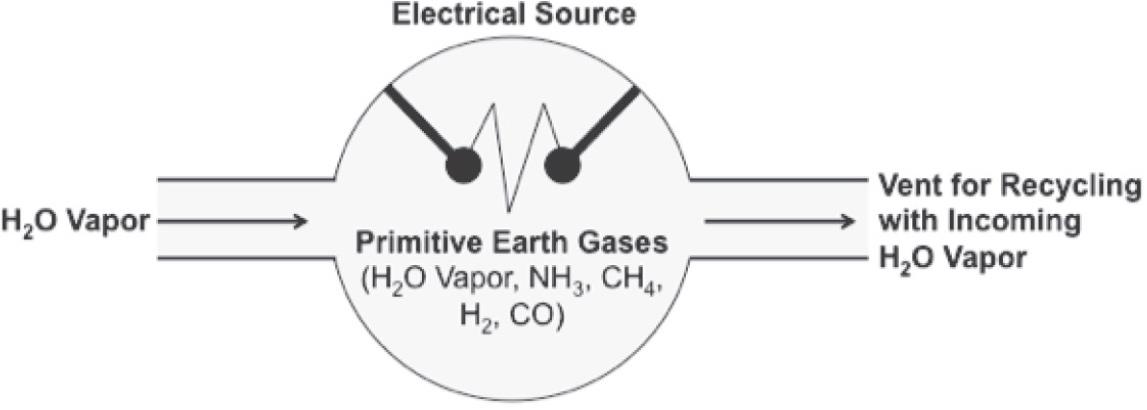
After one day of exposure, the mixture in the flask had turned pink in color, and later analysis showed that at least 10% of the carbon had been transformed into simple and complex organic compounds including at least 11 different amino acids and some basic sugars. No nucleic acids were detected in the mixture.
4. Which of the following contradicts the hypothesis of the experiment that life may have arisen from the formation of complex molecules in the conditions of the primitive Earth?
(A) Complex carbon-based compounds were generated after only one day of exposure to simulated primitive Earth conditions.
(B) Nucleic acid compounds such as DNA and RNA were not detected in the mixture during the experiment.
(C) Over half of known amino acids involved in life were detected in the mixture during the experiment.
(D) Basic sugar molecules were generated and detected in the mixture during the experiment.
5. Some amino acids, such as cysteine (shown below) and methionine, could not be formed in this experiment. Which of the following best explains why these molecules could not be detected?
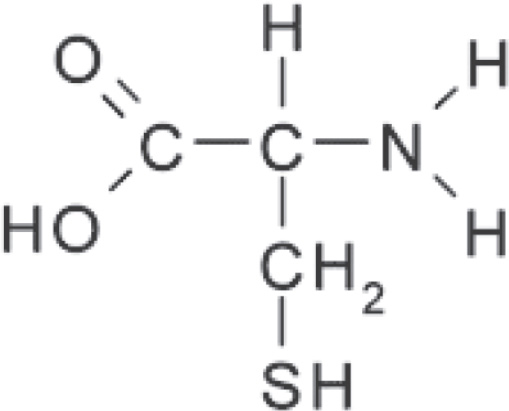
(A) The chemical reactions necessary to create amino acids such as cysteine and methionine require more energy than the simulated lightning provided in the experiment.
(B) The chemical reactions necessary to create amino acids such as cysteine and methionine require enzymes for catalysis to occur, which were not included in the experiment.
(C) Sulfur-based compounds were not included in the experiment.
(D) Nitrogen-based compounds were not included in the experiment.
6. A scientist believes that the Miller-Urey experiment failed to yield the remaining amino acids and the nucleic acids because of the absence of critical chemical substrates that would have existed on the primordial Earth due to volcanism. All of the following basic compounds which are associated with volcanism should be added in a follow-up Miller-Urey experiment EXCEPT:
(A) H2S gas
(B) SiO2 (silica)
(C) SO2
(D) H3PO4 (phosphoric acid)
7. Catabolism refers to breaking down complex macromolecules into their basic components. Many biological processes use hydrolysis for catabolism. Hydrolysis of proteins could directly result in
(A) free water
(B) adenine
(C) cholesterol
(D) dipeptides
8. Which of the following contain both hydrophilic and hydrophobic properties and are often found in cell plasma membranes?
(A) Nucleotides
(B) Phospholipids
(C) Water
(D) Amino acids
9. Maltotriose is a trisaccharide composed of three glucose molecules linked through α-1,4 glycosidic linkages formed via dehydration synthesis. What would the formula be for maltotriose?
(A) C18H36O18
(B) C18H10O15
(C) C18H32O16
(D) C3H6O3
10. A radioactive isotope of hydrogen, 3H, is called tritium. Tritium differs from the more common form of hydrogen because
(A) it contains two neutrons and one proton in its nucleus
(B) it contains one neutron and two protons in its nucleus
(C) it differs by its atomic number
(D) it is radioactive and therefore gives off one electron
REFLECT
Respond to the following questions:
• Which topics from this chapter do you feel you have mastered?
• Which content topics from this chapter do you feel you need to study more before you can answer multiple-choice questions correctly?
• Which content topics from this chapter do you feel you need to study more before you can effectively compose a free response?
• Was there any content that you need to ask your teacher or another person about?
Chapter 5
Cells
LIVING THINGS
All living things are composed of cells. According to the cell theory, the cell is life’s basic unit of structure and function. This simply means that the cell is the smallest unit of living material that can carry out all the activities necessary for life.
It may seem strange that we are still made of tiny little cells. Why not just be a living thing that is one giant cell?
One reason for this is because compartments allow more specialization. As we will see, compartmentalization is an important part of the organization of unicellular and multicellular organisms.
The second reason we can’t just be giant cells is the surface area-to-volume ratio. There is always lots of exchange going on between the inside of things and the outside. This ratio must be kept small so that there is lots of space to do these exchanges.
Therefore, even the largest things are still made of cells.
TYPES OF CELLS AND ORGANELLES
For centuries, scientists have known about cells. However, it wasn’t until the development of the electron microscope that scientists were able to figure out what cells do. We now know that there are two distinct types of cells: prokaryotic cells and eukaryotic cells.
MICROSCOPES AND THE STUDY OF CELLS
Cells are studied using different types of microscopes. Light microscopes, (for example, the compound microscopes commonly found in labs) are used to study stained or living cells. They can magnify the size of an organism up to 1,000 times. Electron microscopes are used to study detailed structures of a cell that cannot be easily seen or observed by light microscopy. They are capable of resolving structures as small as a few nanometers in length, such as individual virus particles or the pores on the surface of the nucleus.
A prokaryotic cell, which is a lot smaller than a eukaryotic cell, is relatively simple. Bacteria, for example, are prokaryotic cells. The inside of the cell is filled with a substance called cytoplasm. The genetic material in a prokaryote is one continuous, circular DNA molecule that is found free in the cell in an area called the nucleoid (this is not the same as a nucleus!). Most prokaryotes have a cell wall composed of peptidoglycans that surrounds a lipid layer called the plasma membrane. Prokaryotes also have ribosomes (though smaller than those found in eukaryotic cells), and they may also have one or more flagella, which are long projections used for motility (movement). Eukaryotic cells are more complex than prokaryotic cells. Fungi, protists, plants, and animals are eukaryotes.
Eukaryotic cells are more complex than prokaryotic cells. Fungi, protists, plants, and animals are eukaryotes. Eukaryotic cells are organized into many smaller structures called organelles. Some of these organelles are the same structures seen in prokaryotic cells, but many are uniquely eukaryotic. A good way to remember the difference is that prokaryotes do not have any membrane-bound organelles. Their only membrane is the plasma membrane.
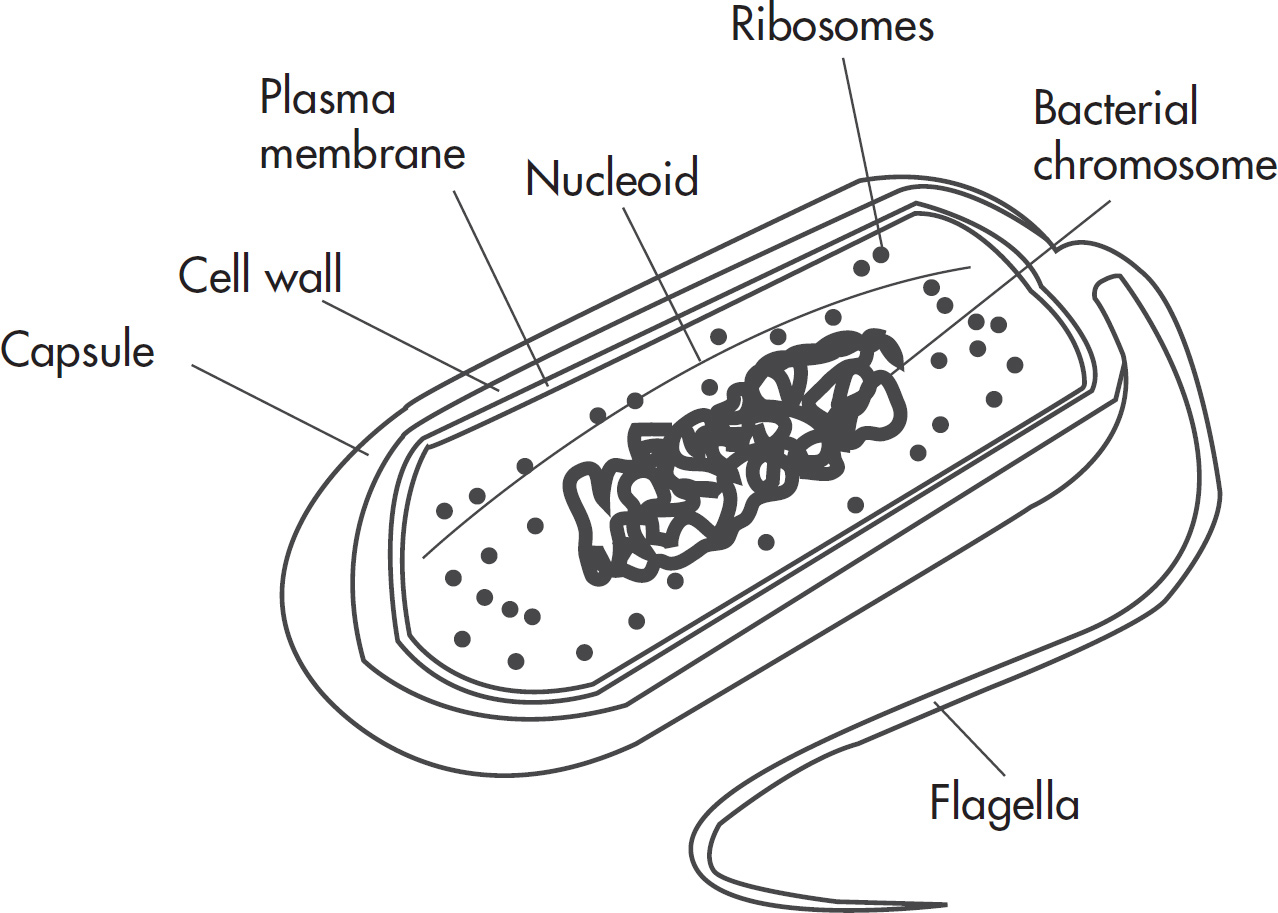
Organelles
A eukaryotic cell is like a microscopic factory. It’s filled with organelles, each of which has its own special tasks. Let’s take a tour of a eukaryotic cell and focus on the structure and function of each organelle. Here’s a picture of a typical animal cell and its principal organelles.
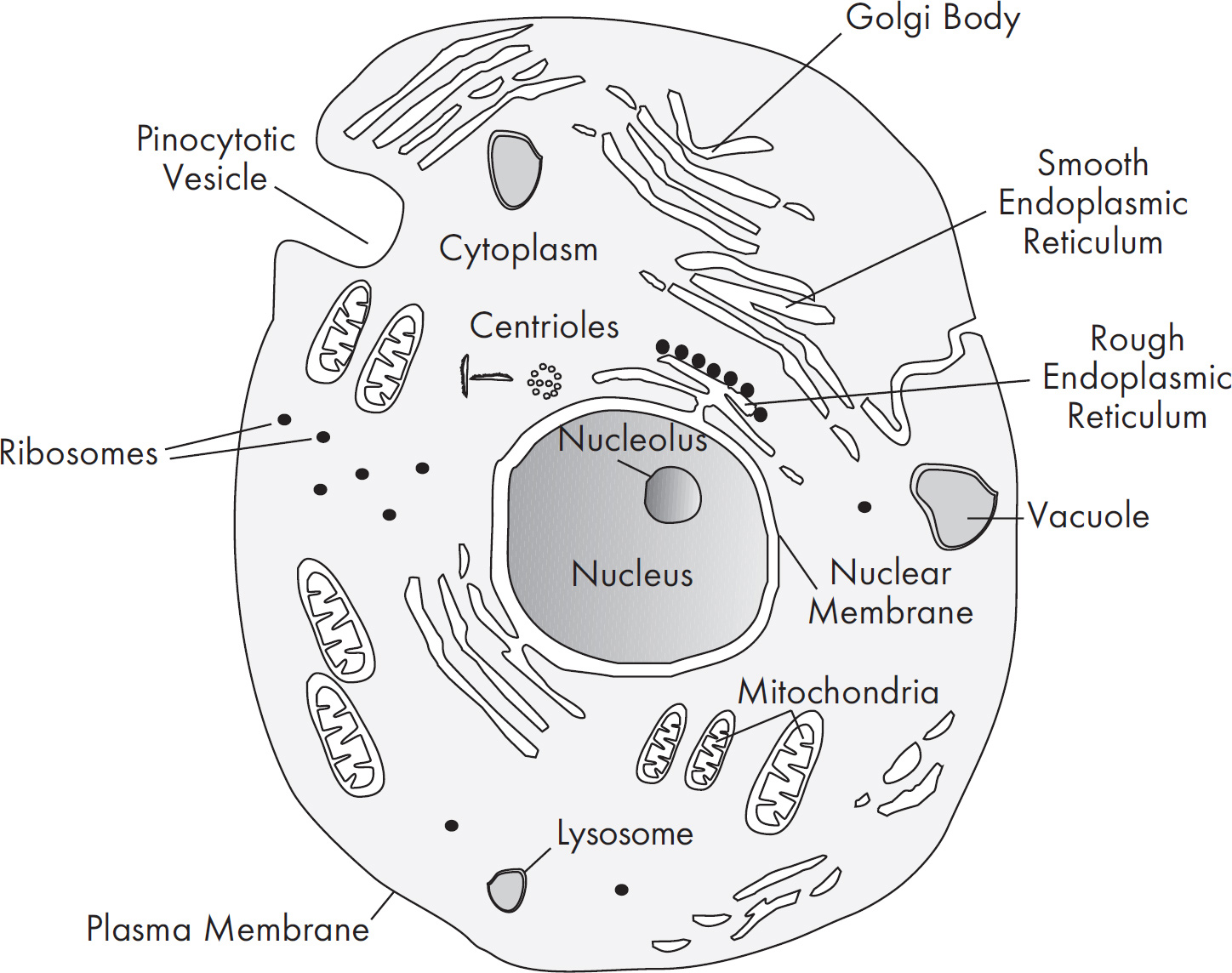
Plasma Membrane
The cell has an outer envelope known as the plasma membrane. Although the plasma membrane appears to be a simple, thin layer surrounding the cell, it’s actually a complex, double-layered structure made up of phospholipids and proteins. The hydrophobic fatty acid tails face inward and the hydrophilic phosphate heads face outward. It is called a phospholipid bilayer since two lipid layers are forming a hydrophobic sandwich.
The plasma membrane is important because it regulates the movement of substances into and out of the cell. The membrane itself is semipermeable, meaning that only certain substances, namely small hydrophobic molecules (such as O2 and CO2), pass through it unaided. Anything large and/or hydrophilic can only pass through the membrane via special tunnels (discussed more later in this chapter).
Many proteins are associated with the cell membrane. Some of these proteins are loosely associated with the lipid bilayer (peripheral proteins). They are located on the inner or outer surface of the membrane. Others are firmly bound to the plasma membrane (integral proteins). These proteins are amphipathic, which means that their hydrophilic regions extend out of the cell or into the cytoplasm, while their hydrophobic regions interact with the tails of the membrane phospholipids. Some integral proteins extend all the way through the membrane (transmembrane proteins).
This arrangement of phospholipids and proteins is known as the fluid-mosaic model. This means that each layer of phospholipids is flexible, and it is a mosaic because it is peppered with different proteins and carbohydrate chains. Remember, anything hydrophilic should not go through the hydrophobic interior. This means that the phospholipids on one side should never flip-flop to the other side of the membrane (because that would require their polar heads to pass through the hydrophobic area).
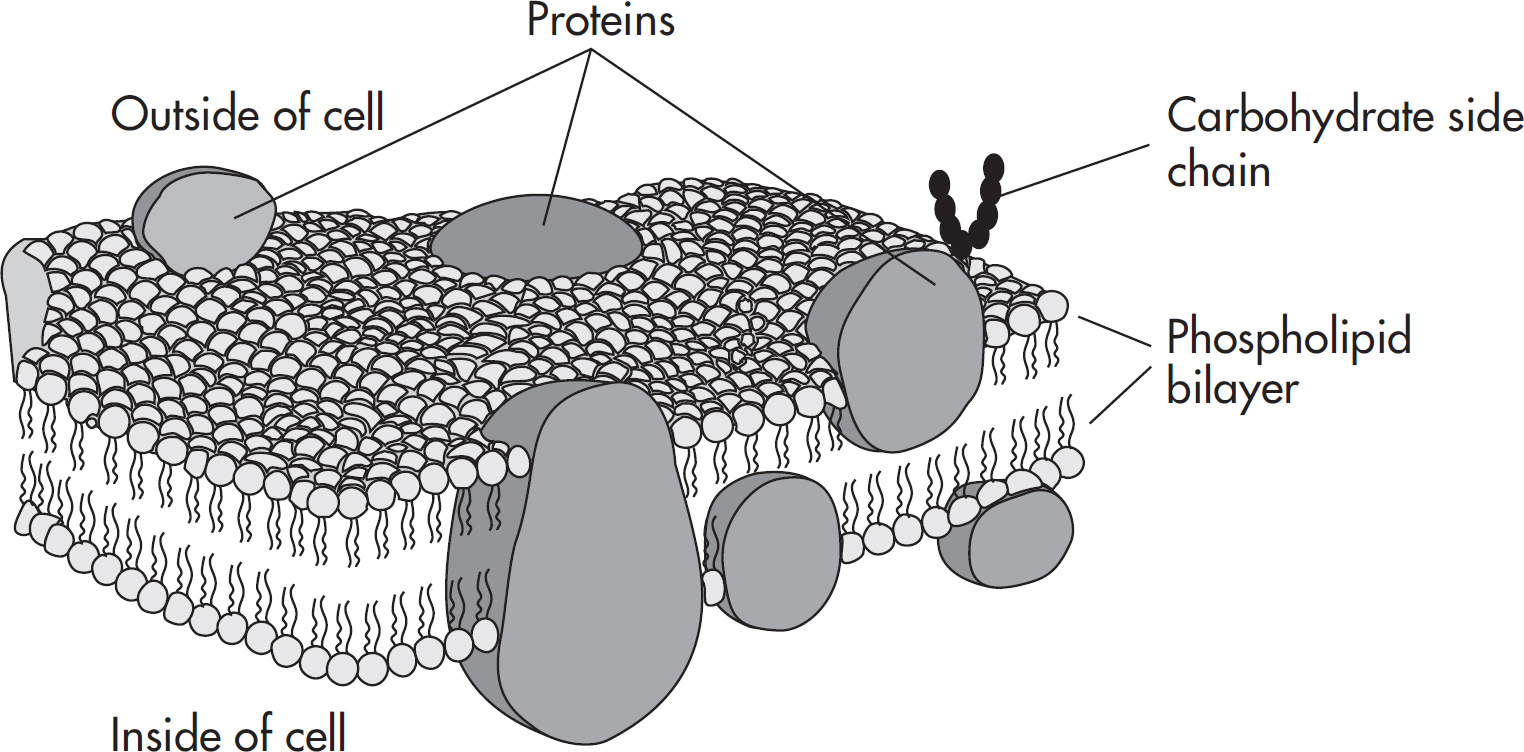
Why should the plasma membrane need so many different proteins? It’s because of the number of activities that take place in or on the membrane. Generally, plasma membrane proteins fall into several broad functional groups. Some membrane proteins form junctions between adjacent cells (adhesion proteins). Others serve as docking sites for arrivals at the cell, such as hormones (receptor proteins). Some proteins form pumps that use ATP to actively transport solutes across the membrane (transport proteins). Others form channels that selectively allow the passage of certain ions or molecules (channel proteins). Finally, some proteins, such as glycoproteins, are exposed on the extracellular surface and play a role in cell recognition and adhesion (recognition and adhesion proteins).
Attached to the surface of some proteins are carbohydrate side chain. They are found only on the outer surface of the plasma membrane. As mentioned above, cholesterol molecules are also found in the phospholipid bilayer because they help stabilize membrane fluidity in animal cells.
The Nucleus
The nucleus, which is usually the largest organelle, is the control center of the cell. The nucleus not only directs what goes on in the cell, it is also responsible for the cell’s ability to reproduce. It’s the home of the hereditary information—DNA—which is organized into large structures called chromosomes. The most visible structure within the nucleus is the nucleolus, which is where rRNA is made and ribosomes are assembled.
Ribosomes
The ribosomes are the sites of protein synthesis. Their job is to manufacture all the proteins required by the cell or secreted by the cell. Ribosomes are round structures composed of two subunits. The structure is composed of RNA and proteins. Ribosomes can be either free floating in the cell or attached to another structure called the endoplasmic reticulum (ER).
Endoplasmic Reticulum (ER)
The endoplasmic reticulum (ER) is a continuous channel that extends into many regions of the cytoplasm. The region of the ER that is attached to the nucleus and “studded” with ribosomes is called the rough ER (RER). Proteins generated in the rough ER are trafficked to or across the plasma membrane, or they are used to build Golgi bodies, lysosomes, or the ER. The region of the ER that lacks ribosomes is called the smooth ER (SER). The smooth ER makes lipids, hormones, and steroids and breaks down toxic chemicals.
Golgi Bodies
The Golgi bodies, which look like stacks of flattened sacs, also participate in the processing of proteins. Once the ribosomes on the rough ER have completed synthesizing proteins, the Golgi bodies modify, process, and sort the products. They’re the packaging and distribution centers for materials destined to be sent out of the cell. They package the final products in little sacs called vesicles, which carry the products to the plasma membrane. Golgi bodies are also involved in the production of lysosomes.
Mitochondria
Another important organelle is the mitochondrion. The mitochondria are often referred to as the “powerhouses” of the cell. They’re power stations responsible for converting the energy from organic molecules into useful energy for the cell. The energy molecule in the cell is adenosine triphosphate (ATP).
The mitochondrion is usually an easy organelle to recognize because it has a unique oblong shape and a characteristic double membrane consisting of an inner portion and an outer portion. The inner mitochondrial membrane forms folds known as cristae and separates the innermost area called the matrix from the intermembrane space. The outer membrane separates the intermembrane space from the cytoplasm. As we’ll see later, most of the production of ATP is done on the cristae.
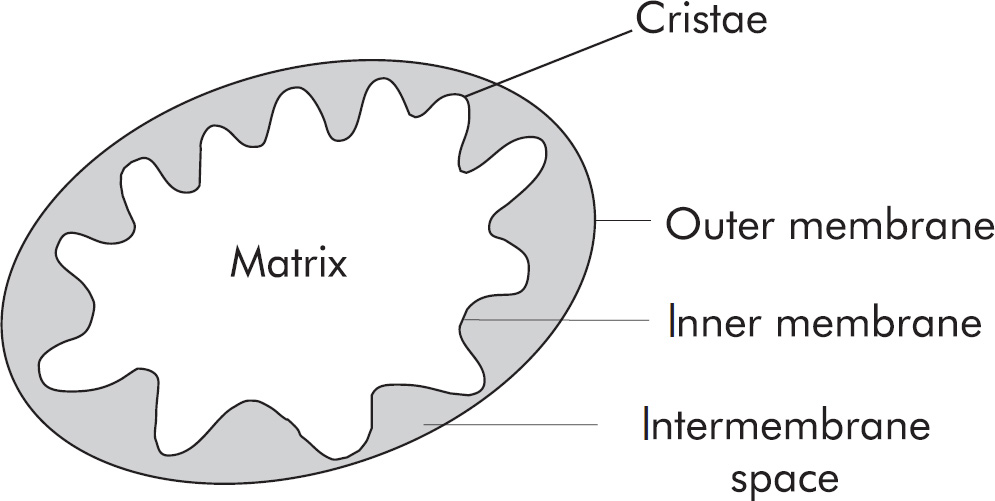
Lysosomes
Throughout the cell are small, membrane-bound structures called lysosomes. These tiny sacs carry digestive enzymes, which they use to break down old, worn-out organelles, debris, or large ingested particles. Lysosomes make up the cell’s clean up crew, helping to keep the cytoplasm clear of unwanted flotsam. Lysosomes contain hydrolytic enzymes that only function at acidic pH, which is enclosed inside the lumen of the lysosome.
Centrioles
The centrioles are small, paired, cylindrical structures that are often found within microtubule organizing centers (MTOCs). Centrioles are most active during cellular division. When a cell is ready to divide, the centrioles produce microtubules, which pull the replicated chromosomes apart and move them to opposite ends of the cell. Although centrioles are common in animal cells, they are not found in most types of plant cells.
Vacuoles
In Latin, the term vacuole means “empty cavity.” But vacuoles are far from empty. They are fluid-filled sacs that store water, food, wastes, salts, or pigments.
Peroxisomes
Peroxisomes are organelles that detoxify various substances, producing hydrogen peroxide (H2O2) as a byproduct. They also contain enzymes that break down hydrogen peroxide into oxygen and water. In animals, they are common in the liver and kidney cells.
Cytoskeleton
Have you ever wondered what actually holds the cell together and enables it to keep its shape? The shape of a cell is determined by a network of fibers called the cytoskeleton. The most important fibers you’ll need to know are microtubules and microfilaments.
Microtubules, which are made up of the protein tubulin, participate in cellular division and movement. These small fibers are an integral part of three structures: centrioles, cilia, and flagella. We’ve already mentioned that centrioles help chromosomes separate during cell division. Cilia and flagella are threadlike structures best known for their locomotive properties in single-celled organisms. The beating motion of cilia and flagella structures propels these organisms through their watery environments.
The two classic examples of organisms with these structures are the Euglena, which gets about using its whiplike flagellum, and the Paramecium, which is covered in cilia. The rhythmic beating of the Paramecium’s cilia enables it to motor about in waterways, ponds, and microscope slides in your biology lab. You’ve probably already checked these out in lab, but here’s what they look like.
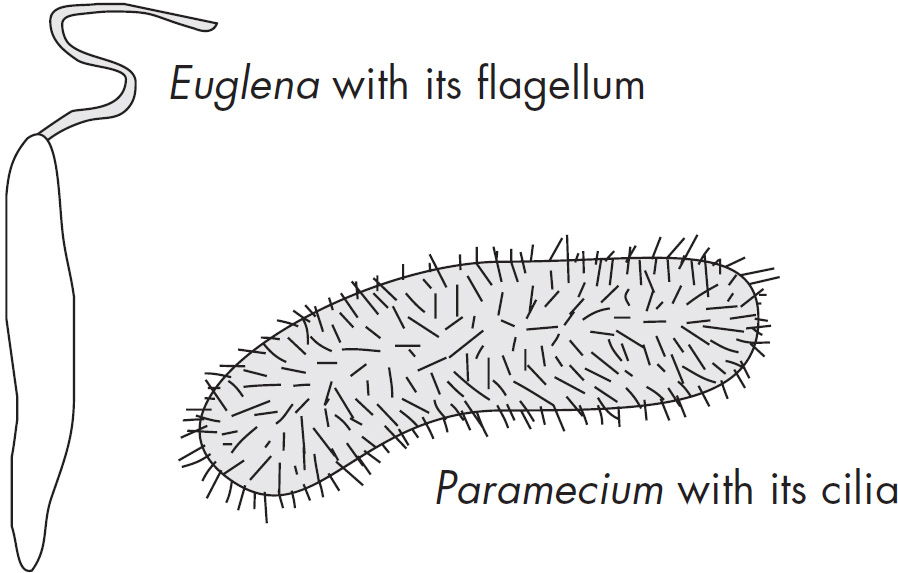
Though we usually associate such structures with microscopic organisms, they aren’t the only ones with cilia and flagella. As you probably know, these structures are also found in certain human cells. For example, the cells lining your respiratory tract possess cilia that sweep constantly back and forth (beating up to twenty times per second), helping to keep dust and unwanted debris from descending into your lungs. And every sperm cell has a type of flagellum, which enables it to swim through the female reproductive organs to fertilize the waiting ovum.
Microfilaments, like microtubules, are important for movement. These thin, rodlike structures are composed of the protein actin. Actin monomers are joined together and broken apart as needed to allow microfilaments to grow and shrink. Microfilaments assist during cytokinesis, muscle contraction, and formation of pseudopodia extensions during cell movement.
Plant Cells Versus Animal Cells
Plant cells contain most of the same organelles and structures seen in animal cells, with several key exceptions. Plant cells, unlike animal cells, have a protective outer covering called the cell wall (made of cellulose). A cell wall is a rigid layer just outside of the plasma membrane that provides support for the cell. It is found in plants, protists, fungi, and bacteria. (In fungi, the cell wall is usually made of chitin, a modified polysaccharide. Chitin is also a principle component of an arthropod’s exoskeleton.) In addition, plant cells possess chloroplasts (organelles involved in photosynthesis). Chloroplasts contain chlorophyll, the light-capturing pigment that gives plants their characteristic green color. Another difference between plant and animal cells is that most of the cytoplasm within a plant cell is usually taken up by a large vacuole that crowds the other organelles. In mature plants, this vacuole contains the cell sap. A full vacuole in a plant is a sign that it is not dehydrated. Dehydrated plants cannot fill their vacuoles and they wilt. Plant cells also differ from animal cells in that plant cells do not contain centrioles.
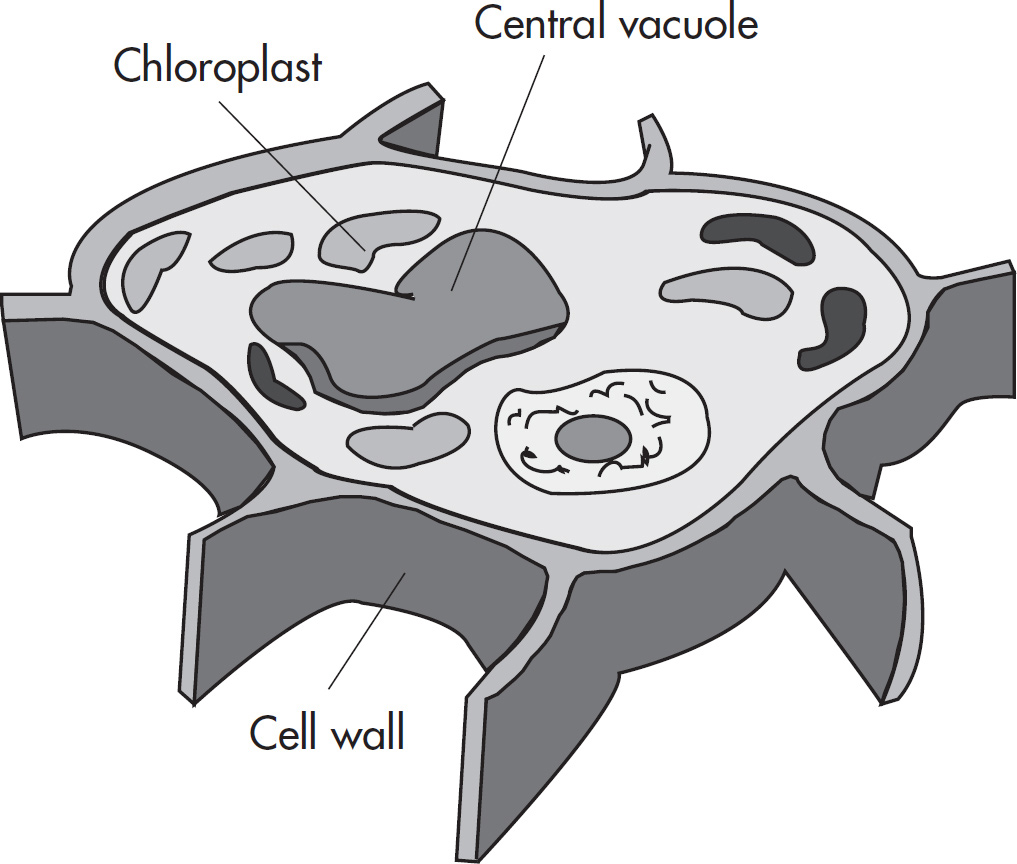
To help you remember the differences among prokaryotes, plant cells, and animal cells, we’ve put together this simple table. Make sure you learn it! The testing board is bound to ask you which cells contain which structures:
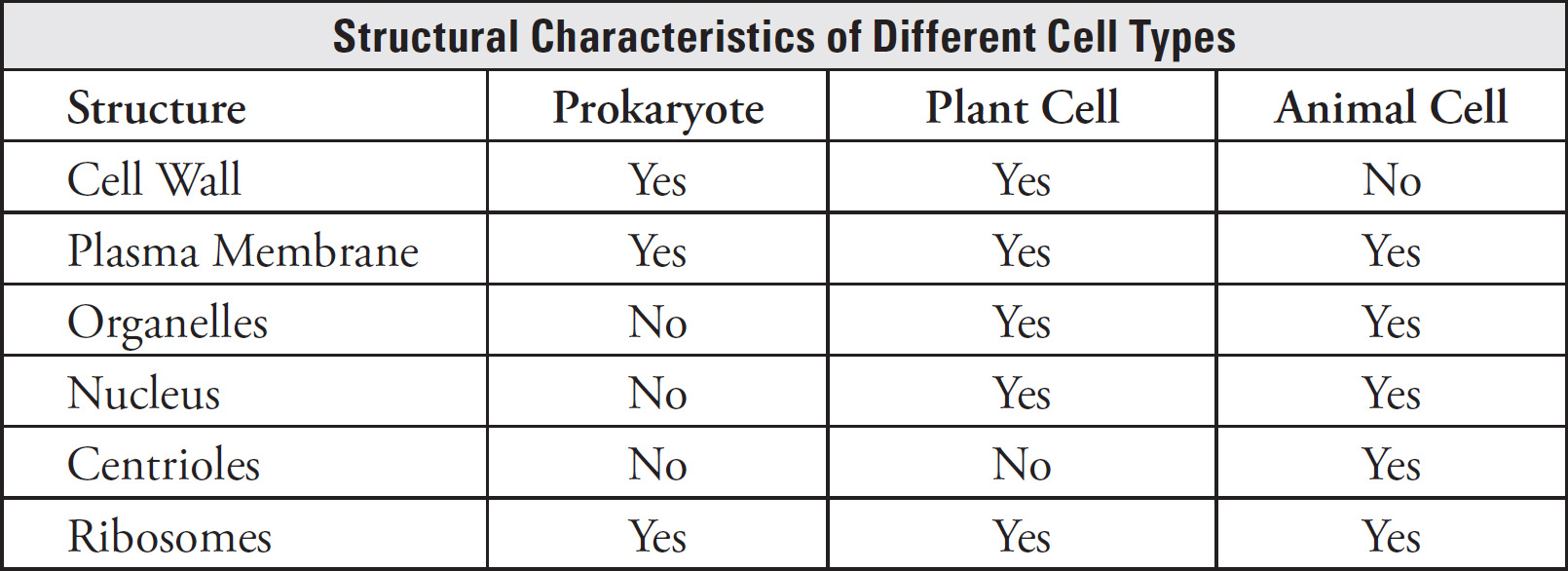
Why do we need to know about the structure of cells? Because biological structure is often closely related to function. (Watch out for this connection: it’s a favorite theme for the AP Biology Exam.) And, more importantly, because the testing board likes to test you on it!
Transport: The Traffic Across Membranes
We’ve talked about the structure of cell membranes; now let’s discuss how molecules and fluids pass through the plasma membrane. What are some of the patterns of membrane transport? The ability of molecules to move across the cell membrane depends on two things: (1) the semipermeability of the plasma membrane and (2) the size and charge of particles that want to get through.
Since the plasma membrane is composed primarily of phospholipids, lipid-soluble substances cross the membrane without any resistance. Why? Because “like dissolves like.” The lipid bilayer is a hydrophobic sandwich, and only hydrophobic things can pass that zone. If a substance is hydrophilic, the bilayer won’t let it pass without assistance, called facilitated transport.
Facilitated transport depends upon a number of proteins that act as tunnels through the membrane. Channels are very specialized types of tunnels that only let certain things through.
The most famous of these are aquaporins, which are water-specific channels. Although water is polar, there are typically sufficient aquaporins for water to traverse the membrane whenever it wishes without forming traffic jams. However, without aquaporins, no water would be able to cross the membrane.
Passive Transport: Simple and Facilitated Diffusion
Now we know how things cross the membrane, but let’s talk about why they cross the membrane. The simple answer is: “To get to the other side.” If there is a high concentration of something in one area, it will move to spread out and diffuse into an area with a lower concentration, even if that means entering or exiting a cell. In other words, the substance moves down a concentration gradient. This is called diffusion. When the molecule that is diffusing is hydrophobic, the diffusion is called simple diffusion. When the diffusion requires the help of a channel-type protein, it is called facilitated diffusion. Anytime that a substance is moving by diffusion, it is called passive transport because there is no outside energy required to cause the movement. It’s like riding a bicycle downhill. The molecules are just “going with the flow.”
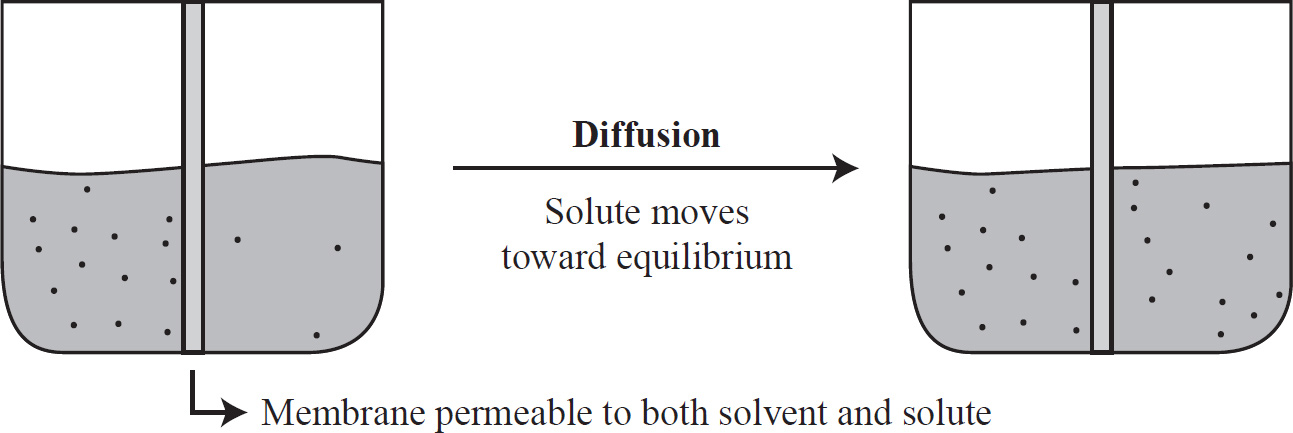
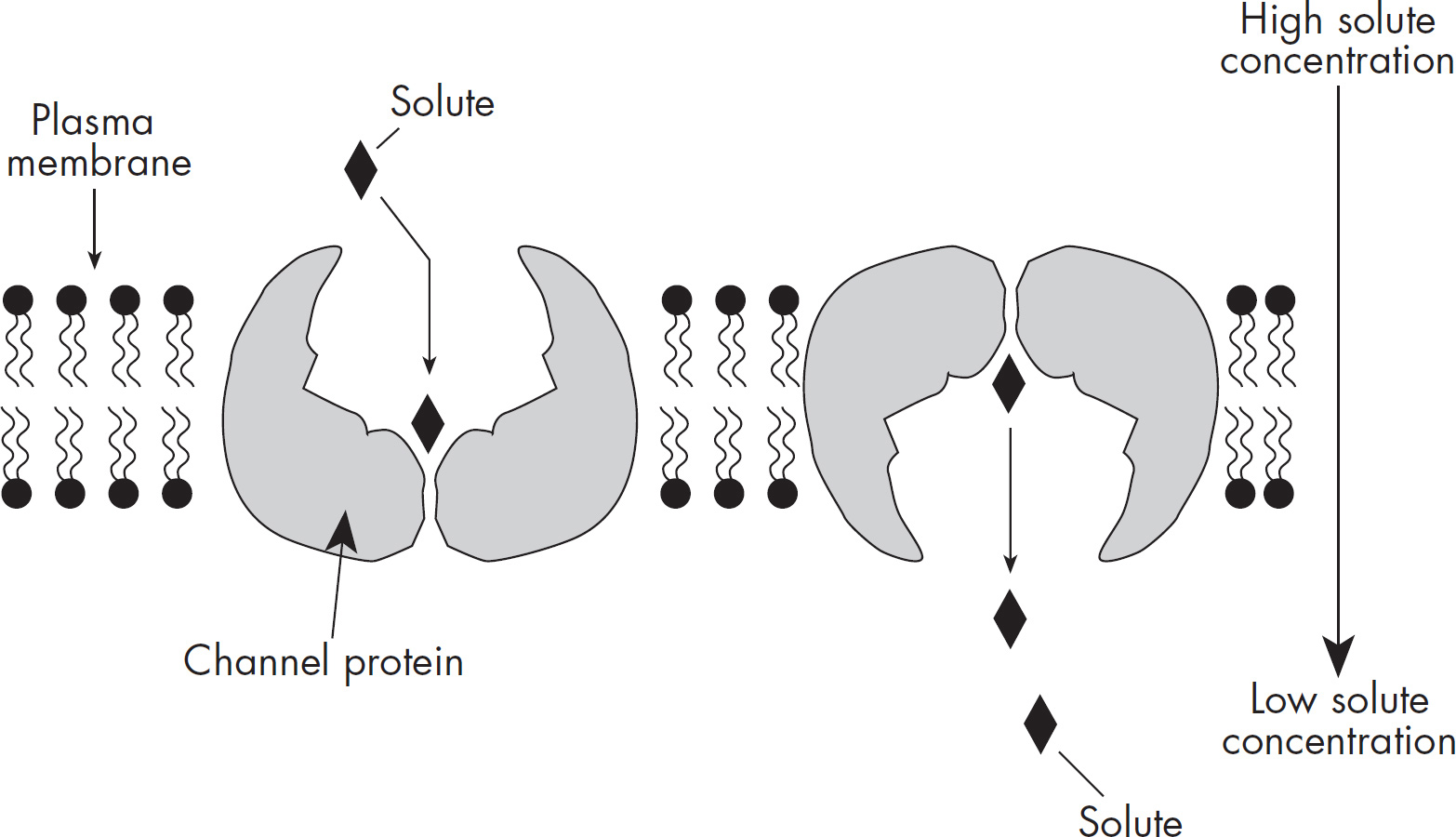
Osmosis
When the molecule that is diffusing is water, the process is called osmosis. Water always wants to move from an area where it is most concentrated to an area where it is least concentrated. Water is not usually pure water, but since it is so great at dissolving things, it usually exists as a solution. A solution is made when a liquid solvent dissolves solute particles. So, if we rethink water moving down its concentration gradient, this is the same as saying that it wants to go from where there is less solute (ie. watery solution) to where there is more solute (ie. concentrated solution). It’s as if the water is moving to dilute the concentrated solute particles.
Often, situations discussing osmosis set it up so that there are two areas and the water can flow freely, but the solute cannot. For example, if a chamber containing water and a chamber containing a sucrose solution are connected by a semipermeable membrane that allows water but not sucrose to cross, diffusion of sucrose between the chambers cannot occur. In this case, osmosis draws water into the sucrose chamber to reduce the sucrose concentration. This will reduce the total volume of the water chamber. Water will flow into the sucrose chamber until the concentration is the same across the membrane.
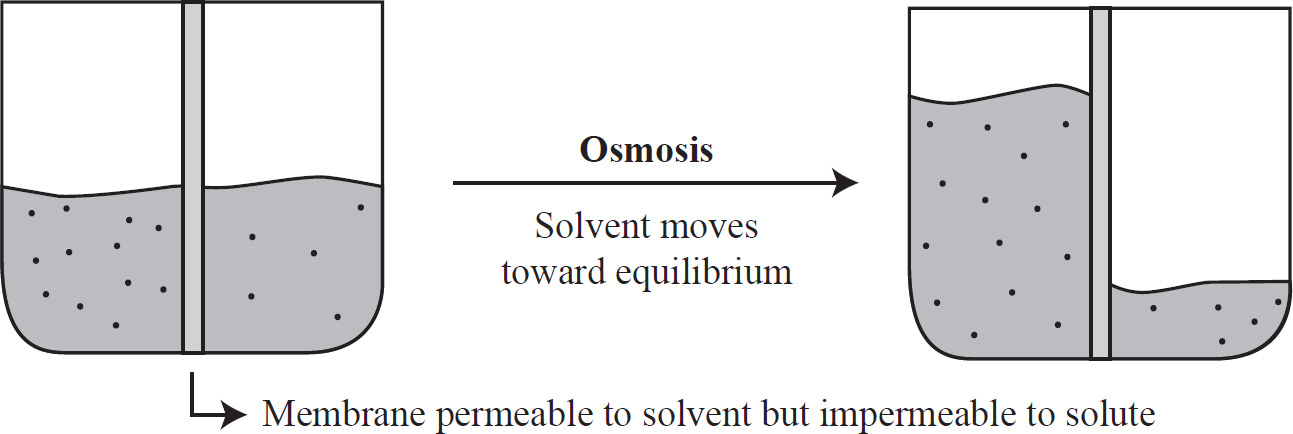
In both diffusion and osmosis, the final result is that solute concentrations are the same on both sides of the membrane. The only difference is that in diffusion the membrane is permeable to solute, and in osmosis it is not.
The term tonicity is used to describe osmotic gradients. If the environment is isotonic to the cell, the solute concentration is the same inside and outside. A hypertonic solution has more total dissolved solutes than the cell, while a hypotonic solution has less. The tendency of water to move down its concentration gradient (into cells) can be a powerful force; it can cause cells to explode and can overcome gravity. You may also hear the terms isosmotic, hyperosmotic, and hyposmotic. These words are usually used when comparing two solutions, while tonicity terms are used when comparing a solution to a cell. Other than this, the definitions are similar; isotonic and isosmotic both describe situations where the concentration is the same on either side of a membrane, for example. The only difference is what type of membrane.
Water potential (Ψ) is the measure of potential energy in water and describes the eagerness of water to flow from an area of high water potential to an area of low water potential. It is affected by two factors: pressure potential (Ψp) and solute potential (Ψs). The equation for solute (or osmotic) potential is listed on the AP Biology Equations and Formulas sheet and describes the effect of solute concentration on water flow. Adding a solute lowers the water potential of a solution, causing water to be less likely to leave this solution and more likely to flow into it. This makes sense because the added solute makes the solution more concentrated, and the water is now unlikely to diffuse away. The more solute molecules present, the more negative the solute potential is.
A red blood cell dropped into a hypotonic solution (such as distilled water) will expand because water will move into the cell, an area of lower water potential. Eventually, the red blood cell will pop. If a similar experiment is done with a plant cell, water will still move into the cell, but the cell wall will exert pressure, increasing the water potential and limiting the gain of water. Water potential also explains how water moves from soil into plant roots and how plants transport water from roots to leaves to support photosynthesis.
The concentration of a solution can be calculated by dividing the number of moles of solute by the volume (in liters) of solution. Highly concentrated (or stock) solutions can be diluted to make less concentrated solutions. The equation to do this is on the AP Biology Equations and Formulas sheet.
CiVi = CfVf
This equation tells you how much (Vi) of a concentrated solution (at Ci) is required to make a more dilute solution (Cf) of a certain volume (Vf). The amount of solvent added will be the final volume of the solution (Vf) minus whatever amount of stock solution you need to add. For example, if you have a bottle of 2 M solution of Tris buffer and would like to make 50 mL of 0.1M Tris.
CiVi = CfVf
(2 M)(Vi) = (0.1 M)(0.050 L)
Vi = 0.0025 L, or 2.5 mL
Thus, you would add 2.5 mL of the concentrated (or stock) Tris solution to a new tube or bottle and then add water (the solvent) up to a final volume of 50 mL. In other words, you should add 47.5 mL of water.
Active Transport
Suppose a substance wants to move in the opposite direction—from a region of lower concentration to a region of higher concentration. A transport protein can help usher the substance across the plasma membrane, but it’s going to need energy to accomplish this. This time it’s like riding a bicycle uphill. Compared with riding downhill, riding uphill takes a lot more work. Movement against the natural flow is called active transport.
But where does the protein get this energy? Some proteins in the plasma membrane are powered by ATP. The best example of active transport is a special protein called the sodium-potassium pump. It ushers out three sodium ions (Na+) and brings in two potassium ions (K+) across the cell membrane. These pumps depend on ATP to get ions across that would otherwise remain in regions of higher concentration. Primary active transport occurs when ATP is directly utilized to transport something. Secondary active transport occurs when something is actively transported using the energy captured from the movement of another substance flowing down its concentration gradient.
We’ve now seen that small substances can cross the cell membrane by:
• simple diffusion
• facilitated transport
• active transport
Endocytosis
When the particles that want to enter a cell are just too large, the cell uses a portion of the cell membrane to engulf the substance. The cell membrane forms a pocket, pinches in, and eventually forms either a vacuole or a vesicle. This is called endocytosis.
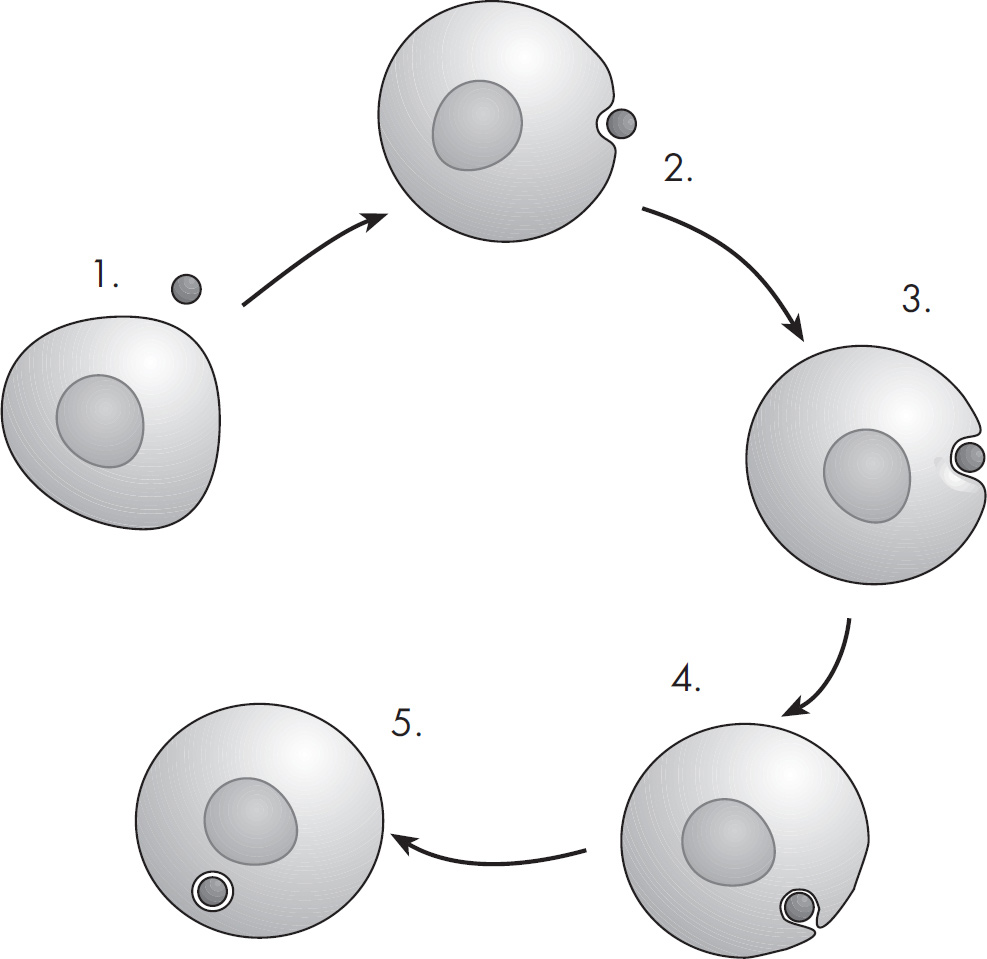
Three types of endocytosis exist: pinocytosis, phagocytosis, and receptor- mediated endocytosis. In pinocytosis, the cell ingests liquids (“cell-drinking”). In phagocytosis, the cell takes in solids (“cell-eating”). A special type of endocytosis, receptor-mediated endocytosis, involves cell surface receptors that work in tandem with a endocytic pits that are lined with a protein called clathrin. When a particle, or ligand, binds to one of these receptors, the ligand is brought into the cell by the invagination or “folding in” of the cell membrane. A vesicle then forms around the incoming ligand and carries it into the cell’s interior.
Bulk Flow
Other substances move by bulk flow. Bulk flow is the one-way movement of fluids brought about by pressure. For instance, the movement of blood through a blood vessel or movement of fluids in xylem and phloem of plants are examples of bulk flow.
Dialysis
Dialysis is the diffusion of solutes across a selectively permeable membrane. Special membranes that have holes of a certain size within them can be used to sort substances by using the process of diffusion. Kidney dialysis is a specialized process by which the blood is filtered using machines and concentration gradients. Things present at high levels will naturally diffuse out of the blood; dialysis just gives them the opportunity.
Exocytosis
Sometimes large particles are transported out of the cell. In exocytosis, a cell ejects waste products or specific secretion products, such as hormones, by the fusion of a vesicle with the plasma membrane. Think of it as reverse endocytosis.
Cell Junctions
When cells come in close contact with each other, they develop specialized intercellular junctions that involve their plasma membranes as well as other components. These structures may allow neighboring cells to form strong connections with each other, prevent passage of materials, or establish rapid communication between adjacent cells. There are three types of intercellular contacts in animal cells: desmosomes, gap junctions, and tight junctions.
Desmosomes hold adjacent animal cells tightly to each other, like a rivet. They consist of a pair of discs associated with the plasma membrane of adjacent cells, plus the intercellular protein filaments that cross the small space between them. Intermediate filaments within the cells are also attached to the discs (see figure on the next page).
Gap junctions are protein complexes that form channels in membranes and allow communication between the cytoplasm of adjacent animal cells for the transfer of small molecules and ions. Gap junctions are found in the heart and in compact bone and smooth muscle.
Tight junctions are tight connections between the membranes of adjacent animal cells. They’re so tight that there is no space between the cells. Cells connected by tight junctions seal off body cavities and prevent leaks. Tight junctions are found in the small intestine.
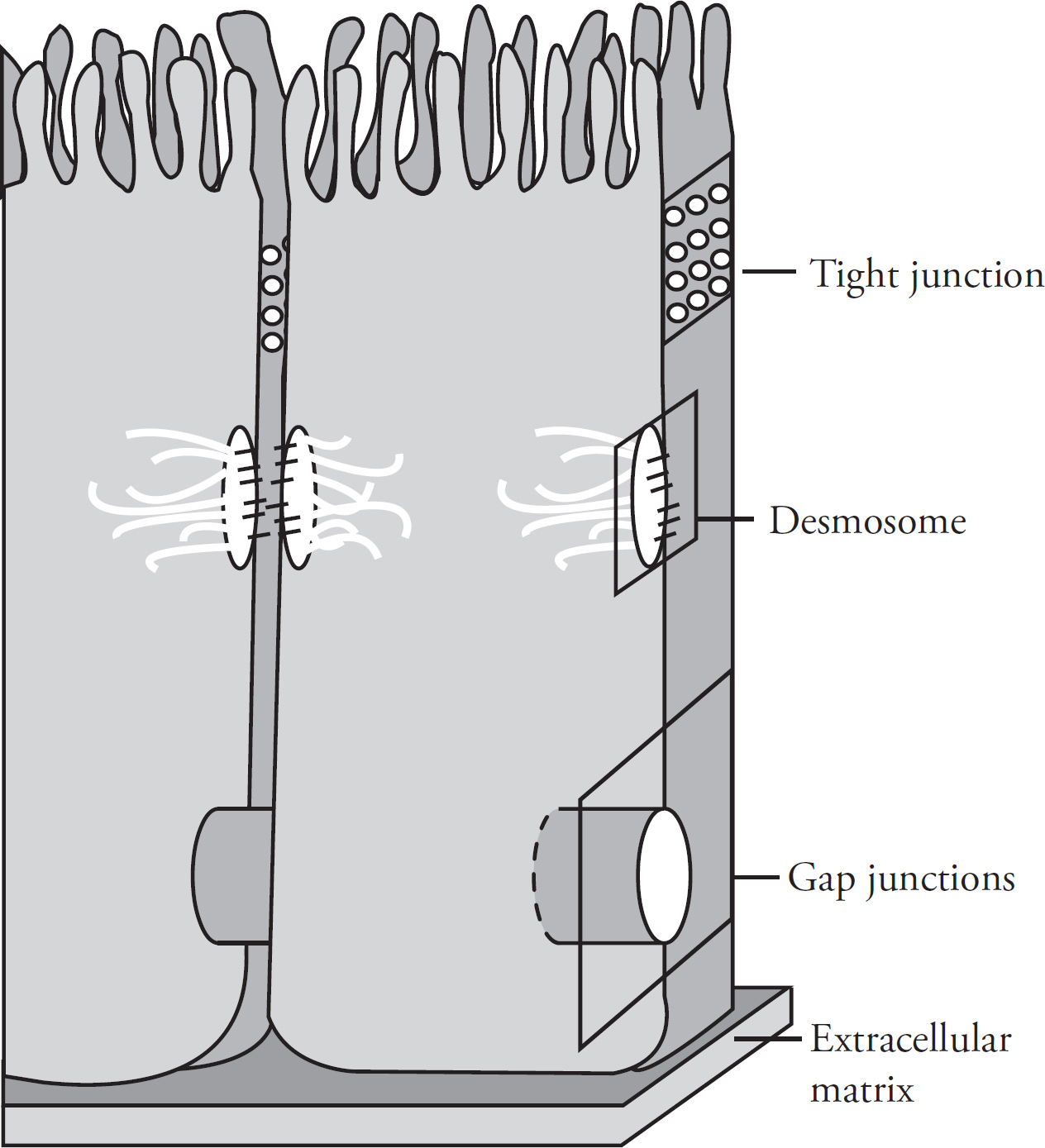
Cell Communication
Communication is important for organisms and involves transduction of stimulatory or inhibitory signals from other cells, organisms, or the environment. Unicellular organisms detect and respond to environmental signals. For example, they can use cell communication to locate a suitable mate, adapt metabolism rates in response to changes in nutrient availability, or make their numbers known to other members of their species (a phenomenon known as quorum sensing).
Taxis is the movement of an organism in response to a stimulus and can be positive (towards the stimulus) or negative (away from the stimulus). Taxes are innate behavioral responses, or instincts. Chemotaxis is movement in response to chemicals. For example, bacteria can control flagella rotation to direct their motion, thus avoiding repellents (such as poisons), or helping them find favorable locations with high concentrations of attractants (such as food). In our bodies, neutrophils use chemotaxis to respond to an infection and are the first responders to inflammation.
The cells of multi-celled organisms must communicate with one another to coordinate the activities of the organism as a whole. Signaling can be short-range (affecting only nearby cells) or long-range (affecting cells throughout the organism). It can be done by cell junctions or signaling molecules called ligands that bind to receptors and trigger a response. Hormones and neurotransmitters are common signaling molecules, and they will be covered in Chapter 11.
Signal transduction is the process by which an external signal is transmitted to the inside of a cell. It usually involves the following three steps:
1. a signaling molecule binding to a specific receptor
2. activation of a signal transduction pathway
3. production of a cellular response
Hydrophobic signaling molecules, like cholesterol-based steroid hormones, can diffuse across the plasma membrane. These molecules can bind receptors inside the cell and often travel to the nucleus where they regulate gene expression.
For signaling molecules that cannot enter the cell, a plasma membrane receptor is required. Plasma membrane receptors form an important class of integral membrane proteins that transmit signals from the extracellular space into the cytoplasm. Each receptor binds a particular molecule in a highly specific way. There are three classes of membrane receptors.
1. Ligand-gated ion channels in the plasma membrane open an ion channel upon binding a particular ligand. An example is the ligand-gated sodium channel on the surface of a skeletal muscle cell at the neuromuscular junction. This channel opens in response to acetylcholine, and a massive influx of sodium depolarizes the muscle cell and causes it to contract.
2. Catalytic (or enzyme-linked) receptors have an enzymatic active site on the cytoplasmic side of the membrane. Enzyme activity is initiated by ligand binding at the extracellular surface. The insulin receptor is an example of an enzyme-linked receptor. After binding insulin, enzymatic activity initiates a complex signaling pathway that allows the cell to grow, synthesize lipids, and import glucose.
3. A G-protein-linked receptor does not act as an enzyme, but instead will bind different version of a G-protein (often GTP or GDP) on the intracellular side when a ligand is bound extracellularly. This causes activation of secondary messengers within the cell. One important second messenger is cyclic AMP (cAMP). It is known as a “universal hunger signal” because it is the second messenger of the hormones epinephrine and glucagon, which cause energy mobilization. Secondary messengers are usually small molecules that can move quickly through the cell. They can be made and destroyed quickly and help the signal amplify throughout the cell.
Signal transduction in eukaryotic cells usually involves many steps and complex regulation. In contrast, bacterial cells usually use a simpler two-component regulatory system in transduction pathways.
CELL COMMUNICATION IN PLANTS
Plants do not have a nervous system but can produce several proteins found in animal nervous systems, such as certain neurotransmitter receptors. Plants can also generate electrical signals in response to environmental stimuli, and this can affect flowering, respiration, photosynthesis, and wound healing. Light receptors are common in plants and help link environmental cues to biological processes such as seed germination, the timing of flowering, and chlorophyll production. Some plants can also use chemicals to communicate with nearby plants. For example, wounded tomatoes produce a volatile chemical as an alarm signal. This warns nearby plants and allows them to prepare a defense or immune response.
TROPISMS
Plants need light. Notice that all the plants in your house tip toward the windows. This movement toward the light is known as phototropism. As you also know, plants generally grow up and down: the branches grow upward, while the roots grow downward into the soil, seeking water. This tendency to grow toward or away from the earth is called gravitropism. All of these tropisms are examples of behavior in plants.
A tropism is a turning response to a stimulus. Tropisms are different from taxes, which are changes in motility. Remember, plants are immobile.
There are four basic tropisms in plants. They’re easy to remember because their prefixes indicate the stimuli to which plants react.
1. Phototropism refers to how plants respond to sunlight. For example, plants always bend toward light.
2. Gravitropism refers to how plants respond to gravity. Stems exhibit negative gravitropism (i.e., they grow up, away from the pull of gravity), whereas roots exhibit positive gravitropism (i.e., they grow downward into the earth).
3. Thigmotropism refers to how plants respond to touch. For example, ivy grows around a post or trellis.
4. Photoperiodism refers to how plants respond to the light/dark cycles and the seasonal changes in the length of day. Animals also often respond to this.
KEY TERMS
cells
light microscopes
electron microscopes
eukaryotic cells
prokaryotic cells
cytoplasm
nucleoid
plasma membrane
flagella
organelles
phosholipid bilayer
peripheral proteins
integral proteins
transmembrane proteins
fluid-mosaic model
adhesion proteins
receptor proteins
transport proteins
channel proteins
recognition and adhesion proteins
carbohydrate side chains
chromosomes
nucleolus
ribosomes
endoplasmic reticulum (ER)
rough ER
smooth ER
Golgi bodies
vesicles
mitochondria
adenosine triphosphate (ATP)
lysosomes
centrioles
microtubule organizing centers (MTOCs)
vacuoles
peroxisomes
cytoskeleton
microtubules
microfilaments
tubulin
cilia
Euglena
Paramecium
cell wall
chitin
chloroplasts
cell sap
facilitated transport
aquaporins
simple diffusion
facilitated diffusion
passive transport
simple diffusion (or passive transport)
osmosis
tonicity
isotonic
hypertonic
hypotonic
water potential
solute potential
solutes
channel proteins
active transport
sodium-potassium pump
endocytosis
pinocytosis
phagocytosis
receptor-mediated endocytosis
bulk flow
dialysis
exocytosis
intercellular junctions
desmosomes
gap junctions
tight junctions
ligands
receptors
signal transduction
ligand-gated ion channel
catalytic receptor
G-protein-linked receptor
phototropism
gravitropism
thigmotropism
photoperiodism
Summary
 Living things are composed of cells, the smallest unit of living organisms. Cells may be categorized into prokaryotes, which do not have a nucleus or membrane bound organelles, and eukaryotes, which have a nucleus and membrane-bound organelles. Features of prokaryotic cells (like bacteria, for example) can include the following:
Living things are composed of cells, the smallest unit of living organisms. Cells may be categorized into prokaryotes, which do not have a nucleus or membrane bound organelles, and eukaryotes, which have a nucleus and membrane-bound organelles. Features of prokaryotic cells (like bacteria, for example) can include the following:
• plasma membrane
• flagella and cilia
• cell wall
• nucleoid region
• ribosomes
• circular chromosome
• cytoplasm
 Components of the eukaryotic cells include the following:
Components of the eukaryotic cells include the following:
• nucleus
• nucleolus
• rough endoplasmic reticulum
• smooth endoplasmic reticulum
• Golgi body
• ribosomes
• mitochondria
• vacuole
• lysosome
• peroxisomes
• centrioles (animals only)
• cytoplasm
• plasma membrane
• cytoskeleton (microtubules, microfilaments)
• central vacuole (plants only)
• cell wall (plants only)
 The plasma membrane is composed of a phospholipid bilayer. The hydrophobic nature of this membrane is responsible for its selective permeability. Transport through the membrane is dependent on size and polarity of the molecules and concentration gradients. There are a few different types of cellular transport:
The plasma membrane is composed of a phospholipid bilayer. The hydrophobic nature of this membrane is responsible for its selective permeability. Transport through the membrane is dependent on size and polarity of the molecules and concentration gradients. There are a few different types of cellular transport:
• simple diffusion
• facilitated transport
• active transport
• bulk flow
• dialysis
• exocytosis
• endocytosis (pinocytosis, phagocytosis, receptor-mediated endocytosis)
 Cells are connected by tight junctions, desmosomes, and gap junctions, which all function to connect adjacent cells.
Cells are connected by tight junctions, desmosomes, and gap junctions, which all function to connect adjacent cells.
 Sometimes cells do not need to pass any molecules through the membrane, but instead they just pass a message. This is signal transduction. A ligand binds to a receptor on the outside of the cell and causes changes to the inside of the cell. Ligand-gated ion channels, catalytic receptors, and G-protein-linked receptors are common examples.
Sometimes cells do not need to pass any molecules through the membrane, but instead they just pass a message. This is signal transduction. A ligand binds to a receptor on the outside of the cell and causes changes to the inside of the cell. Ligand-gated ion channels, catalytic receptors, and G-protein-linked receptors are common examples.
 Response to signals and stimuli is important for all living things. In plants, phototropism, gravitropism, thigmotropism, and photoperiodism are all ways that plants grow according to environmental signals.
Response to signals and stimuli is important for all living things. In plants, phototropism, gravitropism, thigmotropism, and photoperiodism are all ways that plants grow according to environmental signals.
Chapter 5 Drill
Answers and explanations can be found in Chapter 16.
1. Movement of substances into the cell is largely dependent on the size, polarity, and concentration gradient of the substance. Which of the following represents an example of active transport of a substance into a cell?
(A) Diffusion of oxygen into erythrocytes (red blood cells) in the alveolar capillaries of the lungs
(B) Influx of sodium ions through a voltage-gated ion channel in a neuron cell during an action potential
(C) The sodium-potassium pump, which restores resting membrane potentials in neurons through the use of ATP
(D) Osmosis of water into an epithelial cell lining the lumen of the small intestine
2. The development of electron microscopy has provided key insights into many aspects of cellular structure and function, which had previously been too small to be seen. All of the following would require the use of electron microscopy for visualization EXCEPT
(A) the structure of a bacteriophage
(B) the matrix structure of a mitochondrion
(C) the shape and arrangement of bacterial cells
(D) the pores on the nuclear membrane
3. Vibrio cholerae (shown below) are highly pathogenic bacteria that are associated with severe gastrointestinal illness and are the causative agent of cholera. In extreme cases, antibiotics are prescribed that target bacterial structures that are absent in animal cells. Which of the following structures is most likely targeted by antibiotic treatment?
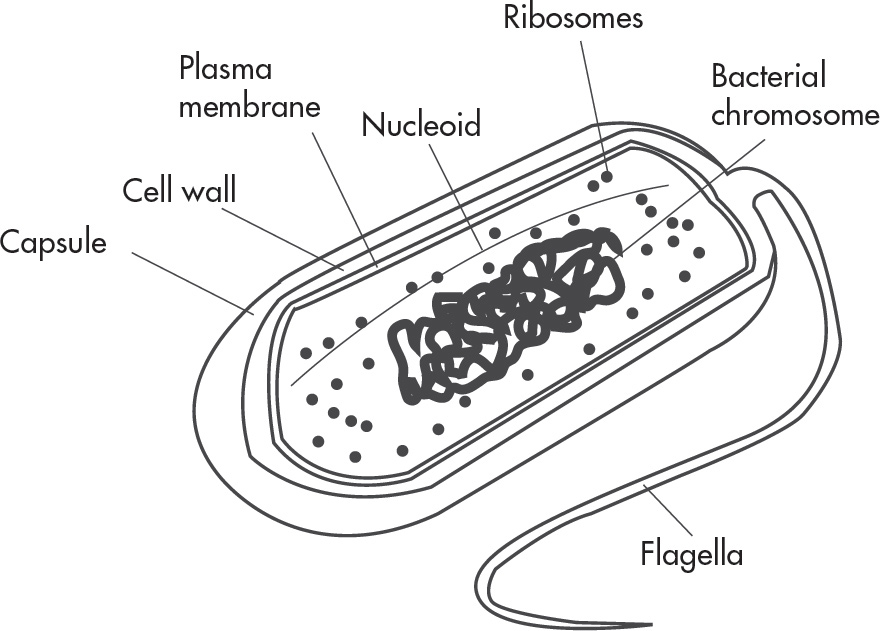
(A) Cytoplasm
(B) Plasma membrane
(C) Ribosomes
(D) Cell wall
Question 4 is based on the information and table below.
A new unicellular organism has recently been identified living in thermal pools in Yellowstone National Park. The thermal pools have average temperatures of 45°C, a pH of approximately 3.2, and high concentrations of sulfur-containing compounds. To identify the organism, a microbiologist performs a series of tests to evaluate its structural organization. The table below summarizes the microscopy data of the newly identified organism.
| Cellular Structure | Analysis |
| Plasma Membrane | Present |
| Cell Wall | Present, very thick |
| Mitochondria | Absent |
| Ribosomes | Present, highly abundant |
| Flagella | Present, peritrichous organization |
4. This organism is most likely a new species of which of the following?
(A) Algae
(B) Protozoa
(C) Bacteria
(D) Fungi
Question 5 represents a question requiring a numeric answer.
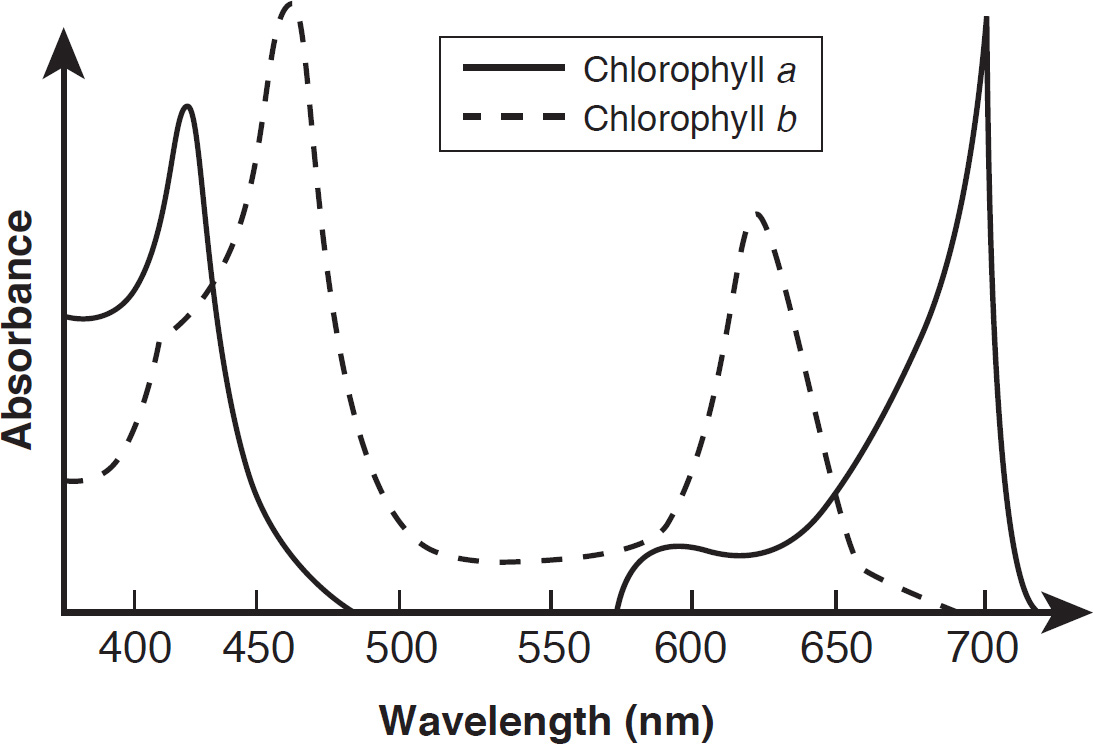
5. Chlorophyll is a green pigment present in the chloroplasts of algae and plants. It is essential for catalyzing the light-dependent cycles in photosynthesis. A scientist purifies both forms of chlorophyll (a and b) from plant chloroplasts and evaluates them for light absorption using a spectrophotometer. Using the spectrophotometer data provided above, at what wavelength is the absorbance of chlorophyll a at its maximum? Give your answer to the nearest whole number.

6. In a eukaryotic cell, which of the following organelles directly work together?
(A) Nuclear envelope, nucleolus, vacuoles, centrioles
(B) Ribosomes, rough endoplasmic reticulum, Golgi bodies, plasma membrane
(C) Mitochondria, ribosomes, lysosomes, chloroplasts
(D) Centrioles, nucleolus, smooth endoplasmic reticulum, lysosomes
7. A lipid-soluble hormone, estrogen, is secreted from the ovaries. This molecule travels through the body via the bloodstream. A researcher was interested in reducing estrogen’s effect in order to determine the response of decreased estrogen on the organism. Which of the following is a valid strategy for reducing effects of estrogen on the whole research organism?
(A) Treat with a competitive inhibitor drug that blocks all receptors at the plasma membrane
(B) Treat with lipid-soluble testosterone
(C) Treat with a lipid-soluble noncompetitive inhibitor that specifically reduces estrogen binding to the intracellular receptor
(D) Remove ovaries of the organism
8. Which type of cell would be the most useful for studying the endoplasmic reticulum?
(A) Neurons firing action potentials
(B) Insulin making cells in the pancreas
(C) Bacterial cells in a colony
(D) White blood cells
9. Paramecium is a single-celled protist that lives in freshwater habitats. In these conditions, Paramecium has evolved strategies to handle the potential consequences of inhabiting this hypotonic environment. One of these strategies could be
(A) contractile vacuoles, which expel water forcefully
(B) increased aquaporins in its cellular membrane
(C) many cilia covering its surface
(D) salt receptors on its surface to seek out less concentrated areas
10. A co-transporter is something that moves two substances across a membrane, one passively and the other actively. The Na+K+ ATPase transports sodium and potassium ions across the plasma membrane against their concentration gradients. This pump is not considered a co-transporter because
(A) ATP is produced through this transporter
(B) both ions are moved via active transport
(C) both ions are moved via passive transport
(D) ATP hydrolysis does not occur during transport
REFLECT
Respond to the following questions:
• Which topics from this chapter do you feel you have mastered?
• Which content topics from this chapter do you feel you need to study more before you can answer multiple-choice questions correctly?
• Which content topics from this chapter do you feel you need to study more before you can effectively compose a free response?
• Was there any content that you need to ask your teacher or another person about?
Chapter 6
Cellular Energetics
BIOENERGETICS
In Chapter 4, we discussed some of the more important organic molecules. But what makes these molecules so important? Glucose, starch, and fat are all energy-rich. However, the energy is packed in the chemical bonds holding the molecules together. To carry out the processes necessary for life, cells must find a way to release the energy in these bonds when they need it and store it away when they don’t. The study of how cells accomplish this is called bioenergetics. Generally, bioenergetics is the study of how energy from the sun is transformed into energy in living things.
During chemical reactions, such bonds are either broken or formed. This process involves energy, no matter in which direction we go. Every chemical reaction involves a change in energy.
ADENOSINE TRIPHOSPHATE (ATP)
We’ve all heard the expression “nothing in life is free.” The same holds true in nature. Here’s a fundamental principle of energy that it is necessary to address.
Energy cannot be created or destroyed. In other words, the sum of energy in the universe is constant.
This rule is called the first law of thermodynamics. As a result, the cell cannot take energy out of thin air. Rather, it must harvest it from somewhere.
The second law of thermodynamics states that energy transfer leads to less organization. That means the universe tends toward disorder (or entropy).
TYPES OF REACTIONS
Exergonic reactions are those in which the products have less energy than the reactants. Simply put, energy is given off during the reaction.
Let’s look at an example. The course of a reaction can be represented by an energy diagram. Here’s an energy diagram for an exergonic reaction.
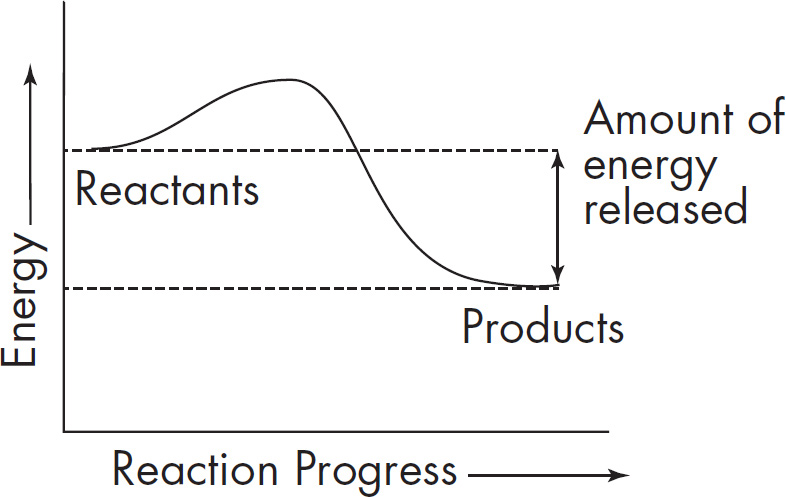
You’ll notice that energy is represented along the y-axis. Based on the diagram, we see our reaction released energy. An example of an exergonic reaction is when food is oxidized in mitochondria of cells and then releases the energy stored in the chemical bonds.
Reactions that require an input of energy are called endergonic reactions. You’ll notice that the products have more energy than the reactants.
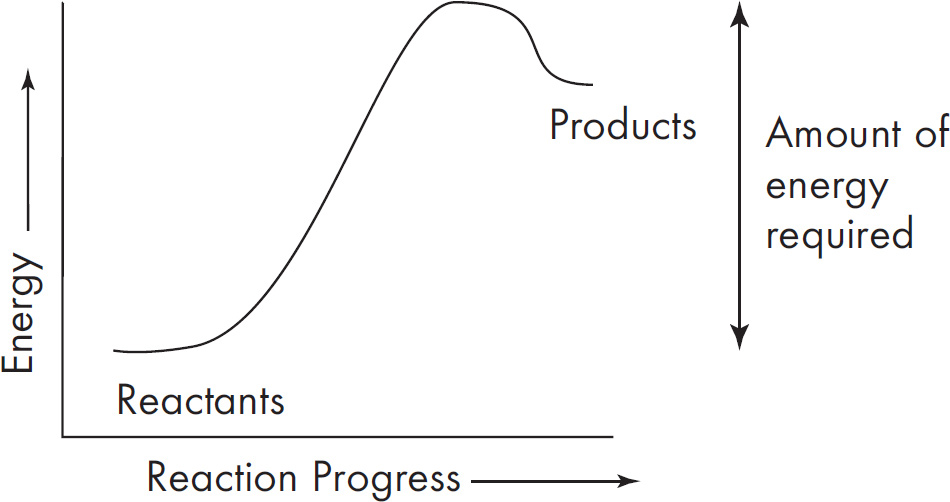
The products gained energy in the form of heat. An example is when plants use carbon dioxide and water to form sugars.
GIBBS FREE ENERGY
A practical way to discuss thermodynamics is the mathematical notion of Gibbs free energy.
∆G = ∆H – T∆S
This equation is on the AP Biology Equations and Formulas sheet. T denotes temperature, H denotes enthalpy (the measure of energy in a thermodynamic system), and S denotes entropy. The change in the Gibbs free energy of a reaction determines whether the reaction is favorable (spontaneous, negative ∆G) or unfavorable (nonspontaneous, positive ∆G). In other words, this equation can be used to figure out if the reactants will stay as they are or be converted to products.
Spontaneous reactions, ones that occur without a net addition of energy, have ∆G < 0. They occur with energy to spare. Reactions with a negative ∆G are exergonic (energy exits the system); reactions with a positive ∆G are endergonic. Endergonic reactions only occur if energy is added. In the lab, energy is added in the form of heat; in the body, endergonic reactions are driven by reaction coupling to exergonic or favorable reactions.
Activation Energy
Even though exergonic reactions release energy, the reaction might not occur naturally without a little bit of energy to get things going. This is because the reactants must first turn into an intermediate state, called the transition state, before turning into the products. The transition state is sort of a reactants-products hybrid state that is difficult to achieve. In order to reach this transition state, a certain amount of energy is required. This is called the activation energy.
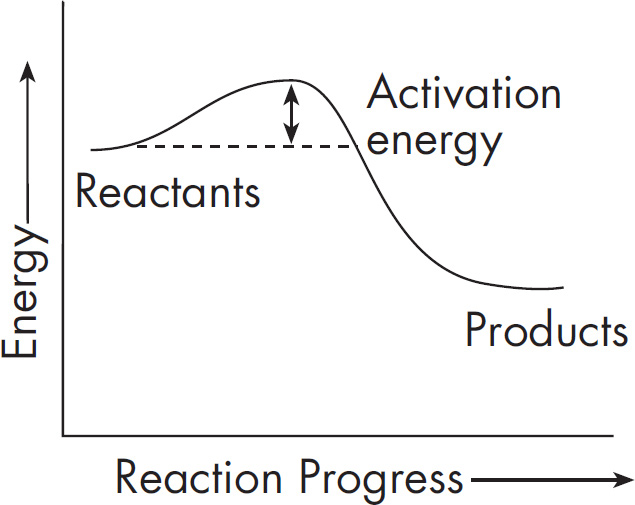
You’ll notice that we needed a little energy to get us going. That’s because chemical bonds must be broken before new bonds can form. This energy barrier—the hump in the graph—is called the activation energy. Once a set of reactants has reached its activation energy, the reaction can occur much faster than it would in the absence of the enzyme.
Enzymes
A catalyst is something that speeds something up. Enzymes are biological catalysts that speed up reactions. They accomplish this by lowering the activation energy and helping the transition state to form. Thus, the reaction can occur more quickly because the tricky transition state is not as much of a hurdle to overcome. Enzymes do NOT change the energy of the starting point or the ending point of the reaction. They only lower the activation energy.
Enzyme Specificity
Most of the crucial reactions that occur in the cell require enzymes. Yet enzymes themselves are highly specific—in fact, each enzyme catalyzes only one kind of reaction. This is known as enzyme specificity. Because of this, enzymes are usually named after the molecules they target. In enzymatic reactions, the targeted molecules are known as substrates. For example, maltose, a disaccharide, can be broken down into two glucose molecules. Our substrate, maltose, gives its name to the enzyme that catalyzes this reaction: maltase.
Many enzymes are named simply by replacing the suffix of the substrate with –ase. Using this nomenclature, maltose becomes maltase.
Enzyme-Substrate Complex
Enzymes have a unique way of helping reactions along. As we just saw, the reactants in an enzyme-assisted reaction are known as substrates. During a reaction, the enzyme’s job is to bring the transition state about by helping the substrate(s) get into position. It accomplishes this through a special region on the enzyme known as an active site.
The enzyme temporarily binds one or more of the substrates to its active site and forms an enzyme-substrate complex. Let’s take a look:
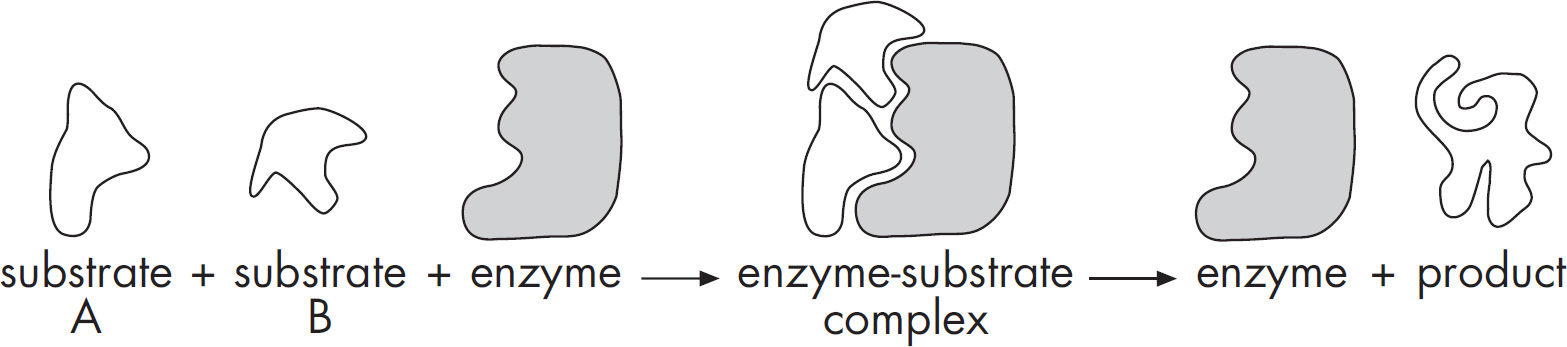
Once the reaction has occurred and the product is formed, the enzyme is released from the complex and restored to its original state. Now the enzyme is free to react again with another bunch of substrates.
By binding and releasing substrates over and over again, the enzyme speeds the reaction along, enabling the cell to release much-needed energy from various molecules. Here is a quick review on the function of enzymes.
Enzymes Do:
• increase the rate of a reaction by lowering the reaction’s activation energy
• form temporary enzyme-substrate complexes
• remain unaffected by the reaction
Enzymes Don’t:
• change the reaction
• make reactions occur that would otherwise not occur at all
Induced Fit
However, scientists have discovered that enzymes and substrates don’t fit together quite so seamlessly. It appears that the enzyme has to change its shape slightly to accommodate the shape of the substrates. This is called induced fit. Sometimes certain factors are involved in activating the enzyme and making it capable of binding the substrate. Because the fit between the enzyme and the substrate must be perfect, enzymes only operate under a strict set of biological conditions.
Enzymes Don’t Always Work Alone
Enzymes sometimes need a little help in catalyzing a reaction. Those factors are known as cofactors. Cofactors can either be organic molecules called coenzymes or they can be inorganic molecules or ions. The inorganic cofactors are usually metal ions (Fe2+, Mg2+). Vitamins are examples of organic coenzymes. Your daily dose of vitamins is important for just this reason: the vitamins are active and necessary participants in crucial chemical reactions.
Factors Affecting Reaction Rates
Enzymatic reactions can be influenced by a number of factors, such as temperature, pH, and the relative amounts of enzyme and substrate.
Temperature
The rate of a reaction increases with increasing temperature, up to a point, because an increase in the temperature of a reaction increases the chance of collisions among the molecules. But too much heat can damage an enzyme. If a reaction is conducted at an excessively high temperature (above 42°C), the enzyme loses its three-dimensional shape and becomes inactivated. Enzymes damaged by heat and deprived of their ability to catalyze reactions are said to be denatured.
Here’s one thing to remember: all enzymes operate at an ideal temperature. For most human enzymes, this temperature is body temperature, 37°C.
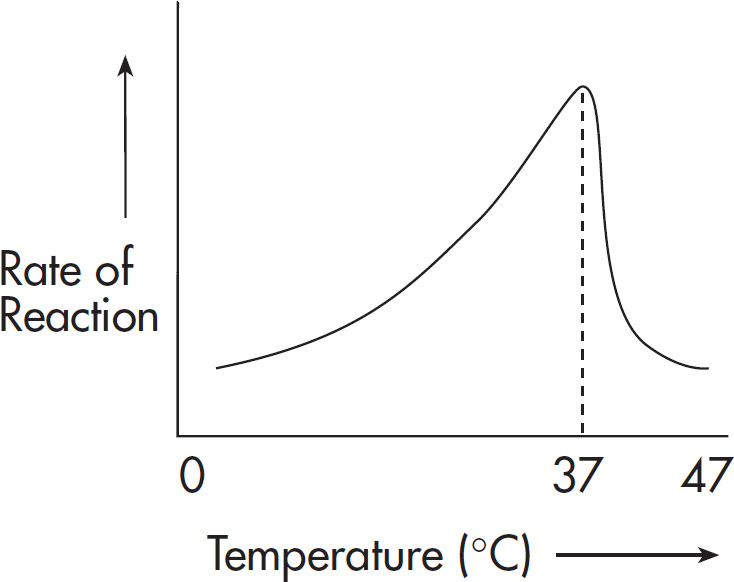
Not all organisms have a constant body temperature. For example, ectotherms depend on the environment to control their varying body temperatures. Q10 is a measure of temperature sensitivity of a physiological process or enzymatic reaction rate. The equation for Q10 is included on the AP Biology Equations and Formulas sheet.
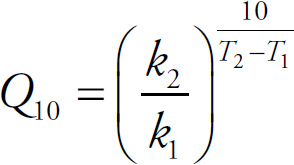
The temperature unit must be in either Celsius or Kelvin, and the same unit must be used for T1 and T2. The two reaction rates (k1 and k2) must also have the same unit. Q10 has no unit; it is the factor by which the rate of a reaction increases. The more temperature-dependent a reaction is, the higher Q10 will be. Reactions with Q10 = 1 are temperature independent.
pH
Enzymes also function best at a particular pH. For most enzymes, the optimal pH is at or near a pH of 7.
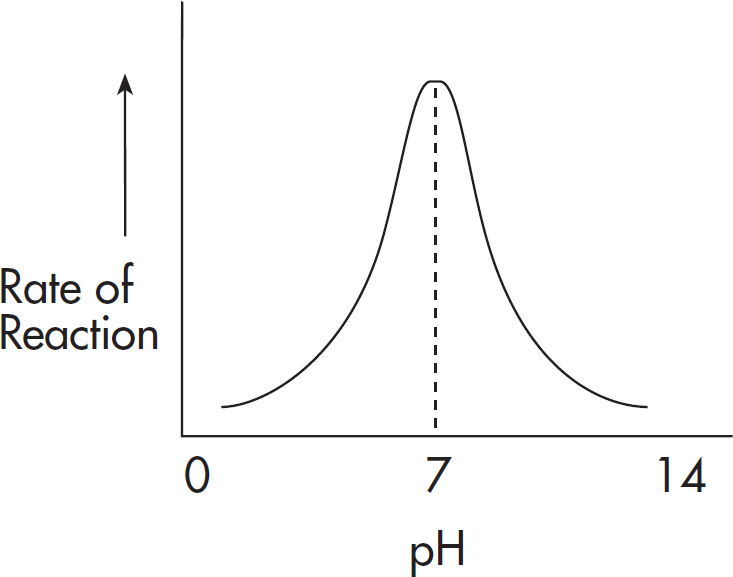
Other enzymes operate at a low pH. For instance, pepsin, the digestive enzyme found in the stomach, is most effective at an extremely acidic pH of 2.
Enzyme Regulation
We know that enzymes control the rates of chemical reactions. But what regulates the activity of enzymes? It turns out that a cell can control enzymatic activity by regulating the conditions that influence the shape of the enzyme. Enzymes can be turned on/off by things that bind to them. Sometimes these things can bind at the active site, and sometimes they bind at other sites, called allosteric sites.
If the substance has a shape that fits the active site of an enzyme (i.e., similar to the substrate), it can compete with the substrate and block the substrate from getting into the active site. This is called competitive inhibition. If there was enough substrate available, the substrate would out-compete the inhibitor and the reaction would occur. You can always tell a competitive inhibitor based on what happens when you flood the system with lots of substrate. If the inhibitor binds to an allosteric site, it is an allosteric inhibitor and it is noncompetitive inhibition. A noncompetitive inhibitor generally distorts the enzyme shape so that it cannot function. The substrate can still bind, but the enzyme will not be able to catalyze the reaction.
REACTION COUPLING
As we just saw, almost everything an organism does requires energy. How, then, can the cell acquire the energy it needs without becoming a major mess? Fortunately, it’s through adenosine triphosphate (ATP).
ATP, as the name indicates, consists of a molecule of adenosine bonded to three phosphates. The great thing about ATP is that an enormous amount of energy is packed into those phosphate bonds, particularly the third bond.
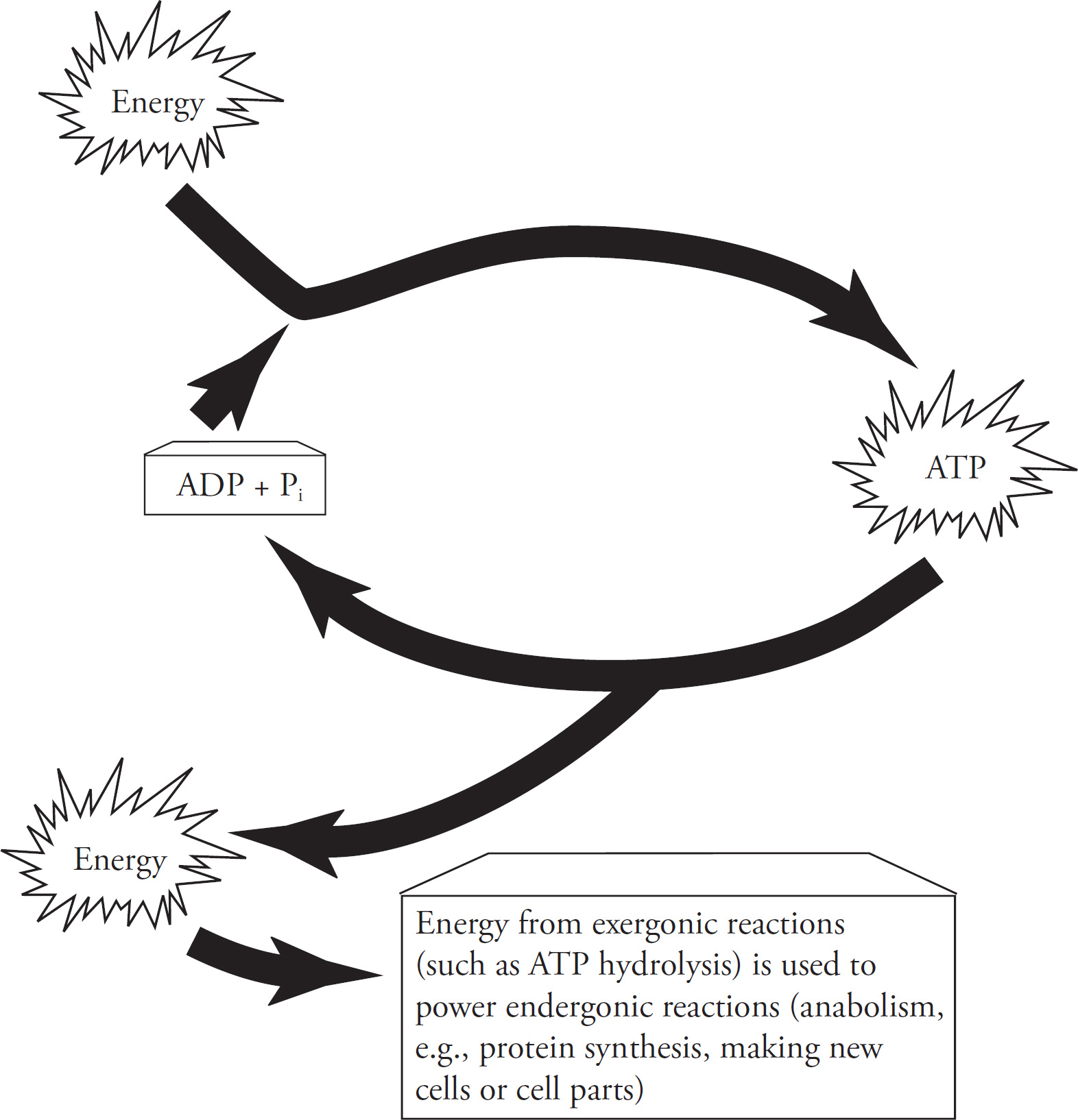
When a cell needs energy, it takes one of these potential-packed molecules of ATP and splits off the third phosphate, forming adenosine diphosphate (ADP) and one loose phosphate (Pi ), while releasing energy in the process.
ATP → ADP + Pi + energy
The energy released from this reaction can then be put to whatever use the cell pleases. Of course, this doesn’t mean that the cell is above the laws of thermodynamics. But within those constraints, ATP is the best source of energy the cell has available. It is relatively neat (only one bond needs to be broken to release that energy) and relatively easy to form. Organisms can use exergonic processes that increase energy, like breaking down ATP, to power endergonic reactions, like building organic macromolecules.
Sources of ATP
But where does all this ATP come from? It can be formed in several ways, but the bulk of it comes from a process called cellular respiration. Cellular respiration is a process of breaking down glucose and making ATP.
In autotrophs, the sugar is made during photosynthesis. In heterotrophs, the glucose comes from the food we eat. We will start by going over photosynthesis, since in plants it is a precursor to cellular respiration.
PHOTOSYNTHESIS
Plants and algae are producers. All they do is bask in the sun, churning out the glucose necessary for life.
As we discussed earlier, photosynthesis is the process by which light energy is converted to chemical energy. Here’s an overview of photosynthesis.
6CO2 + 6H2O → C6H12O6 + 6O2
You’ll notice from this equation that carbon dioxide and water are the raw materials plants use to manufacture simple sugars. But remember, there’s much more to photosynthesis than what’s shown in the simple reaction above. We’ll soon see that this beautifully orchestrated process occurs thanks to a whole host of special enzymes and pigments. But before we turn to the stages in photosynthesis, let’s talk about where photosynthesis occurs.
The leaves of plants contain lots of chloroplasts, which are the primary sites of photosynthesis.
Now let’s look at an individual chloroplast. If you split the membrane of a chloroplast, you’ll find a fluid-filled region called the stroma. Inside the stroma are structures that look like stacks of coins. These structures are the grana.
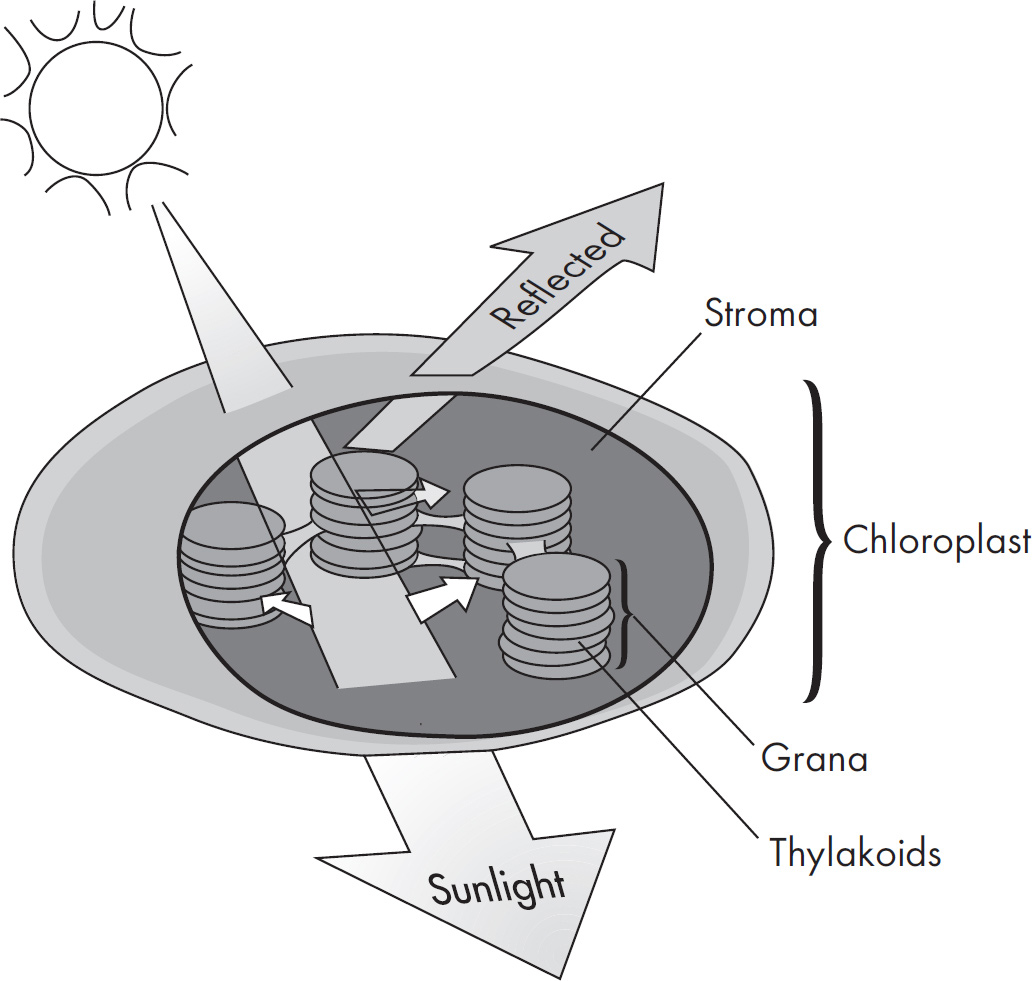
The many disk-like structures that make up the grana are called the thylakoids. They contain chlorophyll, the light-absorbing pigment that drives photosynthesis, as well as enzymes involved in the process.
There are two stages in photosynthesis: the light reactions (also called the light-dependent reactions) and the dark reactions (also called the light-independent reactions). The whole process begins when the photons (or “energy units”) of sunlight strike a leaf, activating chlorophyll and exciting electrons. The activated chlorophyll molecule then passes these excited electrons down to a series of electron carriers, ultimately producing ATP and NADPH. The whole point of the light reaction is to produce two things: (1) energy in the form of ATP and (2) electron carriers, specifically NADPH. Both of these products, along with carbon dioxide, are then used in the dark reaction (light-independent) to make carbohydrates.
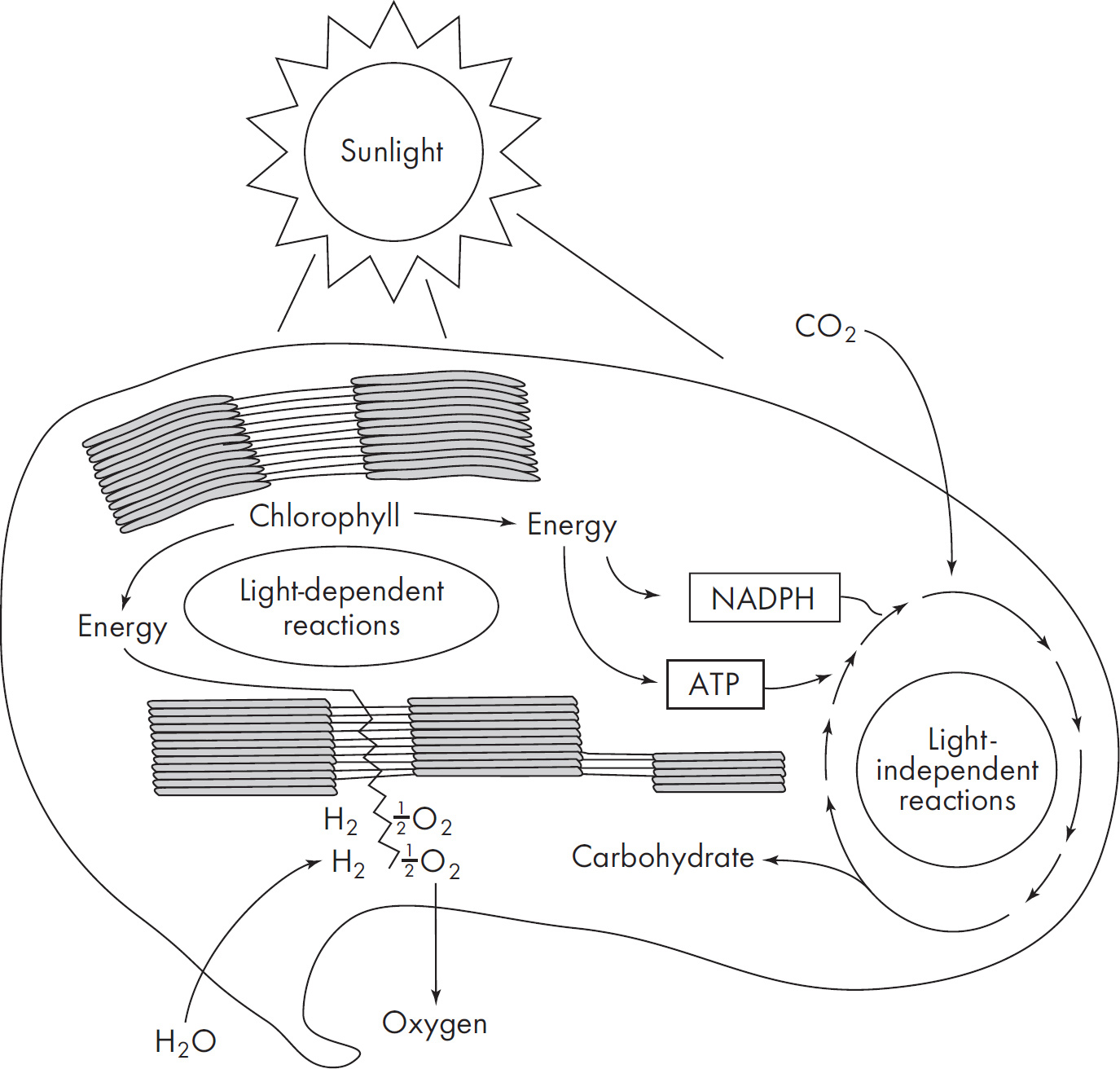
THE LIGHT REACTION
The Pigments
Many light-absorbing pigments participate in photosynthesis. Some of the more important ones are chlorophyll a, chlorophyll b, and carotenoids. These pigments are clustered in the thylakoid membrane into units called antenna complexes.
All of the pigments within a unit are able to “gather” light, but they’re not able to “excite” the electrons. Only one special molecule—located in the reaction center— is capable of transforming light energy to chemical energy. In other words, the other pigments, called antenna pigments, “gather” light and “bounce” the energy to the reaction center.
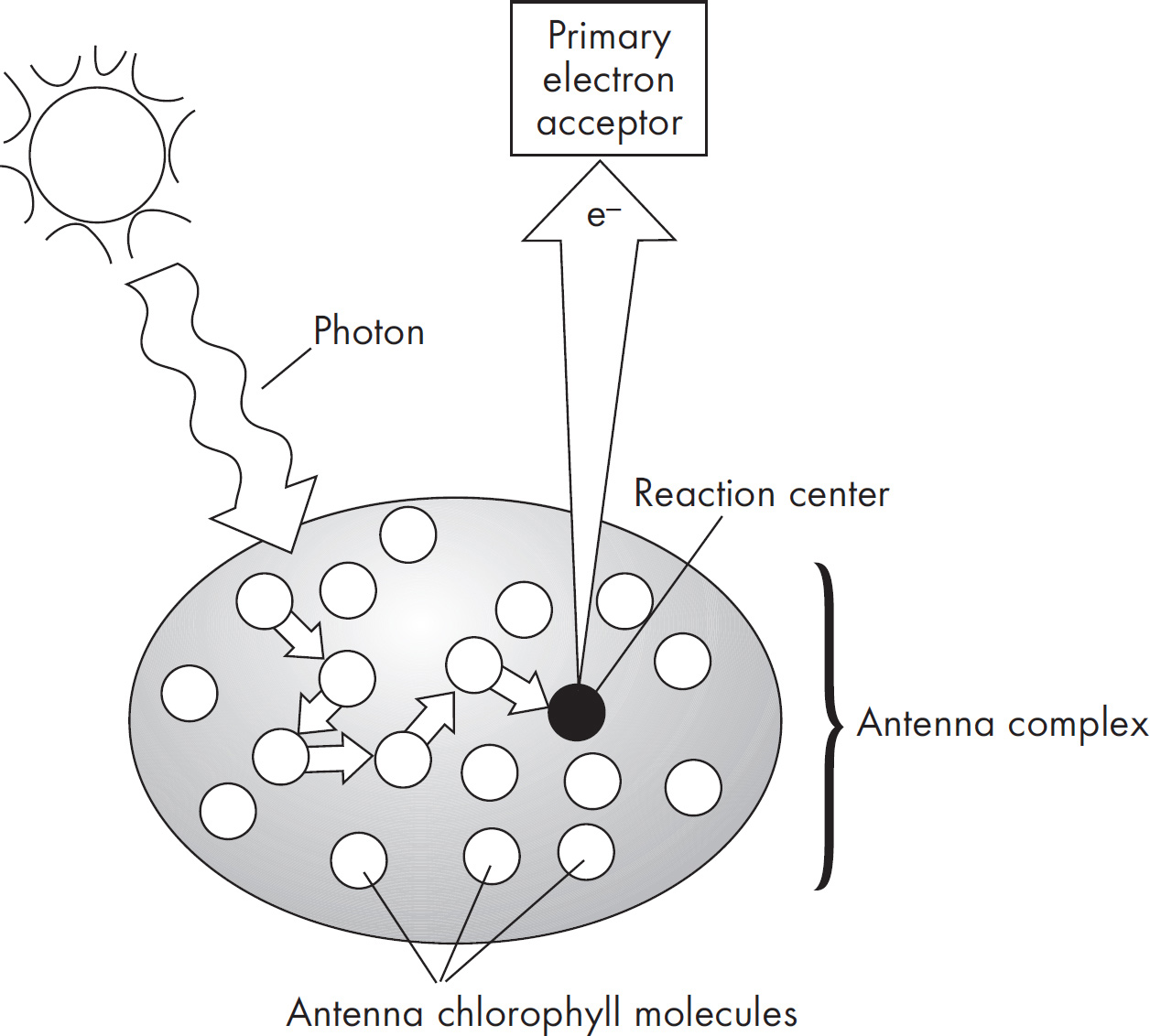
There are two types of reaction centers: photosystem I (PS I) and photosystem II (PS II). The principal difference between the two is that each reaction center has a specific type of chlorophyll—chlorophyll a—that absorbs a particular wavelength of light. For example, P680, the reaction center of photosystem II, has a maximum absorption at a wavelength of 680 nanometers. The reaction center for photosystem I—P700—best absorbs light at a wavelength of 700 nanometers.
When light energy is used to make ATP, it is called photophosphorylation. Autotrophs are using light (that’s photo) and ADP and phosphates (that’s phosphorylation) to produce ATP.
An absorption spectrum shows how well a certain pigment absorbs electromagnetic radiation. Light absorbed is plotted as a function of radiation wavelength. This spectrum is the opposite of an emission spectrum, which gives information on which wavelengths are emitted by a pigment. The absorption spectrum for chlorophyll a, chlorophyll b and carotenoids is below. You can see that chlorophyll a and chlorophyll b absorb blue and red light quite well but do not absorb light in the green part of the spectrum. Light in the green range of wavelengths is reflected, and this is why chlorophyll and many plants are green. Carotenoids absorb light on the blue-green end of the spectrum, but not on the other end. This is why plants rich in carotenoids are yellow, orange or red.
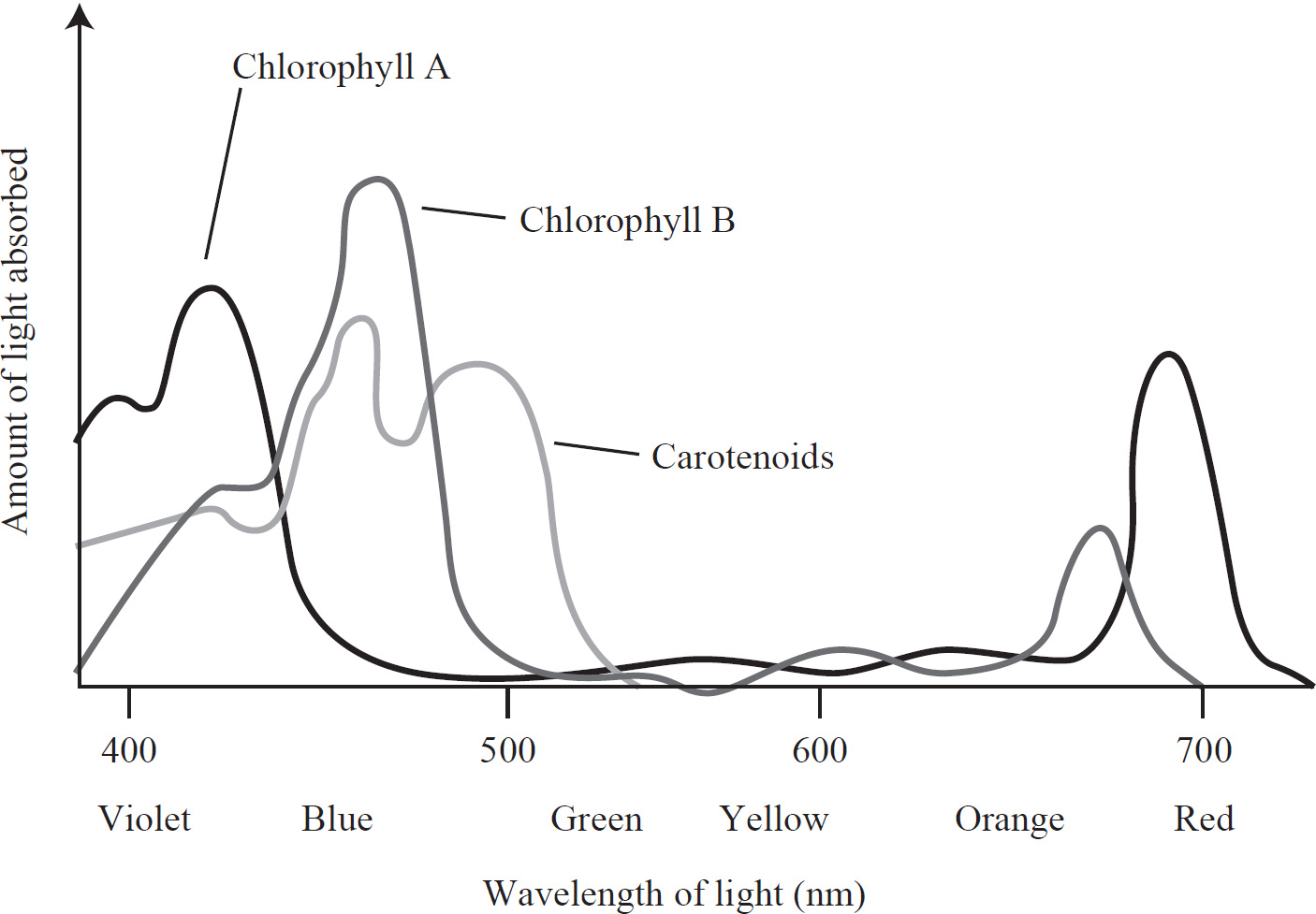
Capturing Energy
When a leaf captures sunlight, the energy is sent to P680, the reaction center for photosystem II. The activated electrons are trapped by P680 and passed to a molecule called the primary acceptor, and then they are passed down to carriers in the electron transport chain and eventually enter photosystem I.
Some of the energy dissipates as electrons move along the chain of acceptors and will be used to “pump” protons across the membrane into the thylakoid lumen. To replenish the electrons in the thylakoid, when the reaction center P680 absorbs light, it also splits water into oxygen, hydrogen ions, and electrons. That process is called photolysis. The electrons from photolysis replace the missing electrons in photosystem II.
Here’s how it works: Hydrogen ions accumulate inside the thylakoids when photolysis occurs. A proton gradient is established. As the hydrogen ions move through ATP synthase, ADP and Pi produce ATP.
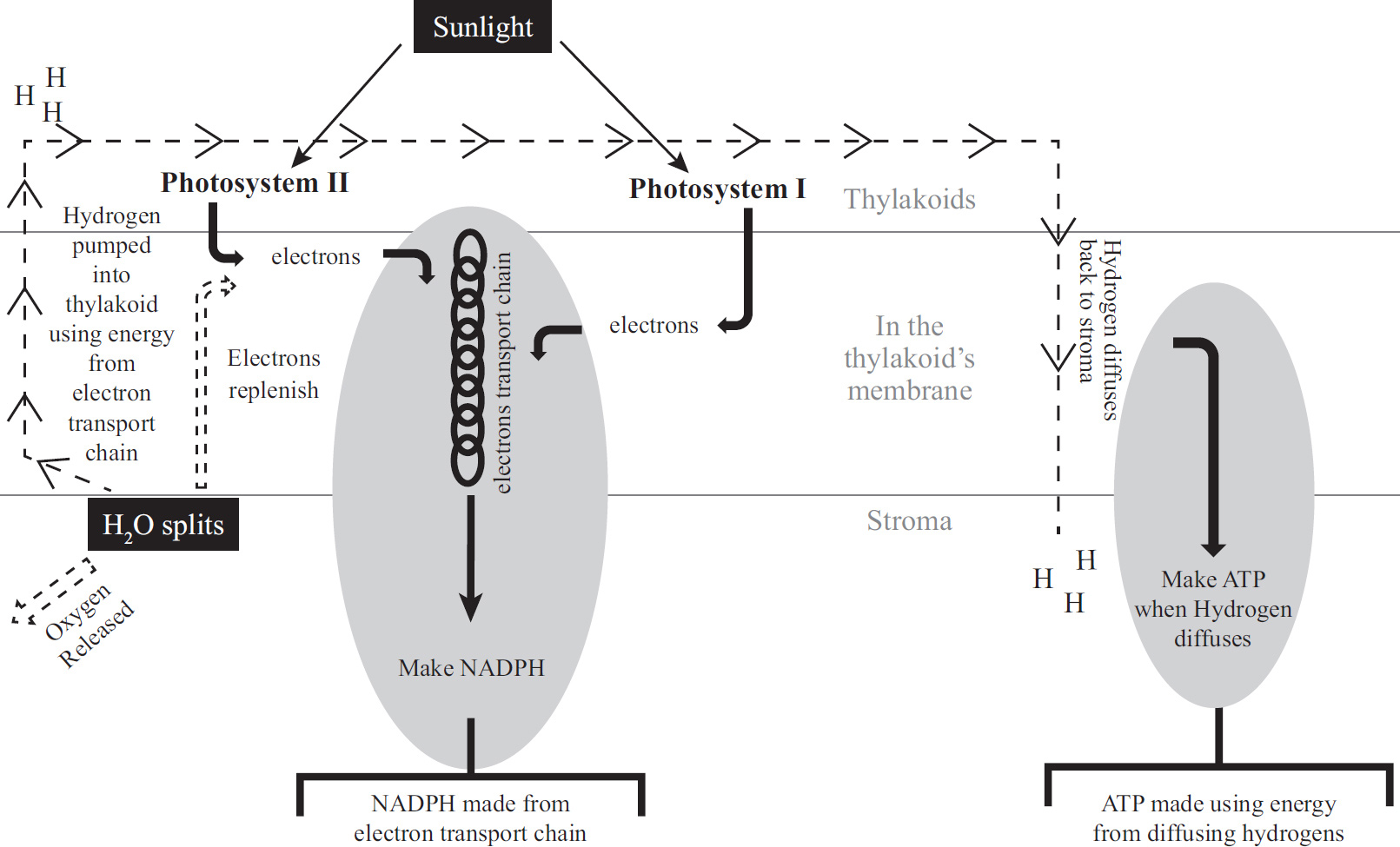
After the electrons leave photosystem II, they go to photosystem I. The electrons are passed through a second electron transport chain until they reach the final electron acceptor NADP+ to make NADPH.
The light reaction occurs in the thylakoids.
• P680 in photosystem II captures light and passes excited electrons down an electron transport chain to produce ATP.
• P700 in photosystem I captures light and passes excited electrons down an electron transport chain to produce NADPH.
• A molecule of water is split by sunlight, releasing electrons, hydrogen, and free O2.
The light-dependent reactions occur in the grana of chloroplasts, where the thylakoids are found. Remember: the light-absorbing pigments and enzymes for the light-dependent reactions are found within the thylakoids.
THE LIGHT-INDEPENDENT REACTION
Now let’s turn to the dark reaction. The dark reaction uses the products of the light reaction—ATP and NADPH—to make sugar. We now have energy to make glucose, but what do plants use as their carbon source? CO2, of course. You’ve probably heard of the term carbon fixation. All this means is that CO2 from the air is converted into carbohydrates. This step occurs in the stroma of the leaf. The dark reactions are also called the Calvin-Benson Cycle (or just Calvin cycle).
Let’s summarize the important facts about the dark reaction:
• The Calvin cycle occurs in the stroma of chloroplasts.
• ATP and NADPH from the light reaction are necessary for carbon fixation.
• CO2 is fixed to form glucose.
CAM Photosynthesis
In some plants, the stomates are closed during the day to reduce excessive water loss from transpiration. You might think that this would prevent these plants from carrying out photosynthesis during the day. However, desert plants have evolved a way to perform photosynthesis when their stomates are closed; it’s called CAM (crassulacean acid metabolism) photosynthesis. In CAM photosynthesis, a process similar to C4 photosynthesis, PEP carboxylase is used to fix CO2 to oxaloacetate, but oxaloacetate is converted to malic acid instead of malate (an ionized form of malate) and sent to the cell’s vacuole. During the day, malic acid is converted back to oxaloacetate, and carbon dioxide is released for photosynthesis. CO2 then enters the Calvin cycle.
The following table summarizes the stages of photosynthesis:
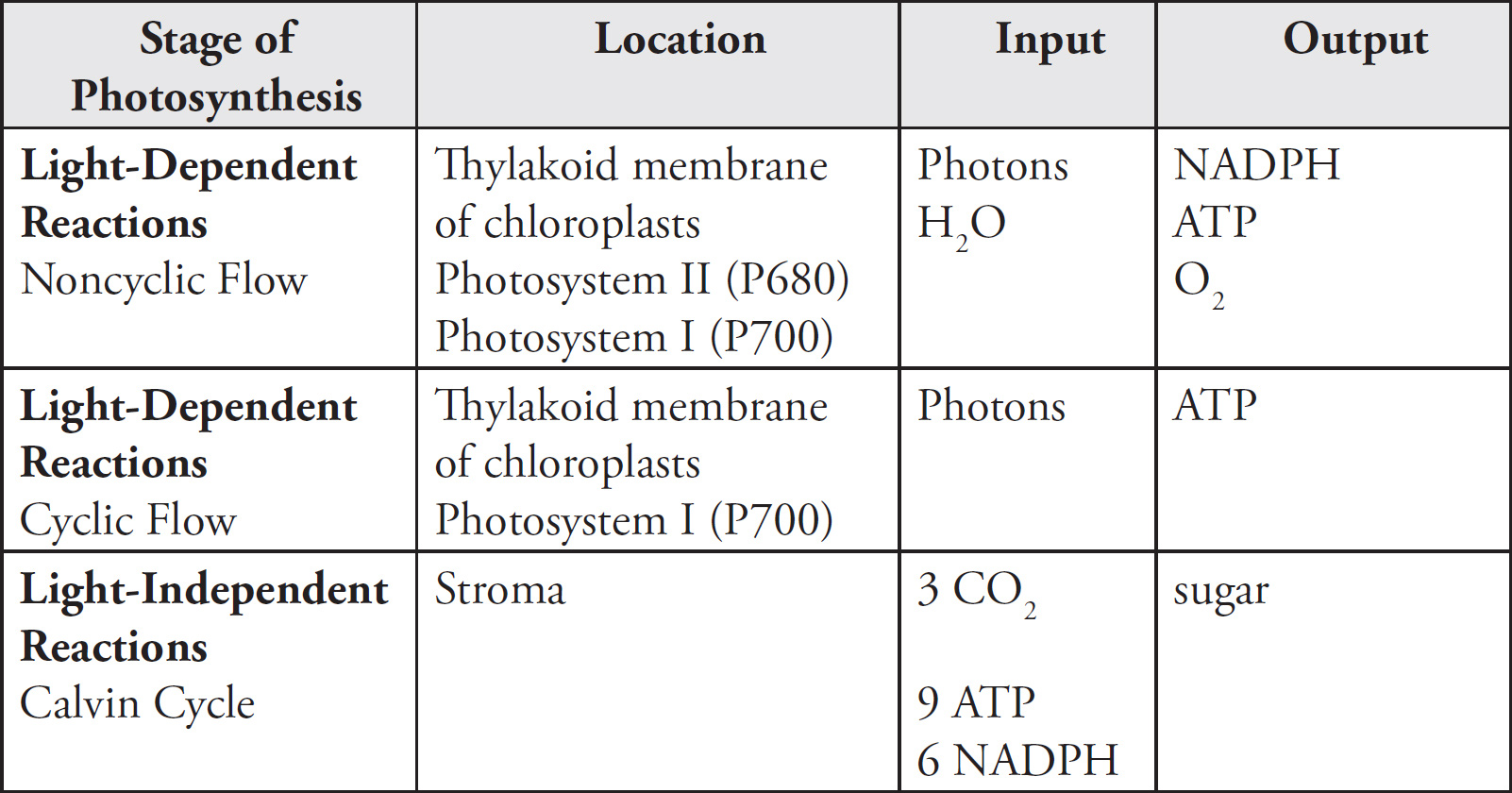
CELLULAR RESPIRATION: THE SHORTHAND VERSION
In the shorthand version, cellular respiration looks something like this.
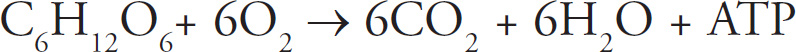
Notice that we’ve taken a sugar, perhaps a molecule of glucose, and combined it with oxygen to produce carbon dioxide, water, and energy in the form of our old friend, ATP. However, as you probably already know, the actual picture of what really happens is far more complicated.
Generally speaking, we can break cellular respiration down into two different approaches: aerobic respiration or anaerobic respiration. If ATP is made in the presence of oxygen, we call it aerobic respiration. If oxygen isn’t present, we call it anaerobic respiration. Let’s jump right in with aerobic respiration.
AEROBIC RESPIRATION
Aerobic respiration consists of four stages:
1. Glycolysis
2. Formation of acetyl-CoA
3. The Krebs (or citric acid) cycle
4. Oxidative phosphorylation (or the electron transport chain + chemiosmosis)
In the first three stages, glucose is broken down, and energy molecules are made. Some of these are ATP, and others are special electron carriers called NADH and FADH2. In the fourth stage, the electron carriers unload their electrons, and the energy is eventually used to make even more ATP.
Stage 1: Glycolysis
The first stage begins with glycolysis, the splitting (-lysis) of glucose (glyco-). Glucose is a six-carbon molecule that is broken into two three-carbon molecules called pyruvic acid. This breakdown of glucose also results in the net production of two molecules of ATP.
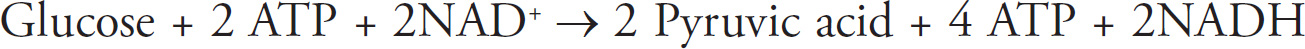
Although we’ve written glycolysis as if it were a single reaction, this process doesn’t occur in one step. In fact, it requires a sequence of enzyme-catalyzed reactions.
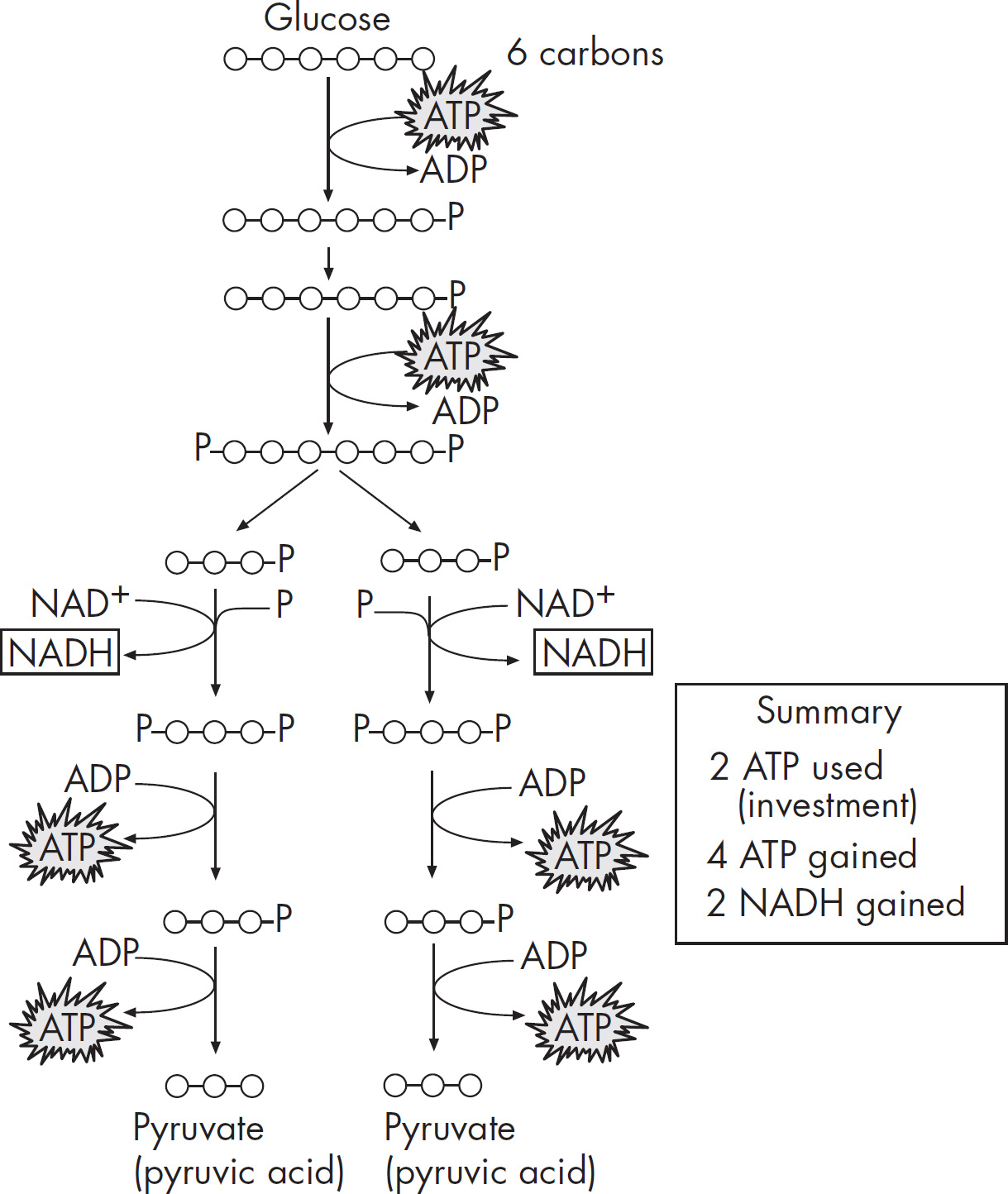
If you take a good look at the reaction above, you’ll see two ATPs are needed to produce four ATP. You’ve probably heard the expression, “You have to spend money to make money.” In biology, you have to invest ATP to make ATP: our investment of two ATP yielded four ATP, for a net gain of two. The ATP molecules generated in glycolysis are created from combining ADP and an inorganic phosphate with the help of an enzyme.
Glycolysis also creates two NADH, which result from the transfer of electrons to the carrier NAD+, which then becomes NADH. NAD+ and NADH are constantly being turned into each other as electrons are being carried and then unloaded.
There are four important tidbits to remember regarding glycolysis:
• occurs in the cytoplasm
• net of 2 ATPs produced
• 2 pyruvic acids formed
• 2 NADH produced
Stage 2: Formation of Acetyl-CoA
Pyruvic acid is transported to the mitochondrion. Each pyruvic acid (a three-carbon molecule) is converted to acetyl coenzyme A (a two-carbon molecule, usually just called acetyl-CoA) and CO2 is released.
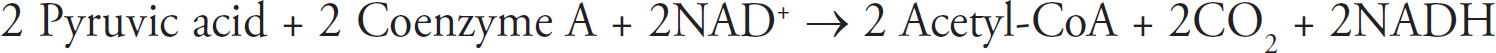
Are you keeping track of our carbons? We’ve now gone from two three-carbon molecules to two two-carbon molecules. The extra carbons leave the cell in the form of CO2. Once again, two molecules of NADH are also produced. This process of turning pyruvic acid into acetyl-CoA is called the pyruvate dehydrogenase complex.
Stage 3: The Krebs Cycle
The next stage is the Krebs cycle, also known as the citric acid cycle. Each of the two acetyl coenzyme A molecules will enter the Krebs cycle, one at a time, and all the carbons will ultimately be converted to CO2. This stage occurs in the matrix of the mitochondria.
The Krebs cycle begins with each molecule of acetyl-CoA produced from the second stage of aerobic respiration combining with oxaloacetate, a four-carbon molecule, to form a six-carbon molecule, citric acid or citrate (hence its name, the citric acid cycle).
Citrate gets turned into several other things, and because the cycle begins with a four-carbon molecule, oxaloacetate, it eventually gets turned back into oxaloacetate to maintain the cycle by joining with the next acetyl-CoA coming down the pipeline.
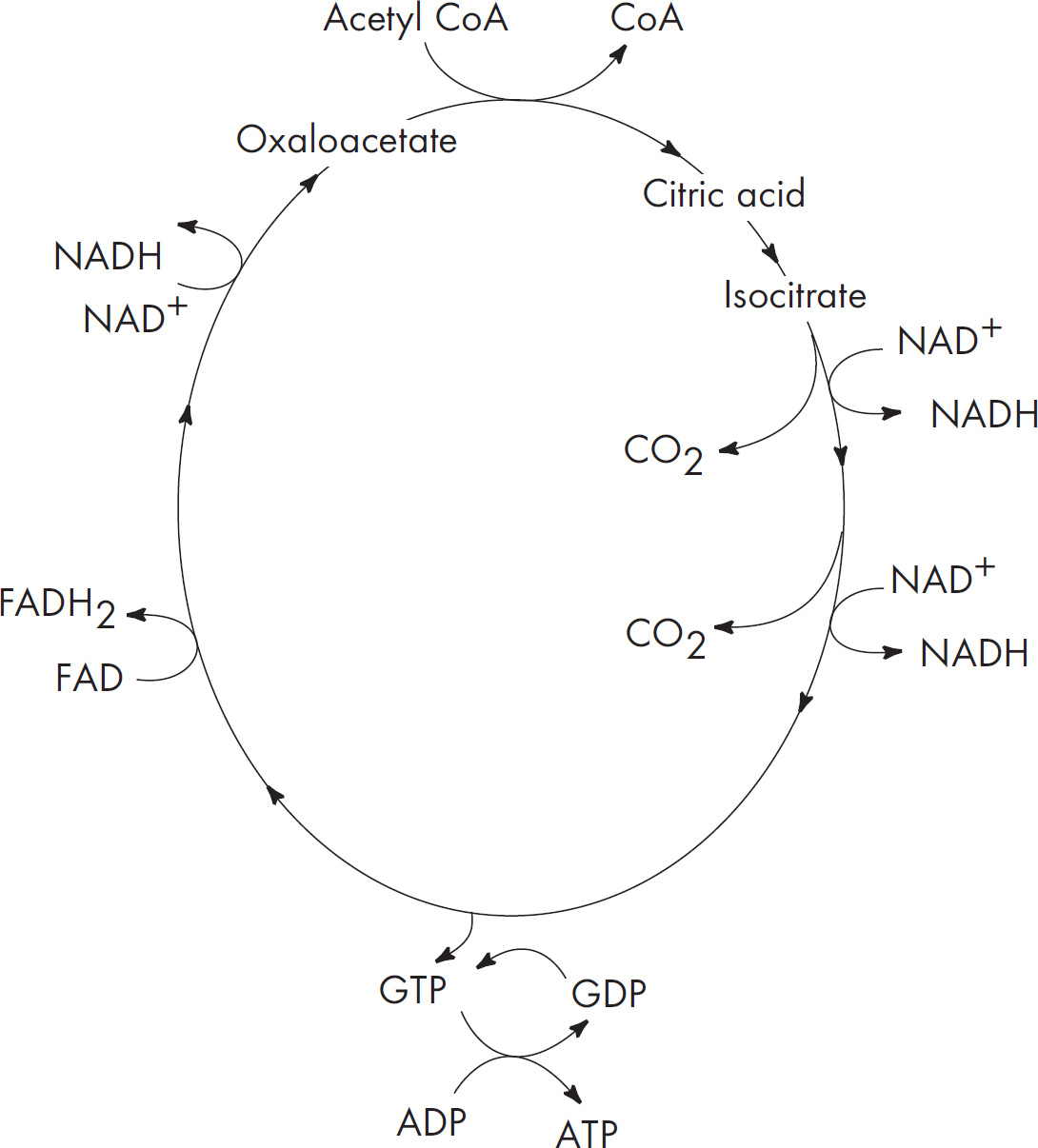
With each turn of the cycle, three types of energy are produced:
• 1 ATP
• 3 NADH
• 1 FADH2
To figure out the total number of products per molecule of glucose, we simply double the number of products—after all, we started off the Krebs cycle with two molecules of acetyl-CoA for each molecule of glucose!
Now we’re ready to tally up the number of ATP produced.
So far, we’ve made only four ATP—two ATP from glycolysis and two ATP from the Krebs cycle.
Although that seems like a lot of work for only four ATP, we have also produced hydrogen carriers in the form of 10NADH and 2FADH2. These molecules will, in turn, produce lots of ATP.
Stage 4: Oxidative Phosphorylation
Electron Transport Chain
As electrons (and the hydrogen atoms to which they belong) are removed from a molecule of glucose, they carry with them much of the energy that was originally stored in their chemical bonds. These electrons—and their accompanying energy—are then transferred to readied hydrogen carrier molecules. In the case of cellular respiration, these charged carriers are NADH and FADH2.
Let’s see how many “loaded” electron carriers we’ve produced. We now have:
• 2 NADH molecules from glycolysis
• 2 NADH from the production of acetyl-CoA
• 6 NADH from the Krebs cycle
• 2 FADH2 from the Krebs cycle
That gives us 12 altogether.
These electron carriers—NADH and FADH2—“shuttle” electrons to the electron transport chain, the resulting NAD+ and FADH can be recycled to be used as carriers again, and the hydrogen atoms are split into hydrogen ions and electrons.
H2 → 2H+ + 2e–
Then, two interesting things occur. First, the high-energy electrons from NADH and FADH2 are passed down a series of protein carrier molecules that are embedded in the cristae, thus it is called the electron transport chain. Some of the carrier molecules in the electron transport chain are NADH dehydrogenase and cytochrome C.
Each carrier molecule hands down the electrons to the next molecule in the chain. The electrons travel down the electron transport chain until they reach the final electron acceptor, oxygen. Oxygen combines with these electrons (and some hydrogens) to form water. This explains the “aerobic” in aerobic respiration. If oxygen weren’t available to accept the electrons, they wouldn’t move down the chain at all, thereby shutting down the whole process of electron transport.
CHEMIOSMOSIS
At the same time that electrons are being passed down the electron transport chain, another mechanism is at work. Remember those hydrogen ions (also called protons) that split off from the original hydrogen atom? The energy released from the electron transport chain is used to pump hydrogen ions across the inner mitochondrial membrane from the matrix into the intermembrane space. The pumping of hydrogen ions into the intermembrane space creates a pH gradient, or proton gradient. The hydrogen ions really want to diffuse back into the matrix. The potential energy established in this gradient is responsible for the production of ATP. This pumping of ions and diffusion of ions to create ATP is chemiosmosis.
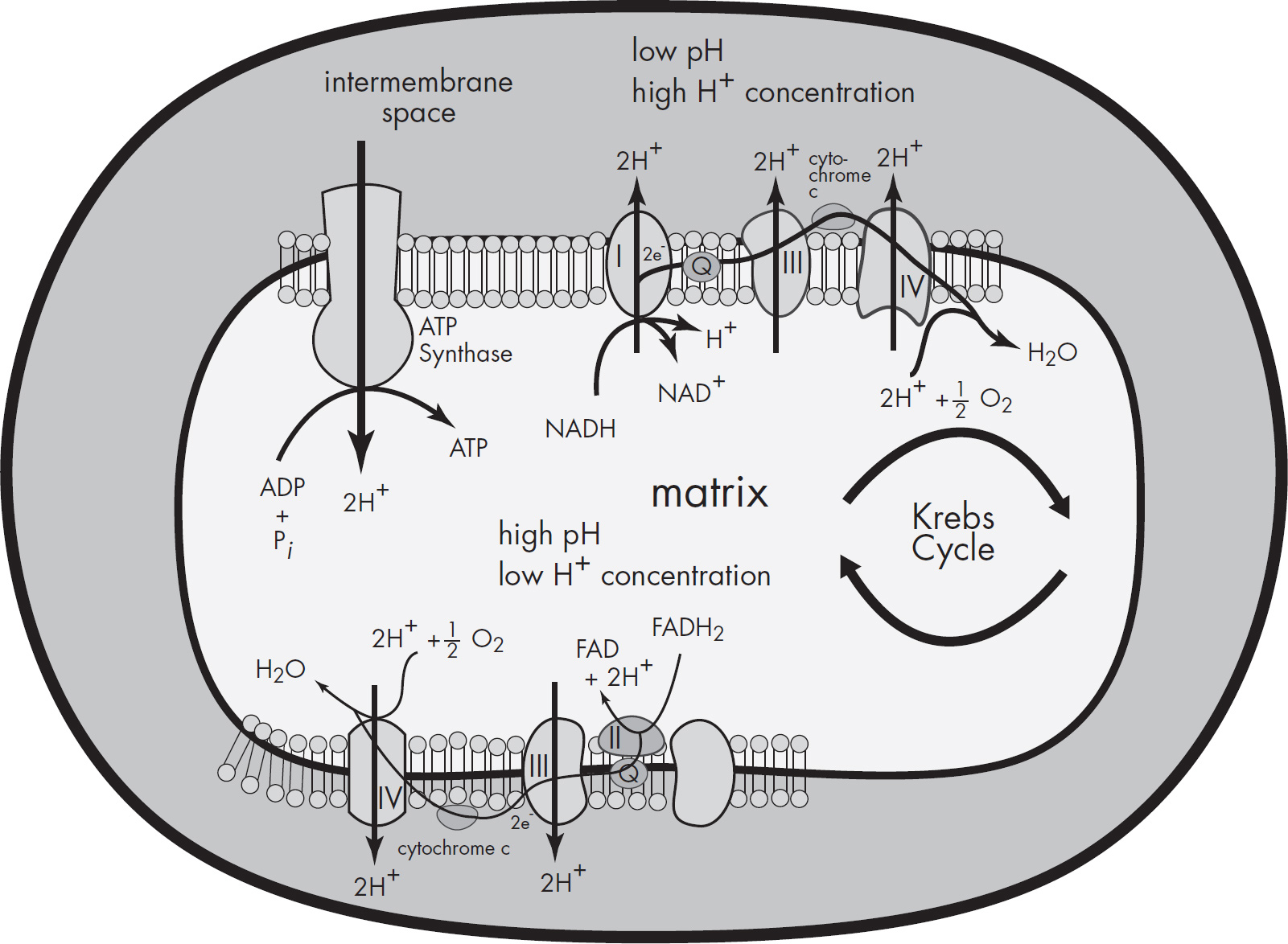
These hydrogen ions can diffuse across the inner membrane only by passing through channels called ATP synthase. Meanwhile, ADP and Pi are on the other side of these protein channels. The flow of protons through these channels produces ATP by combining ADP and Pi on the matrix side of the channel. Overall, this process is called oxidative phosphorylation because when electrons are given up it is called “oxidation” and then ADP is “phosphorylated” to make ATP.
You’re also expected to know the following two things for the AP Biology Exam:
• Every NADH from glycolysis yields 1.5 ATP and every other NADH yields 2.5 ATP.
• Every FADH2 yields 1.5 ATP.
For the AP Biology Exam, all you need to remember are the big steps and the overall outcome.
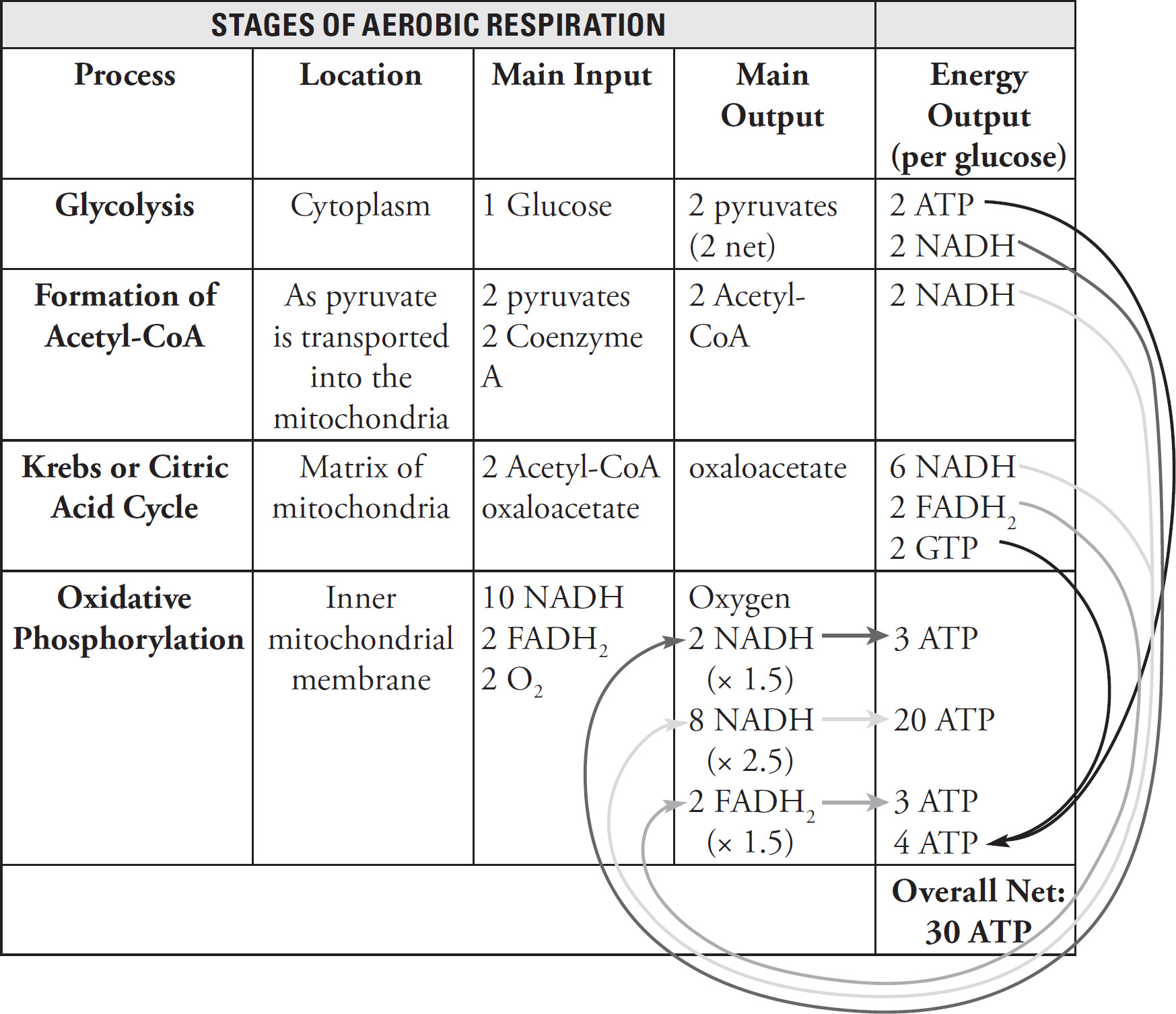
ANAEROBIC RESPIRATION
When oxygen is not available, the anaerobic version of respiration occurs. The electron transport chain stops working, and the electron carriers have nowhere to drop their electrons. The mitochondrial production of acetyl-CoA and the Krebs cycle cease too.
Glycolysis, however, can continue to run. This means that glucose can be broken down to pyruvate to give net two ATP. Only two instead of 30!
Glycolysis also gives two NADH. The pyruvate and the NADH make a deal with each other, and the pyruvate helps the NADH get recycled back into NAD+ and takes its electrons. This creates either lactic acid (in muscles) or ethanol (in yeast). Since these two things are toxic, fermentation is only done in emergencies. Aerobic respiration is a better option. Organisms carry out a process called fermentation. Under anaerobic conditions, pyruvic acid is converted to either lactic acid or ethyl alcohol (or ethanol) and carbon dioxide.
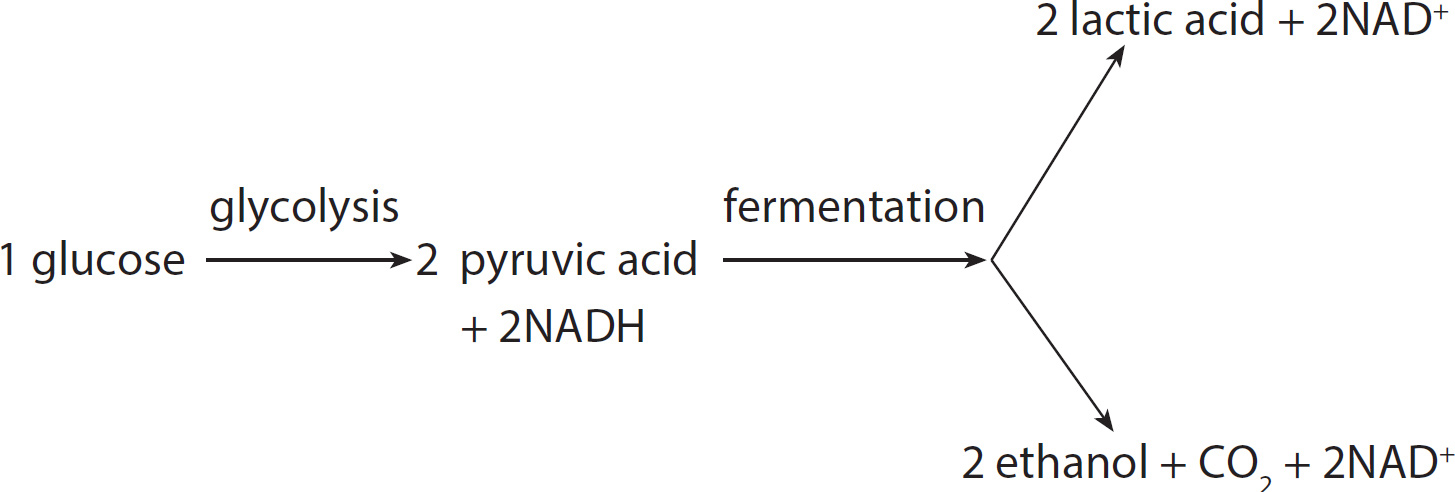
What types of organisms undergo fermentation?
• Yeast cells and some bacteria make ethanol and carbon dioxide.
• Other bacteria produce lactic acid.
Your Muscle Cells Can Ferment
Although human beings are aerobic organisms, they can actually carry out fermentation in their muscle cells. Have you ever had a cramp? If so, that cramp was possibly the consequence of anaerobic respiration.
When you exercise, your muscles require a lot of energy. To get this energy, they convert enormous amounts of glucose to ATP. But as you continue to exercise, your body doesn’t get enough oxygen to keep up with the demand in your muscles. This creates an oxygen debt. What do your muscle cells do? They switch over to anaerobic respiration. Pyruvic acid produced from glycolysis is converted to lactic acid. As a consequence, the lactic acid causes your muscles to ache.
KEY TERMS
bioenergetics
1st Law of Thermodynamics
2nd Law of Thermodynamics
entropy
exergonic reaction
endergonic reaction
Gibbs free energy
enthalpy
transition state
activation energy
enzymes
enzyme specificity
substrates
active site
enzyme-substrate complex
induced fit
coenzymes
cofactors
denatured
Q10
allosteric sites
competitive inhibition
noncompetitive inhibition
allosteric inhibitor
cellular respiration
photosynthesis
stroma
grana
thylakoids
light-dependent reactions
light-independent reactions
photons
chlorophyll a and b
carotenoids
reaction center
antenna pigments
photosystem I
photosystem II
p680
p700
photophosphorylation
absorption spectrum
emission spectrum
photolysis
NADPH
carbon fixation
Calvin-Benson Cycle
CAM photosynthesis
aerobic respiration
NADH
FADH2
anaerobic respiration
glycolysis
pyruvic acid
matrix
acetyl coenzyme A (acetyl-CoA)
Krebs cycle (or citric acid cycle)
oxaloacetate
citric acid
Electron Transport Chain
Chemiosmosis
cytochromes
pH gradient (or proton gradient)
ATP synthase
oxidative phosphorylation
fermentation
lactic acid
ethyl alcohol (or ethanol)
Summary
 Energy cannot be created or destroyed. There are endergonic reactions and exergonic reactions. The change in Gibbs free energy indicates whether the reaction will require energy or release energy.
Energy cannot be created or destroyed. There are endergonic reactions and exergonic reactions. The change in Gibbs free energy indicates whether the reaction will require energy or release energy.
 The study of how cells accomplish biological processes is called bioenergetics.
The study of how cells accomplish biological processes is called bioenergetics.
• Chemical reactions are catalyzed by enzymes.
• Enzymes lower the activation energy of chemical reactions by stabilizing the transition state.
• Enzymes do not change the energy difference between reactants and products, but by lowering the activation energy, they facilitate chemical reactions to occur.
 Enzymes are proteins that are highly specific for their substrate (reactant, which binds at the active site).
Enzymes are proteins that are highly specific for their substrate (reactant, which binds at the active site).
 Enzymes have an optimal, narrow range of temperature and pH in which they have the highest rate of reaction. Outside this range, they may undergo denaturation and, therefore, no longer be active.
Enzymes have an optimal, narrow range of temperature and pH in which they have the highest rate of reaction. Outside this range, they may undergo denaturation and, therefore, no longer be active.
 Enzyme activity may also be regulated or altered by allosteric/noncompetitive inhibitors, competitive inhibitors, and activators.
Enzyme activity may also be regulated or altered by allosteric/noncompetitive inhibitors, competitive inhibitors, and activators.
 Enzymes may require coenzymes or cofactors to help catalyze reactions.
Enzymes may require coenzymes or cofactors to help catalyze reactions.
 ATP is the universal energy molecule in cells. It is created via cellular respiration.
ATP is the universal energy molecule in cells. It is created via cellular respiration.
 Photosynthesis is how plants use energy from sunlight to make sugar.
Photosynthesis is how plants use energy from sunlight to make sugar.
• In the light-dependent reactions, the sunlight is absorbed by chlorophyll and the electrons are passed down a chain and the energy molecules ATP and NADPH are produced. Water is also split into hydrogen and oxygen.
• In the light-independent reactions, sugar is made in the Calvin-Benson Cycle using the energy from ATP and NADPH and the carbon from CO2.
 Cellular respiration consumes oxygen and produces a lot of ATP energy for the cell, using NADH and FADH2 as electron carriers. There are four stages to cellular respiration:
Cellular respiration consumes oxygen and produces a lot of ATP energy for the cell, using NADH and FADH2 as electron carriers. There are four stages to cellular respiration:
• glycolysis
• formation of acetyl-CoA
• Krebs cycle
• oxidative phosphorylation
 Anaerobic respiration occurs when cells lack oxygen to act as a final electron acceptor. In this process, the pyruvates generated by glycolysis are broken down by fermentation to produce lactic acid or ethanol and NAD+ and ATP, which permits glycolysis to continue.
Anaerobic respiration occurs when cells lack oxygen to act as a final electron acceptor. In this process, the pyruvates generated by glycolysis are broken down by fermentation to produce lactic acid or ethanol and NAD+ and ATP, which permits glycolysis to continue.
Chapter 6 Drill
Answers and explanations can be found in Chapter 16.
1. The mitochondrion is a critical organelle structure involved in cellular respiration. Below is a simple schematic of the structure of a mitochondrion. Which of the structural components labeled below in the mitochondrion is the primary location of the Krebs cycle?
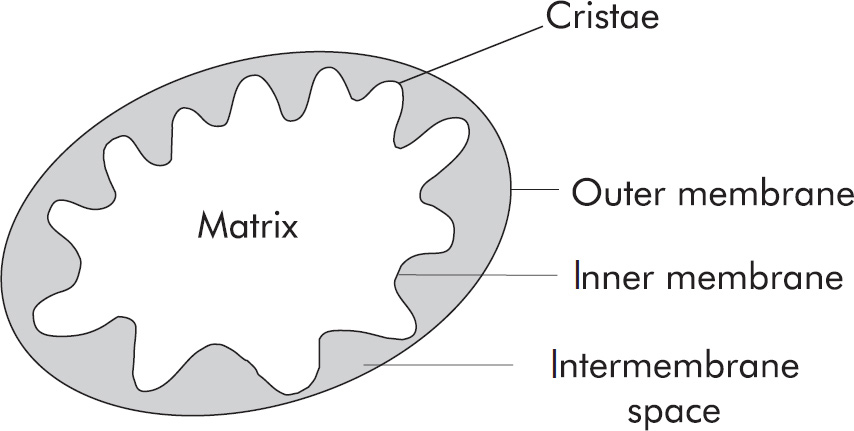
(A) Inner membrane
(B) Matrix
(C) Intermembrane space
(D) Outer membrane
2. Binding of Inhibitor Y as shown below inhibits a key catalytic enzyme by inducing a structural conformation change. Which of the following accurately describes the role of Inhibitor Y?

(A) Inhibitor Y competes with substrates for binding in the active site and functions as a competitive inhibitor.
(B) Inhibitor Y binds allosterically and functions as a competitive inhibitor.
(C) Inhibitor Y competes with substrates for binding in the active site and functions as a non-competitive inhibitor.
(D) Inhibitor Y binds allosterically and functions as a non-competitive inhibitor.
3. A single-step chemical reaction is catalyzed by the addition of an enzyme. Which of the reaction coordinate diagrams accurately shows the effect of the added enzyme (represented by the dashed line) to the reaction?
(A) 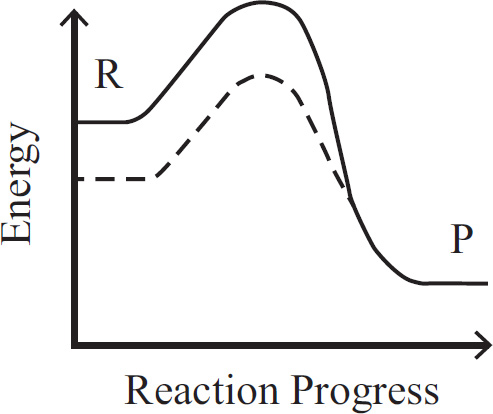
(B) 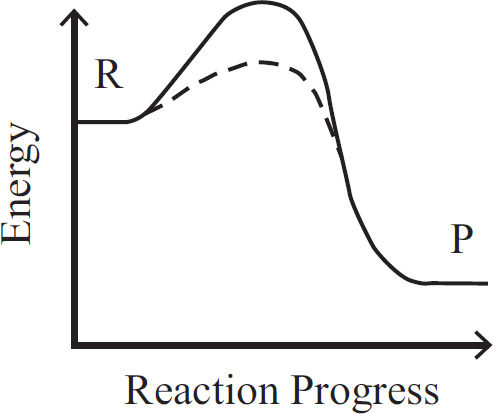
(C) 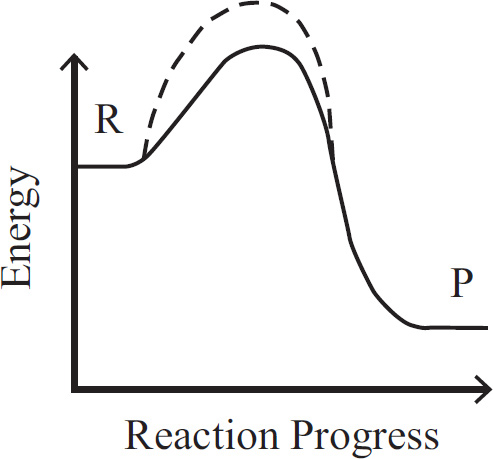
(D) 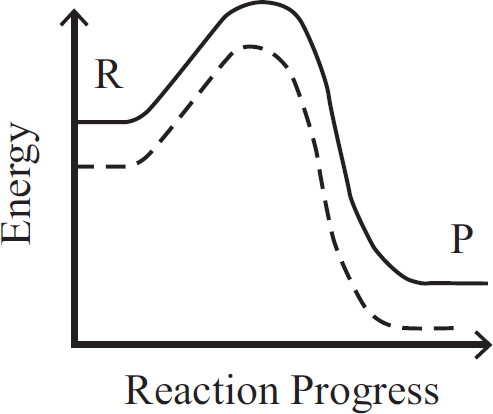
Questions 4-6 refer to the following diagram and paragraph.
Glycolysis (shown below) is a critical metabolic pathway that is utilized by nearly all forms of life. The process of glycolysis occurs in the cytoplasm of the cell and converts 1 molecule of glucose into 2 molecules of pyruvic acid.
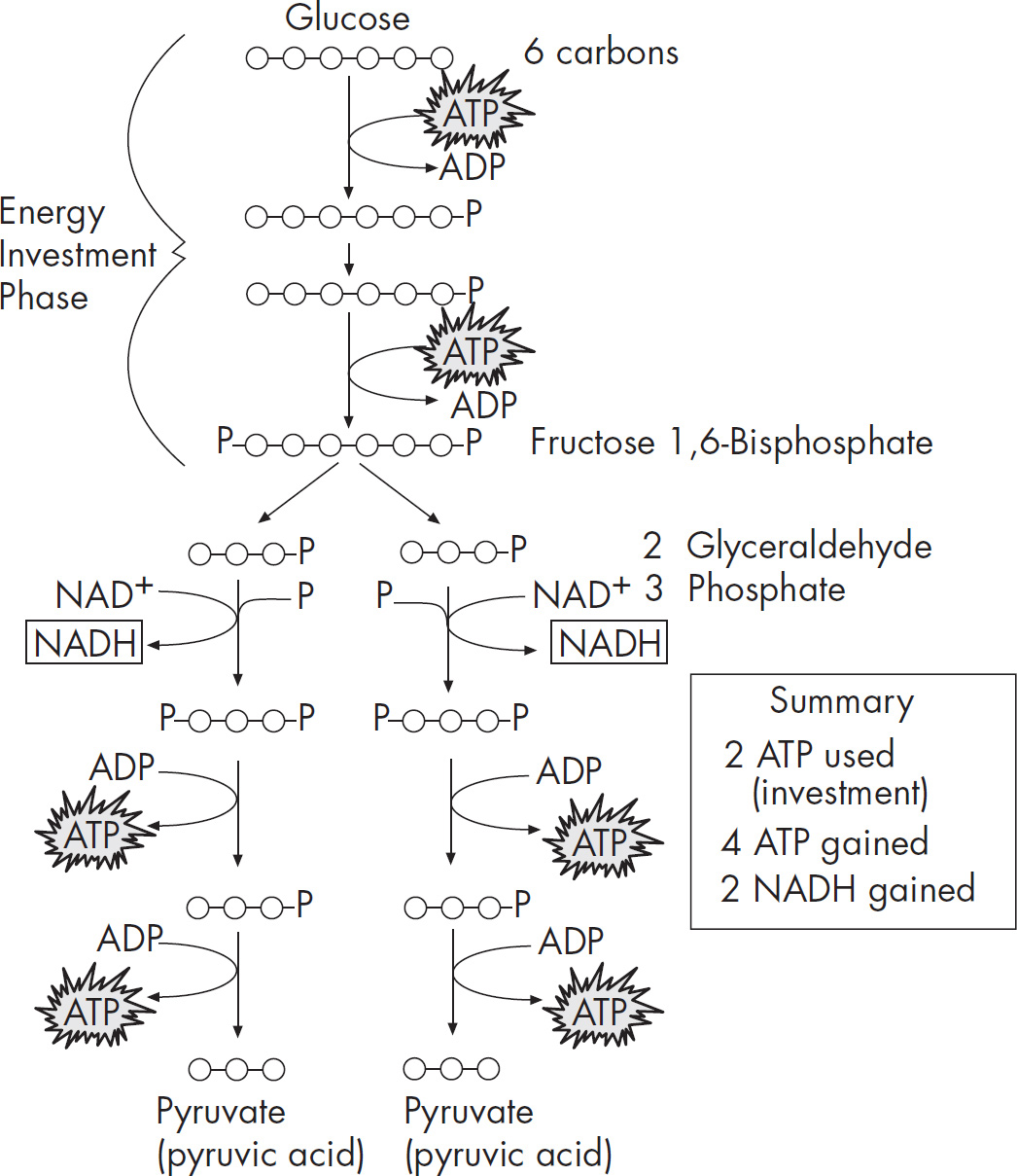
4. How many net ATP would be generated directly from glycolysis from the breakdown of 2 glucose molecules?
(A) 2
(B) 4
(C) 8
(D) 12
5. Glycolysis does not require oxygen to occur in cells. However, under anaerobic conditions, glycolysis normally requires fermentation pathways to occur to continue to produce ATP. Which best describes why glycolysis is dependent on fermentation under anaerobic conditions?
(A) Glycolysis requires fermentation to produce more glucose as a substrate.
(B) Glycolysis requires fermentation to synthesize lactic acid and restore NADH to NAD+.
(C) Glycolysis requires fermentation to generate ATP molecules to complete the early steps of the pathway.
(D) Glycolysis requires fermentation to generate pyruvate for a later step in the pathway.
6. Which of the following most accurately describes the net reaction of glycolysis?
(A) It is an endergonic process because it results in a net increase in energy.
(B) It is an exergonic process because it results in a net increase in energy.
(C) It is an endergonic process because it results in a net decrease in energy.
(D) It is an exergonic process because it results in a net decrease in energy.
7. Taq polymerase, a DNA polymerase derived from thermophilic bacteria, is used in polymerase chain reactions (PCR) in the laboratory. During PCR, Taq catalyzes DNA polymerization, similar to how it would in bacteria. A normal PCR cycle is as follows:
1. Melting/Denaturing |
95°C |
2. Primer Annealing |
50°C |
3. Elongation of DNA |
72°C |
Which of the following conditions likely describes the living environment of Taq bacteria?
(A) Freshwater with acidic pH
(B) Hydrothermal vents reaching temperatures between 70–75°C
(C) Hot springs of 40°C
(D) Tide pools with high salinity
8. The biggest difference between an enzyme-catalyzed reaction and an uncatalyzed reaction is
(A) the free energy between the reactants and the products
(B) the free energy difference between the reactants and the products does not change
(C) the catalyzed reaction would not occur without the enzyme
(D) the energy required to reach the transition state of the reaction
9. Two groups of cells were grown under identical conditions. Mitochondria from each group were isolated and placed in a low pH (approximately pH 6.8) and small molecules were allowed to diffuse across the outer membrane via facilitated diffusion. Both samples were exposed to oxygen bubbles through the growth media. What would you expect to see in terms of ATP production in the sample of cells placed in a low pH, with respect to the control population?
(A) ATP production decreases.
(B) ATP production increases.
(C) ATP production stays the same.
(D) ATP initially increases, then decreases.
10. The second law of thermodynamics states that entropy, or disorder, is constantly increasing in the universe for spontaneous processes. Therefore, how is it possible that organisms exist in very ordered states?
(A) The second law of thermodynamics does not apply to biological life.
(B) Energy can be created or destroyed.
(C) The catabolic reactions are always equal and opposite of the anabolic reactions.
(D) Biological life creates an increase in entropy through dissemination of heat and waste.
REFLECT
Respond to the following questions:
• Which topics from this chapter do you feel you have mastered?
• Which content topics from this chapter do you feel you need to study more before you can answer multiple-choice questions correctly?
• Which content topics from this chapter do you feel you need to study more before you can effectively compose a free response?
• Was there any content that you need to ask your teacher or another person about?
Chapter 7
Molecular Biology
DNA: THE BLUEPRINT OF LIFE
All living things possess an astonishing degree of organization. From the simplest single-celled organism to the largest mammal, millions of reactions and events must be coordinated precisely for life to exist. This coordination is directed by DNA, which is the hereditary blueprint of the cell.
THE MOLECULAR STRUCTURE OF DNA
DNA is made up of repeated subunits of nucleotides. Each nucleotide has a five-carbon sugar, a phosphate, and a nitrogenous base. Take a look at the nucleotide below.
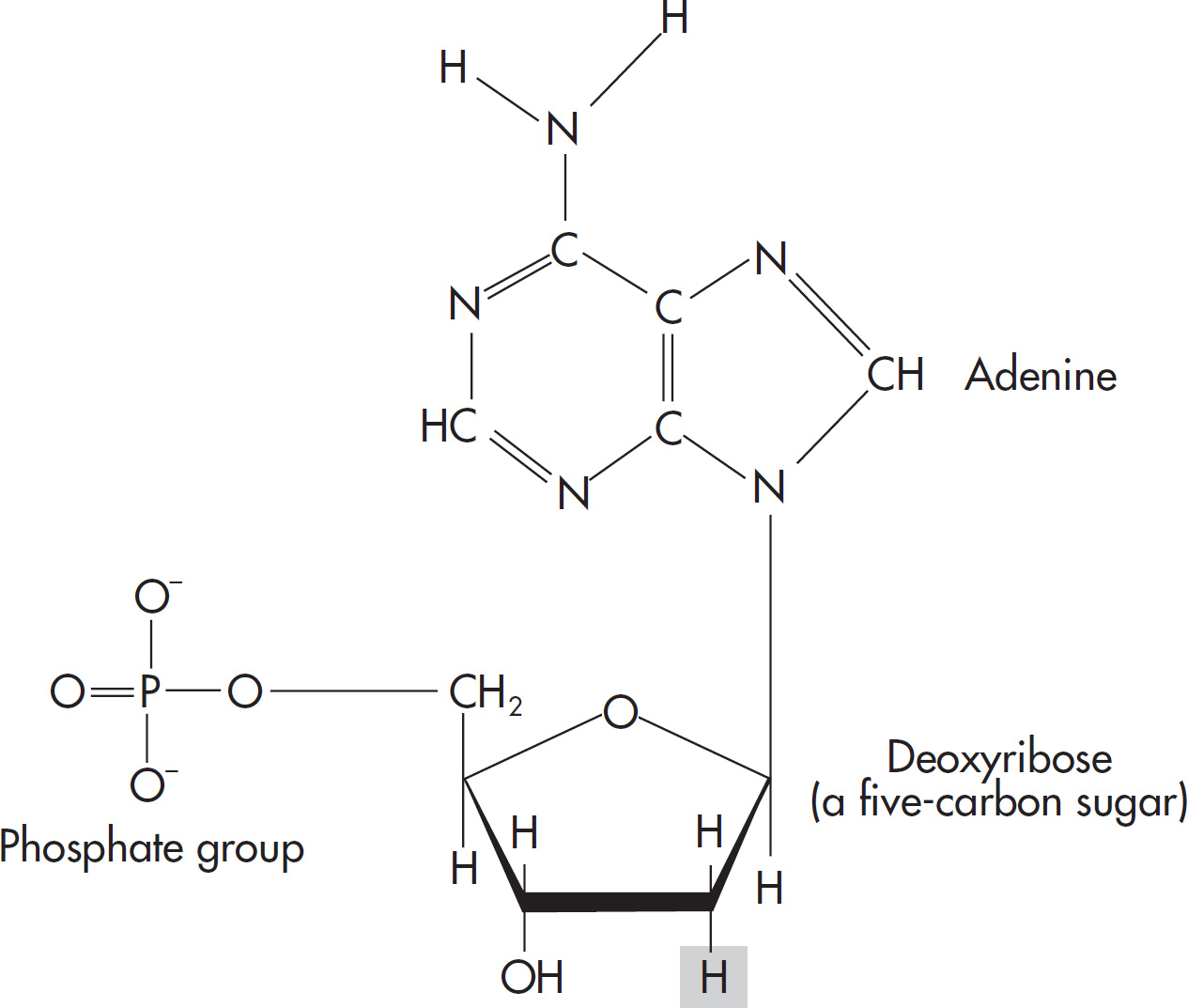
The name of the pentagon-shaped sugar in DNA is deoxyribose. Hence, the name deoxyribonucleic acid. Notice that the sugar is linked to two things: a phosphate and a nitrogenous base. A nucleotide can have one of four different nitrogenous bases:
• adenine—a purine (double-ringed nitrogenous base)
• guanine—a purine (double-ringed nitrogenous base)
• cytosine—a pyrimidine (single-ringed nitrogenous base)
• thymine—a pyrimidine (single-ringed nitrogenous base)
Any of these four bases can attach to the sugar. As we’ll soon see, this is extremely important when it comes to the message of the genetic code in DNA.
The nucleotides can link up in a long chain to form a single strand of DNA. Here’s a small section of a DNA strand.
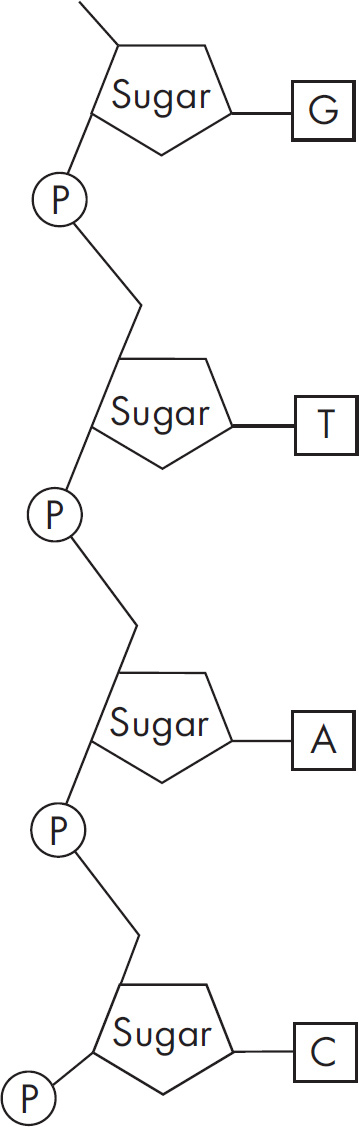
The nucleotides themselves are linked together by phosphodiester bonds between the sugars and the phosphates. This is called the sugar-phosphate backbone of DNA that serves as a scaffold for the bases.
Two DNA Strands
Each DNA molecule consists of two strands that wrap around each other to form a long, twisted ladder called a double helix. The structure of DNA was brilliantly deduced in 1953 by three scientists: Watson, Crick, and Franklin.
Now let’s look at the way two DNA strands get together. Again, think of DNA as a ladder. The sides of the ladder consist of alternating sugar and phosphate groups, while the rungs of the ladder consist of pairs of nitrogenous bases.
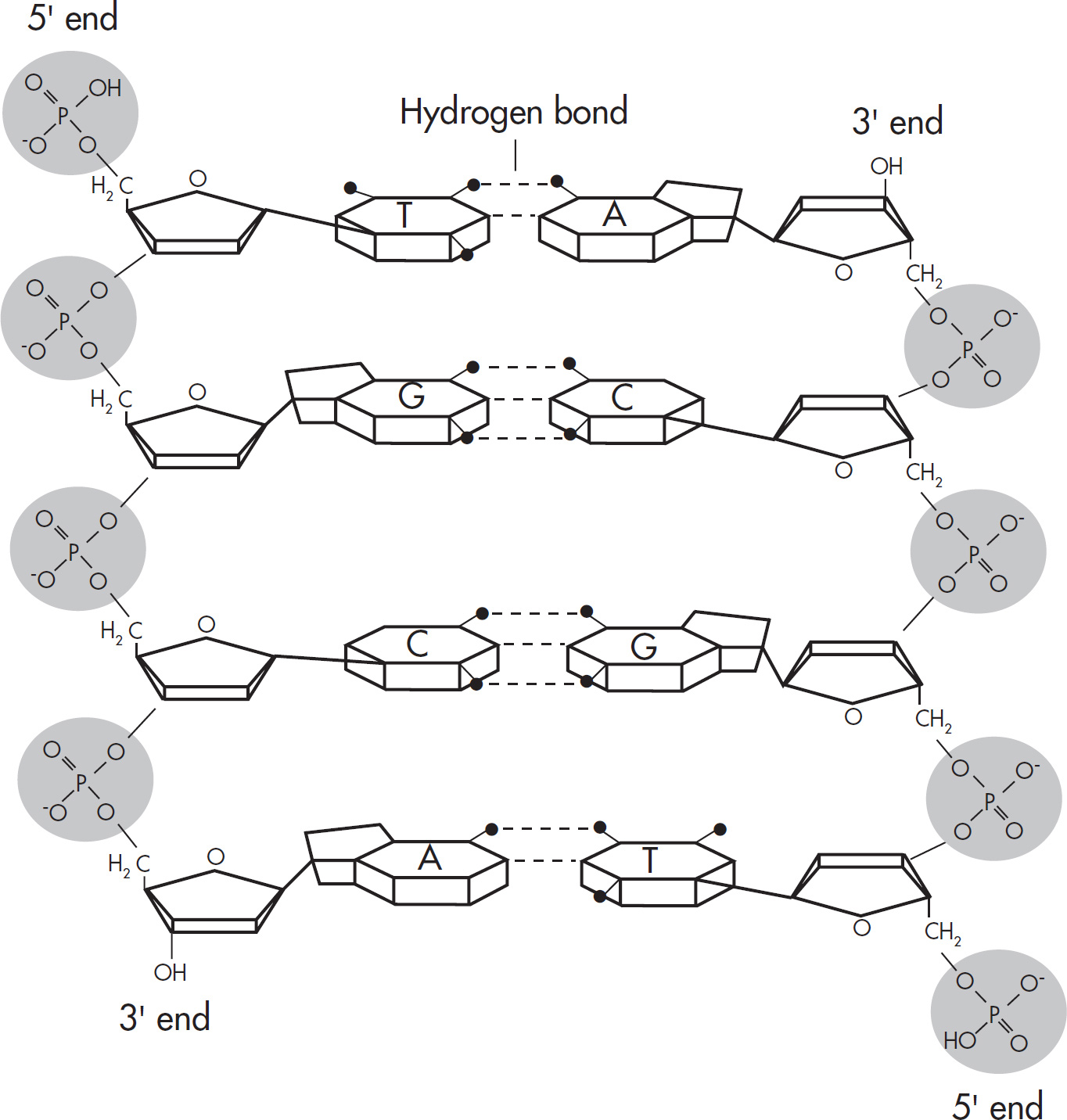
The nitrogenous bases pair up in a particular way. Adenine in one strand always binds to thymine (A–T or T–A) in the other strand. Similarly, guanine always binds to cytosine (G–C or C–G). This predictable matching of the bases is known as base pairing.
The two strands are said to be complementary. This means that if you know the sequence of bases in one strand, you’ll know the sequence of bases in the other strand. For example, if the base sequence in one DNA strand is A–T–C, the base sequence in the complementary strand will be T–A–G.
The two DNA strands run in opposite directions. You’ll notice from the figure above that each DNA strand has a 5’ end and a 3’ end, so-called for the carbon that ends the strand (i.e., the fifth carbon in the sugar ring is at the 5’ end, while the third carbon is at the 3’ end). The 5’ end has a phosphate group, and the 3’ end has an OH, or “hydroxyl,” group. The 5’ end of one strand is always opposite to the 3’ end of the other strand. The strands are therefore said to be antiparallel.
The DNA strands are linked by hydrogen bonds. Two hydrogen bonds hold adenine and thymine together, and three hydrogen bonds hold cytosine and guanine together.
Before we go any further, let’s review the base pairing in DNA:
• Adenine pairs up with thymine (A–T or T–A) by forming two hydrogen bonds.
• Cytosine pairs up with guanine (G–C or C–G) by forming three hydrogen bonds.
GENOME STRUCTURE
The order of the four base pairs in a DNA strand is the genetic code. Like a special alphabet in our cells, these four nucleotides spell out thousands of recipes. Each recipe is called a gene. The human genome has around 20,000 genes.
The recipes of the genes are spread out among the millions of nucleotides of DNA, and all of the DNA for a species is called its genome. Each separate chunk of DNA in a genome is called a chromosome.
Prokaryotes have one circular chromosome, and eukaryotes have linear chromosomes. In eukaryotes, the DNA is further structured, likely since linear chromosomes are more likely to get tangled. To keep it organized, the DNA is wrapped around proteins called histones, and then the histones are bunched together in groups called a nucleosome. Not all DNA is wound-up equally, because it must be unwound in order to be read. How tightly DNA is packaged depends on the section of DNA and also what is going on in the cell at that time. Humans have 23 different chromosomes and they have two copies of each, so each human cell has 46 linear chromosome segments in the nucleus. The DNA of a cell is contained in structures called chromosomes. Chromosomes consist of DNA wrapped around proteins called histones. When the genetic material is in a loose form in the nucleus it is called euchromatin, and its genes are active, or available for transcription. When the genetic material is fully condensed into coils, it is called heterochromatin, and its genes are generally inactive. Situated in the nucleus, chromosomes contain the recipe for all the processes necessary for life, including passing themselves and their information on to future generations. In this chapter, we’ll look at precisely how they accomplish this.
DNA REPLICATION
We said in the beginning of this chapter that DNA is the hereditary blueprint of the cell. By directing the manufacture of proteins, DNA serves as the cell’s blueprint. But how is DNA inherited? For the information in DNA to be passed on, it must first be copied. This copying of DNA is known as DNA replication.
Because the DNA molecule is twisted over on itself, the first step in replication is to unwind the double helix by breaking the hydrogen bonds. This is accomplished by an enzyme called helicase. The exposed DNA strands now form a y-shaped replication fork.
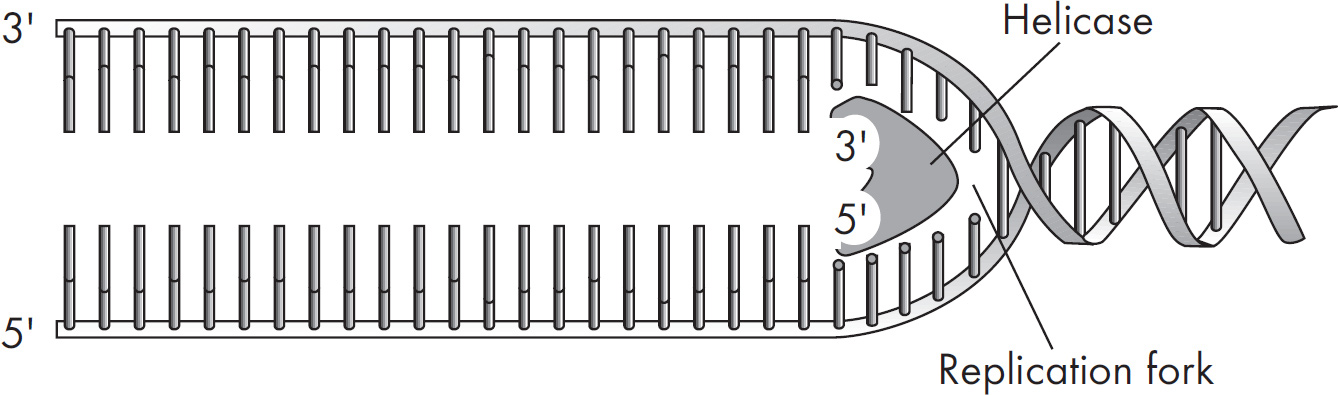
Now each strand can serve as a template for the synthesis of another strand. DNA replication begins at specific sites called origins of replication. Because the DNA helix twists and rotates during DNA replication, another class of enzymes, called DNA topoisomerases, cuts and rejoins the helix to prevent tangling. The enzyme that performs the actual addition of nucleotides alongside the naked strand is DNA polymerase. But DNA polymerase, oddly enough, can only add nucleotides to the 3’ end of an existing strand. Therefore, to start off replication at the 5’ end, an enzyme called RNA primase adds a short strand of RNA nucleotides called an RNA primer. The primer is later degraded by enzymes, and the space is filled with DNA.
One strand is called the leading strand, and it is made continuously. That is, the nucleotides are steadily added one after the other by DNA polymerase. The other strand—the lagging strand—is made discontinuously. Unlike the leading strand, the lagging strand is made in pieces of nucleotides known as Okazaki fragments. Why is the lagging strand made in small pieces?
Normally, nucleotides are added only in the 5’ to 3’ direction since nucleotides can only be added to the 3’ end of the growing chain. However, when the double-helix is “unzipped,” one of the two strands is oriented in the opposite direction—3’ to 5’. Because DNA polymerase doesn’t work in this direction, nucleotides—the Okazaki fragments—need to be added in pieces. You can see in the figure below that the leading strand is being created towards the opening of the helix and the helix continually opens ahead of it to accommodate it. The lagging strand is built in the opposite direction of the way the helix is opening, so it can only build until it hits a previously built stretch. Once the helix unwinds a bit more, it can build another okazaki fragment, and so on. These fragments are eventually linked together by the enzyme DNA ligase to produce a continuous strand. Finally, hydrogen bonds form between the new base pairs, leaving two identical copies of the original DNA molecule.
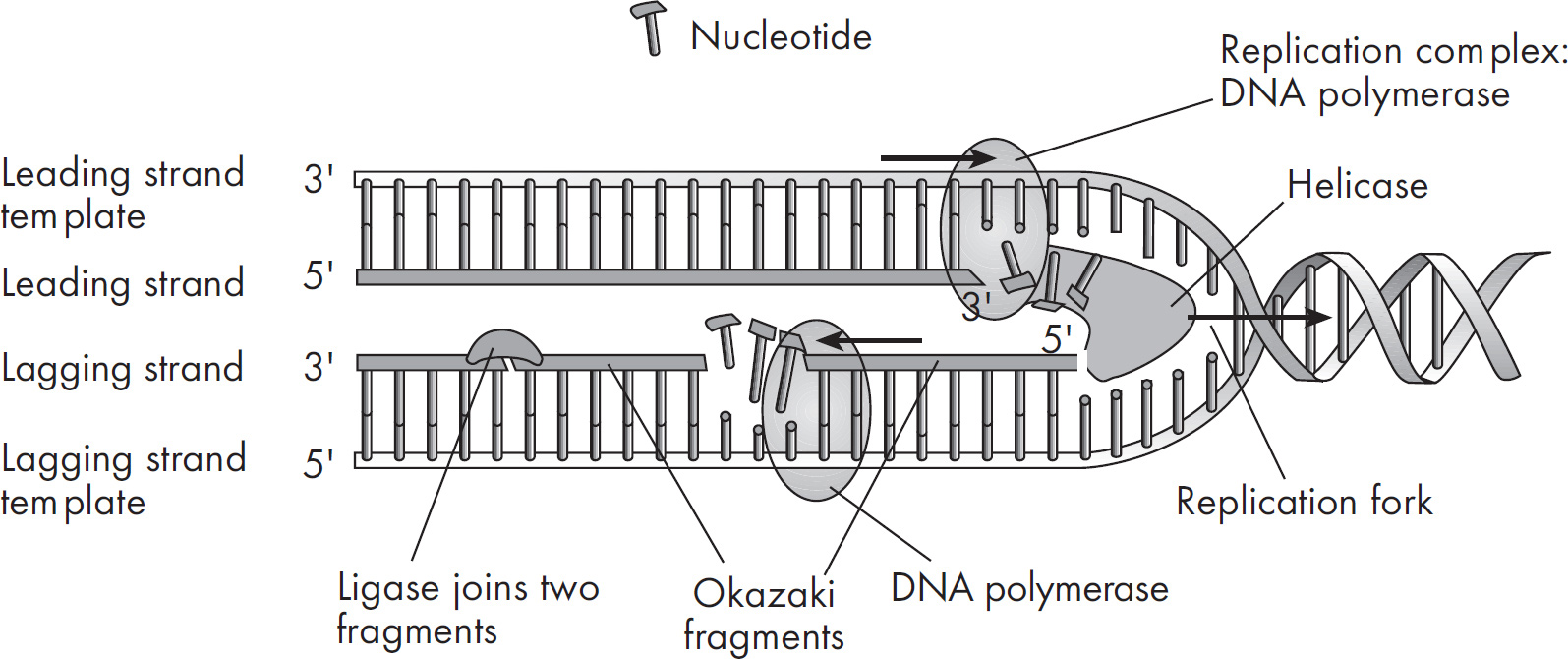
When DNA is replicated, we don’t end up with two entirely new molecules. Each new molecule has half of the original molecule. Because DNA replicates in a way that conserves half of the original molecule in each of the two new ones, it is said to be semiconservative.
An interesting problem is that a few bases at the very end of a DNA molecule cannot be replicated because the DNA polymerase needs to bind. This means that every time replication occurs, the chromosome loses a few base pairs. The genome has compensated for this over time, by putting bits of unimportant (or at least less important) DNA at the ends of a molecule. These ends are called telomeres. They get shorter and shorter over time.
Many enzymes and proteins are involved in DNA replication. The ones you’ll need to know for the AP Biology Exam are DNA helicase, DNA polymerase, DNA ligase, topoisomerase, and RNA primase:
• Helicase unwinds our double helix into two strands.
• Polymerase adds nucleotides to an existing strand.
• Ligase brings together the Okazaki fragments.
• Topoisomerase cuts and rejoins the helix.
• RNA primase catalyzes the synthesis of RNA primers.
Key History
The reason that we know that the DNA is the inheritable material is because of several key experiments. The first turning point was achieved by Avery, MacLeod, and McCarty. They isolated various cellular components from a dead virulent strain of bacteria. Then they followed up on a previous experiment by Griffiths and added each of these cellular components to a stain of living non-virulent bacteria. Only the DNA component of the deadly bacteria was able to change the second bacteria into a deadly strain capable of reproducing. This means that the DNA must be responsible for passing traits and it is inheritable.
The next important experiment was done by Hershey-Chase. They used bacterophages, which are a special type of virus. They labelled the protein parts of some viruses with radiolabelled sulfur and they labelled the DNA parts of other viruses with radiolabelled phosphorus. When the viruses infected bacteria, only the labelled DNA was inside, but they were still able to replicate and make progeny viruses. Thus they proved that only DNA is required to pass on information. The protein was not required.
Central Dogma
DNA’s main role is directing the manufacture of molecules that actually do the work in the body. DNA is just the recipe book, not the chef or the meal. When a DNA recipe is used, scientists say that that DNA is expressed.
The first step of DNA expression is to turn it into RNA. The RNA is then sent out into the cell to perform its job. Often, the job of an RNA is to get turned into a protein. Proteins are the biggest group of “worker-molecules,” and most expressed DNA turns into proteins. These proteins, in turn, regulate everything that occurs in the cell.
The process of making an RNA from DNA is called transcription, and the process of making a protein from an RNA is called translation. Transcription takes place in the nucleus (where the DNA is kept), and translation takes place in the cytoplasm.
Obviously, in prokaryotes, both occur in the cytoplasm because prokaryotes don’t contain a nucleus; this allows transcription and translation to occur at the same time.
The flow of genetic information, the “Central Dogma” of Biology, is therefore:
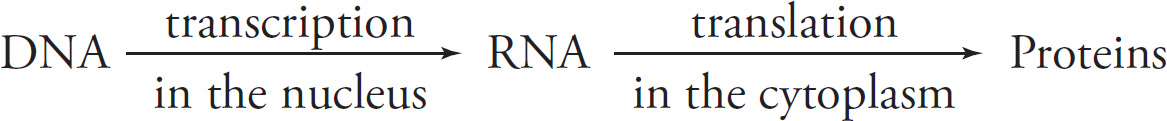
RNA
Before we discuss transcription of RNA, let’s talk about its structure. Although RNA is also made up of nucleotides, it differs from DNA in three ways.
1. RNA is single-stranded, not double-stranded.
2. The five-carbon sugar in RNA is ribose instead of deoxyribose.
3. The RNA nitrogenous bases are adenine, guanine, cytosine, and a different base called uracil. Uracil replaces thymine as adenine’s partner.
Here’s a table to compare DNA and RNA. Keep these differences in mind—the testing board loves to test you on them.
| DIFFERENCES BETWEEN DNA AND RNA | ||
| DNA (double-stranded) | RNA (single-stranded) | |
| Sugar: | deoxyribose | ribose |
| Bases: | adenine guanine cytosine thymine |
adenine guanine cytosine uracil |
There are three main types of RNA: messenger RNA (mRNA), ribosomal RNA (rRNA), and transfer RNA (tRNA). All three types of RNA are key players in the synthesis of proteins:
• Messenger RNA (mRNA) is a temporary RNA version of a DNA recipe that gets sent to the ribosome.
• Ribosomal RNA (rRNA), which is produced in the nucleolus, makes up part of the ribosomes. You’ll recall from our discussion of the cell in Chapter 5 that the ribosomes are the sites of protein synthesis. We’ll see how they function a little later on.
• Transfer RNA (tRNA) shuttles amino acids to the ribosomes. It is responsible for bringing the appropriate amino acids into place at the appropriate time. It does this by reading the message carried by the mRNA.
There is also another class of RNA, called interfering RNAs, or RNAi. These are small snippets of RNA that are naturally made in the body or intentionally created by humans. These interfering RNAs, called siRNA and miRNA, can bind to specific sequences or RNA and mark them for destruction. This will be discussed more later in this chapter. Now that we know about the different types of RNA, let’s see how they direct the synthesis of proteins.
Transcription
Transcription involves making an RNA copy of a bit of DNA code. The initial steps in transcription are similar to the initial steps in DNA replication. The obvious difference is that, whereas in replication we end up with a complete copy of the cell’s DNA, in transcription we end up with only a specific section copied into an mRNA. This is because only the bit of DNA that needs to be expressed will be transcribed. If you wanted to make a cake, but your cookbook couldn’t leave your vault (AKA the nucleus), you wouldn’t copy the entire cookbook! You would only copy down the recipe for the cake.
Since each recipe is a gene, transcription occurs as-needed on a gene-by-gene basis.
The exception to this is prokaryotes because they will transcribe a recipe that can be used to make several proteins. This is called a polycistronic transcript. Eukaryotes tend to have one gene that gets transcribed to one mRNA and translated into one protein. Our transcripts are monocistronic.
Transcription involves three phases: initiation, elongation, and termination. As in DNA replication, the first initiation step in transcription is to unwind and unzip the DNA strands using helicase. Transcription begins at special sequences of the DNA strand called promoters. You can think of a promoter as a docking site or a runway. We will talk about how promoters are involved in regulating transcription later in the chapter.
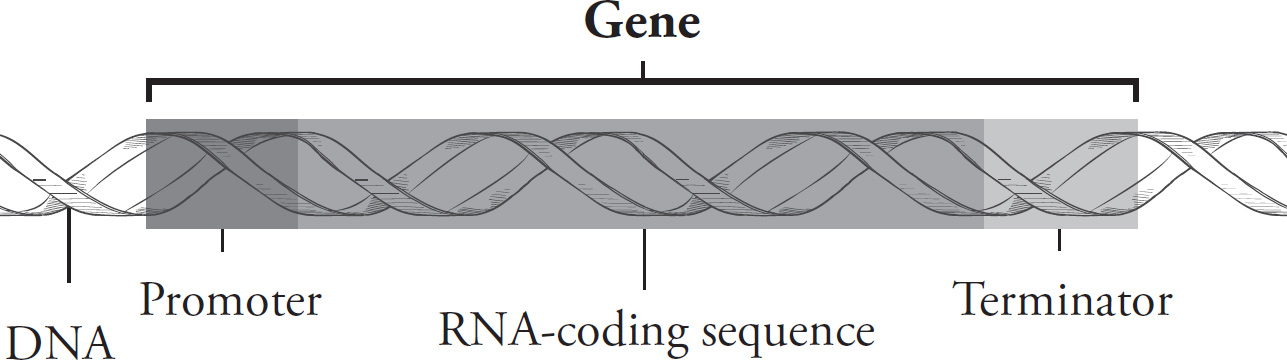
Because RNA is single-stranded, we have to copy only one of the two DNA strands. The strand that serves as the template is known as the antisense strand, the non-coding strand, or the template strand. The other strand that lies dormant is the sense strand, or the coding strand.
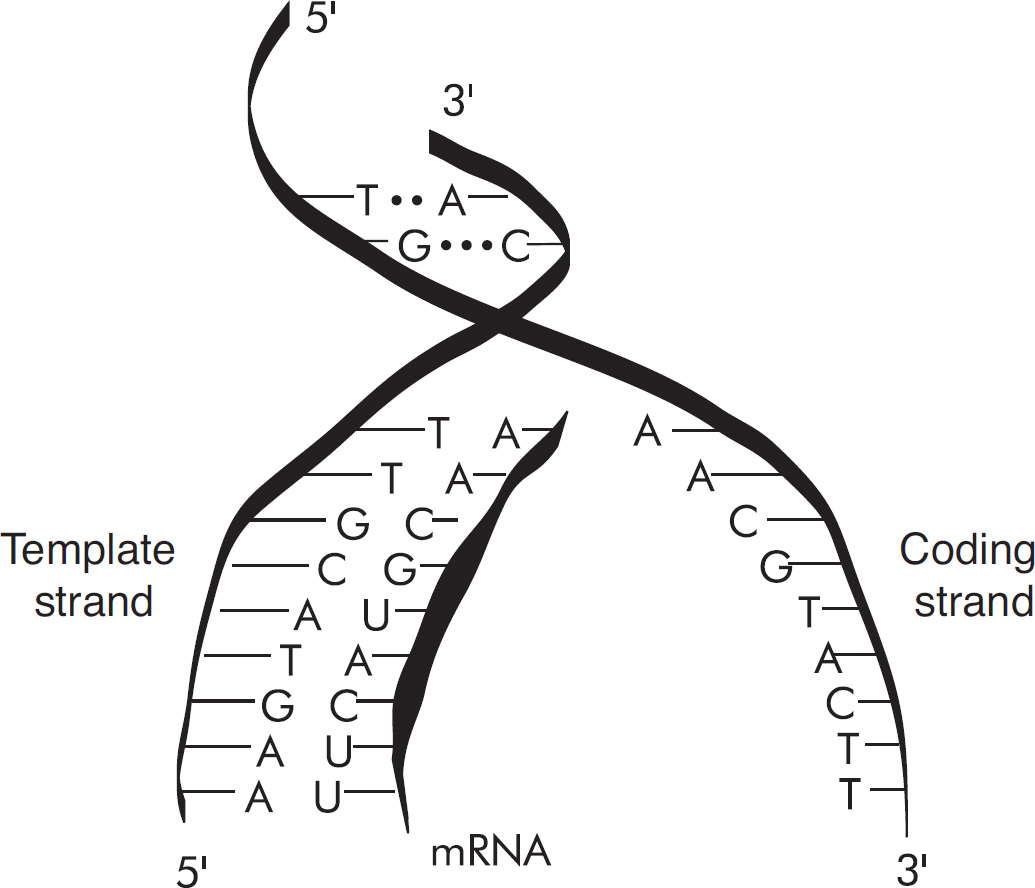
Just as DNA polymerase builds DNA, RNA polymerase builds RNA, and just like DNA polymerase, RNA polymerase only adds nucleotides to the 3’ side (5’ to 3’). This means that the RNA polymerase must bind to the 3’ end of the template strand (which would be the 5’ end of future mRNA).
RNA polymerase doesn’t need a primer, so it can just start transcribing the DNA right off the bat. The promoter region is considered to be “upstream” of the actual coding part of the gene. This way the polymerase can get set up before the bases it needs to transcribe, like a staging area before an official parade starting point. The official starting point is called the start site.
When transcription begins, RNA polymerase travels along and builds an RNA that is complementary to the template strand of DNA. It is just like DNA replication except when DNA has an adenine, the RNA can’t add a thymine since RNA doesn’t have thymine. Instead the RNA adds a uracil to pair with adenine.
Once the mRNA finishes adding on nucleotides and reaches the termination sequence, it separates from the DNA template, completing the process of transcription. The new RNA has now transcribed, or “copied,” the sequence of nucleotide bases directly from the exposed DNA strand. Note that the freshly synthesized RNA is complementary to the template strand, but it is identical to the coding strand (with the substitution of uracil for thymine).
RNA Processing
In prokaryotes, the mRNA is now complete, but in eukaryotes the RNA must be processed before it can leave the nucleus.
In eukaryotes, the freshly transcribed RNA in called an hnRNA (heterogeneous nuclear RNA) and it contains both coding regions and noncoding regions. The regions that express the code that will be turned into protein are exons. The noncoding regions in the mRNA are introns.
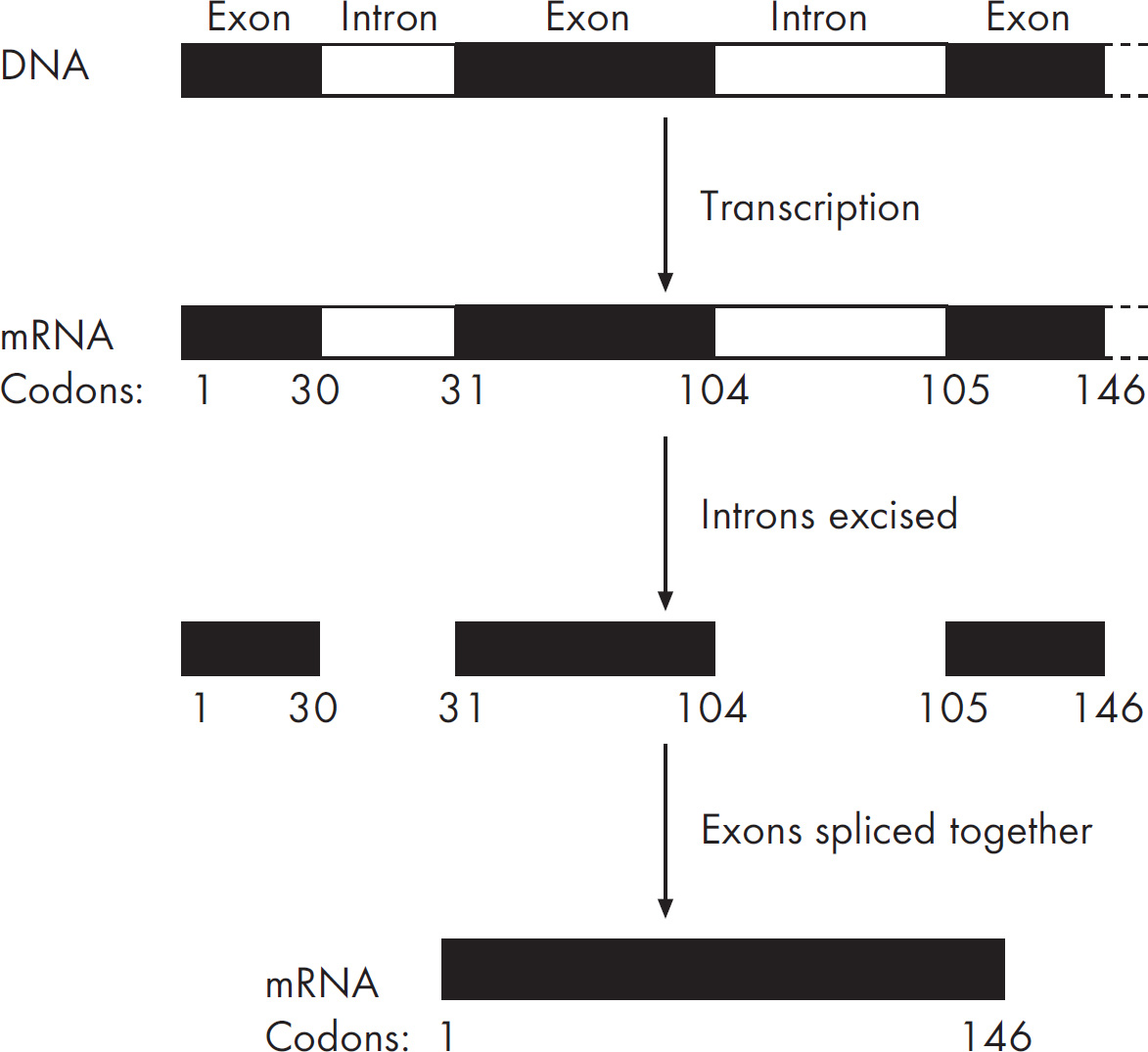
The introns—the intervening sequences—must be removed before the mRNA leaves the nucleus. This is called splicing and it is accomplished by an RNA-protein complex called a spliceosome.
In addition to splicing, a poly(A) tail is added to the 3’ end and a 5’ GTP cap is added to the 5’ end. This process produces a final mRNA that is shorter than the transcribed RNA.
Translation
Translation is the process of turning an mRNA into a protein. Remember, each protein is made of amino acids. The order of the mRNA nucleotides will be read in the ribosome in groups of three. Three nucleotides is called a codon. Each codon corresponds to a particular amino acid. The genetic code is redundant, meaning that certain amino acids are specified by more than one codon.
The mRNA attaches to the ribosome to initiate translation and “waits” for the appropriate amino acids to come to the ribosome. That’s where tRNA comes in. A tRNA molecule has a unique three-dimensional structure that resembles a four-leaf clover:
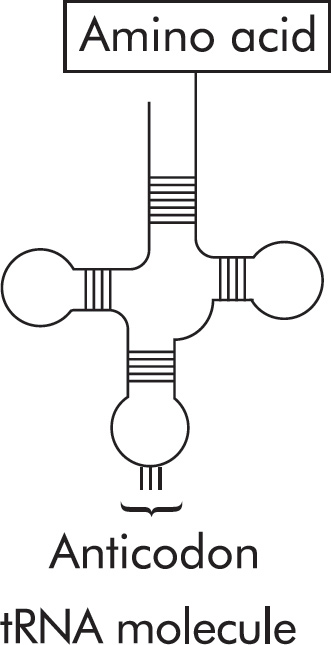
One end of the tRNA carries an amino acid. The other end, called an anticodon, has three nitrogenous bases that can complementarily base pair with the codon in the mRNA. Usually, the normal rules of base pairing are set in stone, but tRNA anticodons can be a bit flexible when they bind with a codon on an mRNA, especially the 3 nucleotide in an anticodon. The third position is said to experience wobble pairing. Things that don’t normally bind will pair up, like guanine and uracil.
Transfer RNAs are the “go-betweens” in protein synthesis. Each tRNA becomes charged and enzymatically attaches to an amino acid in the cell’s cytoplasm and “shuttles” it to the ribosome. The charging enzymes involved in forming the bond between the amino acid and tRNA require ATP.
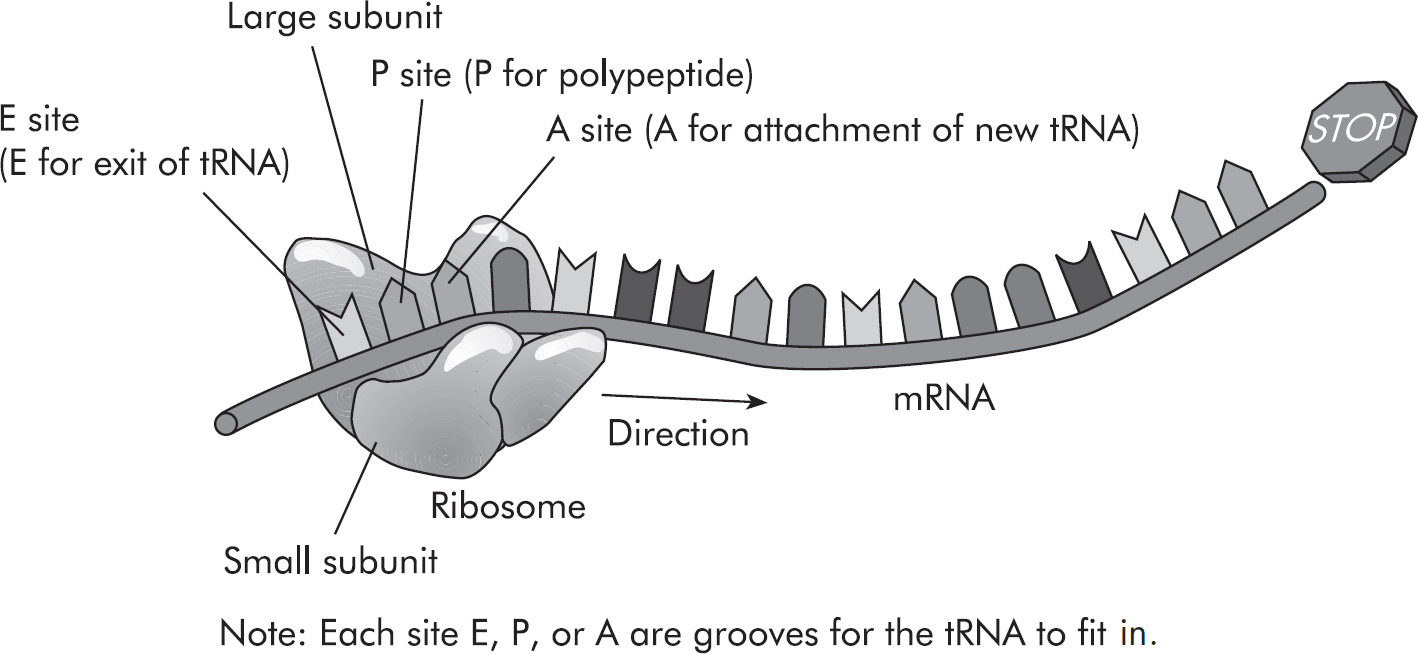
Translation also involves three phases: initiation, elongation, and termination. Initiation begins when a ribosome attaches to the mRNA.
What does the ribosome do? It helps the process along by holding everything in place while the tRNAs assist in assembling polypeptides.
Initiation
Ribosomes contain three binding sites: an A site, a P site, and an E site. The mRNA will shuffle through from A to P to E. As the mRNA codons are read, the polypeptide will be built. In all organisms the start codon for the initiation of protein synthesis is A–U–G, which codes for the amino acid methionine. Proteins can have AUGs in the rest of the protein that code for methionine as well, but the special first AUG of an mRNA is the one that will kick off translation. The tRNA with the complementary anticodon, U–A–C, is methionine’s personal shuttle; when the AUG is read on the mRNA, it delivers the methionine to the ribosome.
Elongation
The addition of amino acids is called elongation. Remember that the mRNA contains many codons, or “triplets,” of nucleotide bases. As each amino acid is brought to the mRNA, it is linked to its neighboring amino acid by a peptide bond. When many amino acids link up, a polypeptide is formed.
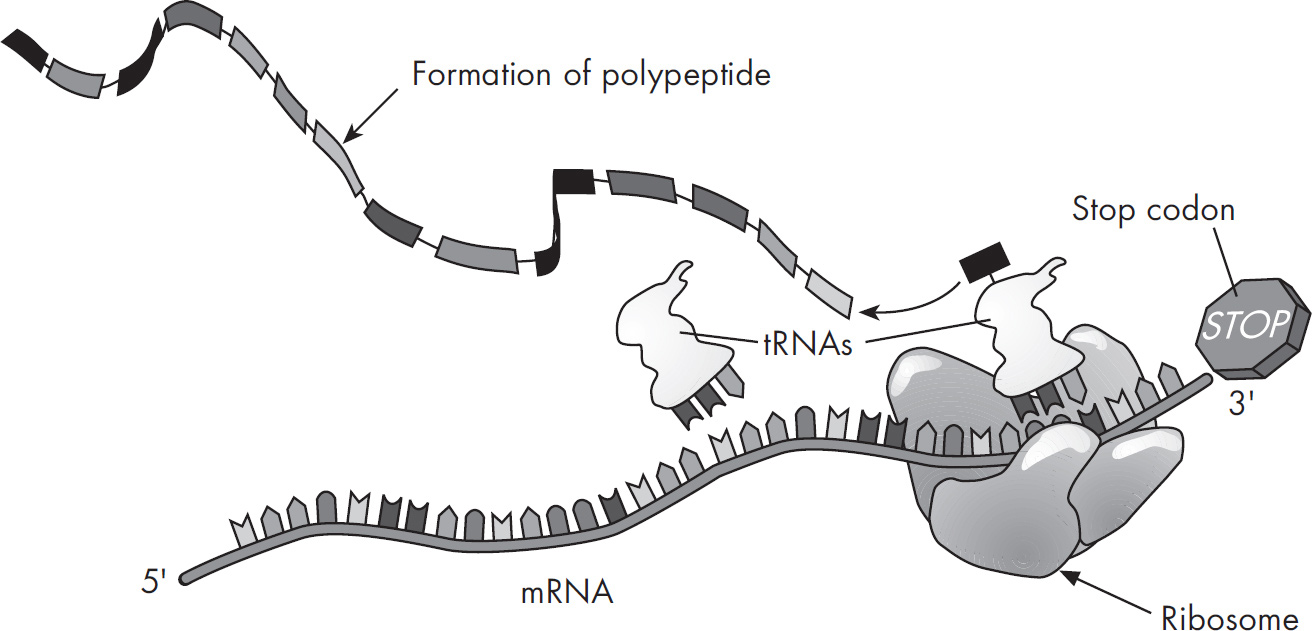
Termination
How does this process know when to stop? The synthesis of a polypeptide is ended by stop codons. A codon doesn’t always code for an amino acid; there are three that serve as a stop codon. Termination occurs when the ribosome runs into one of these three stop codons.
How about a little review?
• In transcription, mRNA is created from a particular gene segment of DNA.
• In eukaryotes, the mRNA is “processed” by having its introns, or noncoding sequences, removed.
• Now, ready to be translated, mRNA proceeds to the ribosome.
• Free-floating amino acids are picked up by tRNA and shuttled over to the ribosome, where mRNA awaits.
• In translation, the anticodon of a tRNA molecule carrying the appropriate amino acid base pairs with the codon on the mRNA.
• As new tRNA molecules match up to new codons, the ribosome holds them in place, allowing peptide bonds to form between the amino acids.
• The newly formed polypeptide grows until a stop codon is reached.
• The polypeptide or protein is released into the cell.
Gene Regulation
What controls gene transcription, and how does an organism express only certain genes? Regulation of gene expression can occur at different times. The largest point is before transcription, or pre-transcriptional regulation. It can also occur post-transcriptionally or post-translationally.
The start of transcription requires the DNA to be unwound and RNA polymerase to bind at the promoter. The process is usually a bit more complicated than that because transcription factors can either encourage or inhibit this from happening. This is often accomplished by making it easier or more difficult for the RNA polymerase to bind or to move to the start site.
The following examples are famous models of regulation. You do not need to memorize them, but you should be familiar with the big picture of how things can come together to regulate transcription. Most of what we know about gene regulation comes from our studies of E. coli. In bacteria, the region of bacterial DNA that regulates gene expression is called an operon. One of the best-understood operons is the lac operon, which controls expression of the enzymes that break down lactose.
The operon consists of four major parts: structural genes, the regulatory gene, the promoter gene, and the operator:
• Structural genes are genes that code for enzymes needed in a chemical reaction. These genes will be transcribed at the same time to produce particular enzymes. In the lac operon, three enzymes (beta galactosidase, galactose permease, and thiogalactoside transacetylase) involved in digesting lactose are coded for.
• The promoter gene is the region where the RNA polymerase binds to begin transcription.
• The operator is a region that controls whether transcription will occur; this is where the repressor binds.
• The regulatory gene codes for a specific regulatory protein called the repressor. The repressor is capable of attaching to the operator and blocking transcription. If the repressor binds to the operator, transcription will not occur. On the other hand, if the repressor does not bind to the operator, RNA polymerase moves right along the operator and transcription occurs. In the lac operon, the inducer, lactose, binds to the repressor, causing it to fall off the operator, and “turns on” transcription.
The following two diagrams show the lac operon when lactose is absent (A) and when lactose is present (B).
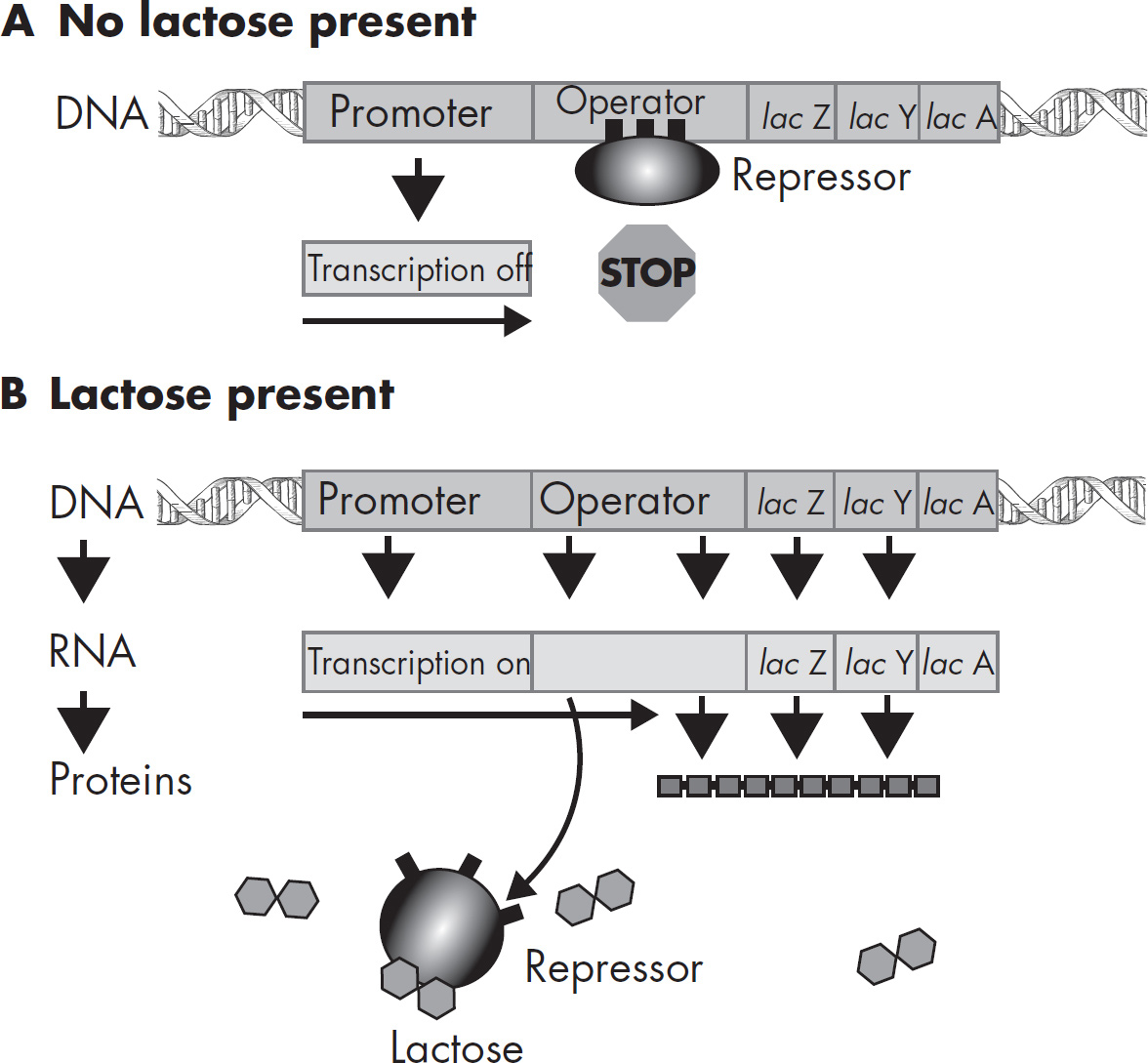
Other operons, such as the trp operon, operate in a similar manner except that this mechanism is continually “turned on” and is only “turned off” in the presence of high levels of the amino acid, tryptophan. Tryptophan is a product of the pathway that codes for the trp operon. When tryptophan combines with the trp repressor protein, it causes the repressor to bind to the operator, which turns the operon “off,” thereby blocking transcription. In other words, a high level of tryptophan acts to repress the further synthesis of tryptophan.
The following two diagrams show the trp operon when tryptophan is absent (C) and when tryptophan is present (D).
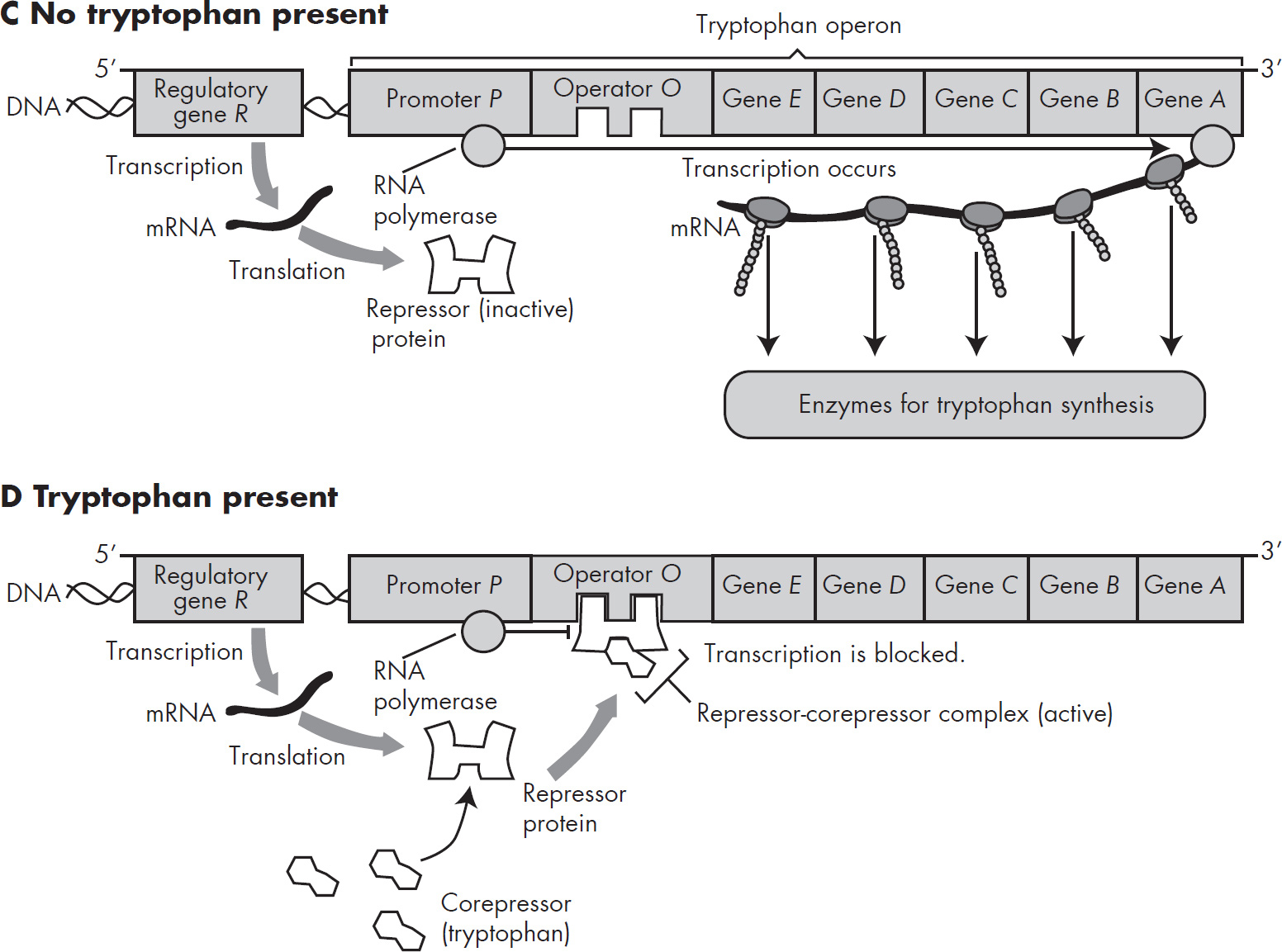
Post-transcriptional regulation occurs when the cell creates an RNA, but then decides that it should not be translated into a protein. This is where RNAi comes into play. RNAi molecules can bind to an RNA via complementary base pairing. This creates a double-stranded RNA (remember RNA is usually single stranded). When a double-stranded RNA is formed, this signals to some special destruction machinery that the RNA should be destroyed. This would prevent it from going on to be translated.
Post-translational regulation can also occur. This would be if a cell has already made a protein, but it doesn’t need to use the protein yet. This is especially common regulation for enzymes because when the cell needs them, it needs them ASAP. It is easier to make them ahead of time and then just turn them on or off as needed. This can involve binding with other proteins, phosphorylation, pH, cleavage, etc. Remember, even if a protein is made, it might need other things to be made (or not made) in order for the protein to be functional.
MUTATIONS
A mutation is a error in the genetic code. Mutations can occur because DNA is damaged and cannot be repaired or because DNA damage is repaired incorrectly. Damage can be caused by chemicals or radiation. It can also occur when a DNA polymerase or an RNA polymerase makes a mistake. DNA polymerases have proofreading abilities, but RNA polymerases do not. This is because RNA is only a temporary molecule and if a mistake is made, it is not as problematic. DNA is important now and in the future since it is passed on from cell to cell in somatic cells and from parent to offspring. If the mistake occurs in a germline cell, that will become a gamete. Mistakes in DNA can last forever.
It is important to know that just having an error in the DNA is not a problem unless that gene is expressed AND the error causes a change in the gene product (RNA or protein). Think of it like an error in a recipe in a cookbook; an error won’t harm anyone unless the recipe is actually made AND the error causes a big change in the recipe. As we will see, a mutation can cause different types of effects.
Base Substitution
Base substitution (point) mutations result when a single nucleotide base is substituted for another. Point mutations can be:
• nonsense mutations—these cause the original codon to become a stop codon, which results in early termination of protein synthesis
• missense mutations—these cause the original codon to be altered and produce a different amino acid
• silent mutations—these happen when a codon that codes for the same amino acid is created and therefore does not change the corresponding protein sequence
Gene Rearrangements
Gene rearrangements involve DNA sequences that have deletions, duplications, inversions, and translocations.
1. Insertions and deletions result in the gain or loss, respectively, of DNA or a gene. They can either involve the addition (insertion) or removal (deletion) of as little as a single base or much larger sequences of DNA. Insertions and deletions may have devastating consequences during protein translation because the introduction or deletion of bases often results in a change in the sequence of codons used by the ribosome (called a frameshift mutation) to synthesize a polyprotein.
Example:

In this example, the insertion of an additional cytosine nucleotide resulted in a frameshift and a premature stop codon.
2. Duplications can result in an extra copy of genes and are usually caused by unequal crossing over during meiosis or chromosome rearrangements. This may result in new traits because one copy can maintain the gene’s original function and one copy may evolve a new function.
3. Inversions can result when changes occur in the orientation of chromosomal regions. This may cause harmful effects if the inversion involves a gene or an important regulatory sequence.
4. Translocations occur when a portion of two different chromosomes (or a single chromosome in two different places) breaks and rejoins in a way that causes the DNA sequence or gene to be lost, repeated, or interrupted.
BIOTECHNOLOGY
Recombinant DNA
Scientists have learned how to harness the central dogma in order to research and cure diseases. Recombinant DNA is generated by combining DNA from multiple sources to create a unique DNA molecule that is not found in nature. A common application of recombinant DNA technology is the introduction of a eukaryotic gene of interest (such as insulin) into a bacterium for production. Bacteria can be hijacked and put to work as little protein factories. This branch of technology that produces new organisms or products by transferring genes between cells is called genetic engineering.
Polymerase Chain Reaction (PCR)
A few years ago, it would take weeks of tedious experiments to identify and study specific genes. Today, thanks to polymerase chain reaction (PCR), we are able to make billions of identical copies of genes within a few hours. To do PCR, the process of DNA replication is slightly modified. In a small PCR tube, DNA, primers, a powerful DNA polymerase, and lots of DNA nucleotides (A’s, C’s, G’s, and T’s) are mixed together.
In a PCR machine, or thermocycler, the tube is heated, cooled, and warmed many times. Each time the machine is heated, the hydrogen bonds break, separating the double-stranded DNA. As it is cooled, the primers bind to the sequence flanking the region of the DNA we want to copy. When it is warmed, the polymerase binds to the primers on each strand and adds nucleotides on each template strand. After this first cycle is finished, there are two identical double-stranded DNA molecules. When the second cycle is completed, these two double-stranded DNA segments will have been copied into four. The process repeats itself over and over, creating as much DNA as needed.
Transformation
Insulin, the protein hormone that lowers blood sugar levels, can now be made for medical purposes by bacteria. Yes, bacteria can be induced to use the universal DNA code to transcribe and translate a human gene! The process of giving bacteria foreign DNA is called transformation.
Genes of interest are first placed into a small circular DNA molecule called a plasmid. Often, a plasmid is pre-designed to have some special helpful genes in it, and together, these are called a vector. Plasmid vectors often contain genes for antibiotic resistance and restriction sites. Plasmids and the gene of interest are cut with the same restriction enzyme (restriction enzymes cut DNA at very specific pre-determined sequences), creating compatible sticky ends. When placed together, the gene is inserted into the plasmid creating recombinant DNA.
The bacteria are then transformed (given the recombinant DNA to take up). In most AP Biology courses, this is done by the heat shock method.
Not all the bacteria will be transformed, but the ones that did not take up the plasmid are not needed. This is where the antibiotic resistance gene on the vector comes in. By growing all the bacteria in the presence of an antibiotic, only those with the resistance gene (AKA those that have been transformed) will survive.
This laboratory technique has not only been used to safely mass-produce important proteins used for medicine, such as insulin, it also plays an important role in the study of gene expression. A technique similar to transformation is transfection, which is putting a plasmid into a eukaryotic cell, rather than a bacteria cell. This is a bit trickier than transformation. Because eukaryotic cells do not grow like bacteria do, transfection is not as useful for making large quantities of something, but sometimes it is important to use eukaryotic cells.
Gel Electrophoresis
An important part of DNA technology is the ability to observe differences in different preparations of DNA.
The DNA fragments can be separated according to their molecular weight using gel electrophoresis. Because DNA and RNA are negatively charged, they migrate through the gel toward the positive pole of the electrical field. The smaller the fragments, the faster they move through the gel. Restriction enzymes are also used to create a molecular fingerprint. The patterns created after cutting with restriction enzymes are unique for each person because each person’s DNA is slightly different. Some people might have sequences that are cut many times, resulting in many tiny fragments, and others might have sequences that are cut infrequently, producing a few large fragments. When restriction fragments between individuals of the same species are compared, the fragments differ in length because of polymorphisms, which are slight differences in DNA sequences. These fragments are called restriction fragment length polymorphisms, or RFLPs. In DNA fingerprinting, RFLPs produced from DNA left at a crime scene are compared to RFLPs from the DNA of suspects.
KEY TERMS
nucleotide
five-carbon sugar
phosphate
nitrogenous base
deoxyribose
adenine
guanine
cytosine
thymine
phosphodiester bonds
double helix
Watson, Crick, and Franklin
base pairing
complementary
antiparallel
hydrogen bonds
gene
genome
chromosome
nucleosome
euchromatin
heterochromatin
DNA replication
helicase
replication fork
origins of replication
topoisomerase
DNA polymerase
RNA primase
RNA primer
leading strand
lagging strand
Okazaki fragments
DNA ligase
semiconservative replication
telomeres
Avery, MacLeod, and McCarty
Hershey-Chase
transcription
translation
Central Dogma
ribose
uracil
messenger RNA (mRNA)
ribosomal RNA (rRNA)
transfer RNA (tRNA)
RNA interference (RNAi)/silencing RNA (siRNA)
polycistronic
monocistronic
promoter
antisense/non-coding/template strand
sense/coding strand
RNA polymerase
start site
exons
introns
spliceosome
poly(A) tail
5’ GTP cap
codon
anticodon
wobble pairing
Initiation
Elongation
Termination
A site, P site, E site
start codon
stop codons
pre-transcriptional regulation
transcription factors
operon
structural genes
promoter genes
operator site
regulatory gene
post-transcriptional regulation
post-translational regulation
mutation
base substitution
nonsense mutation
missense mutation
silent mutation
insertions
deletions
frameshift mutation
duplications
inversions
translocations
recombinant DNA
genetic engineering
polymerase chain reaction (PCR)
transformation
plasmid
vector
restriction enzyme
sticky end
transfection
gel electrophoresis
restriction fragment length polymorphism (RFLPs)
DNA fingerprinting
Summary
 DNA is the genetic material of the cell.
DNA is the genetic material of the cell.
 DNA structure is composed of two antiparallel complementary strands, including:
DNA structure is composed of two antiparallel complementary strands, including:
• Deoxyribose sugar and phosphate backbone with nitrogenous bases and hydrogen bonded together in the middle. The two strands are oriented antiparallel with 3’ and 5’ ends and twist into a double helix.
• Adenine always pairs with thymine, and guanine always pairs with cytosine.
 All the DNA in a cell is the genome. It is divided up into chromosomes, which are divided up into genes that each encode a particular genetic recipe.
All the DNA in a cell is the genome. It is divided up into chromosomes, which are divided up into genes that each encode a particular genetic recipe.
 DNA replication provides two copies of the DNA for cell division, which involves the following steps:
DNA replication provides two copies of the DNA for cell division, which involves the following steps:
• Helicase begins replication by unwinding DNA and separating the strands at the origin of replication.
• Topoisomerases reduce supercoiling ahead of the replication fork.
• RNA primase places an RNA primer down on the template strands.
• DNA polymerase reads the DNA base pairs of a strand as a template and lays down complementary nucleotides to generate a new complementary partner strand.
• Replication proceeds continuously on the leading strand.
• The lagging strand forms with Okazaki fragments, which must be joined later by DNA ligase.
 Transcription involves building RNA from DNA.
Transcription involves building RNA from DNA.
• RNA polymerase reads one of the DNA strands (the anti-sense/non-coding/template strand) and adds RNA nucleotides to generate a single strand of mRNA that will be identical to the other strand of DNA (the coding/sense strand).
• mRNA, rRNA, tRNA, and RNAi are all types of RNA that can be created.
• mRNA gets processed in eukaryotic cells: introns are removed (spliced out), and a 5’ GTP cap and a 3’ poly(A) tail are added.
 Translation occurs in the cytoplasm as the mRNA carries the message to the ribosome.
Translation occurs in the cytoplasm as the mRNA carries the message to the ribosome.
• The mRNA passes through three sites on the ribosome, the A-site, the P-site, and the E-site.
• The mRNA is read in triplets of three nucleotides, called codons. Each tRNA has a region called the anticodon that is complementary to a codon. Sometimes the pairing is not the normal base-pairing. This is called wobble pairing.
• When a tRNA binds, it brings the corresponding amino acid to add to the growing polypeptide.
 Gene expression is regulated primarily by transcription factors influencing transcription (pre-transcriptional regulation), RNAi after transcription (post-transcriptional regulation), and by various things after translation (post-translational regulation). This regulation is dynamic and can either increase or decrease gene expression, RNA levels, and protein levels according to the needs of the cell.
Gene expression is regulated primarily by transcription factors influencing transcription (pre-transcriptional regulation), RNAi after transcription (post-transcriptional regulation), and by various things after translation (post-translational regulation). This regulation is dynamic and can either increase or decrease gene expression, RNA levels, and protein levels according to the needs of the cell.
 Mutations can result from changes in the DNA message, or the mRNA message.
Mutations can result from changes in the DNA message, or the mRNA message.
 Mutations can be small (single nucleotide swaps, additions, or deletions) or large (big chunks or even entire chromosomes that are swapped, duplicated, or deleted).
Mutations can be small (single nucleotide swaps, additions, or deletions) or large (big chunks or even entire chromosomes that are swapped, duplicated, or deleted).
 Some examples of biotechnology are:
Some examples of biotechnology are:
• recombinant DNA
• polymerase chain reaction (PCR)
• transformation of bacteria
• gel electrophoresis
Chapter 7 Drill
Answers and explanations can be found in Chapter 16.
1. A geneticist has discovered a yeast cell, which encodes a DNA polymerase that may add nucleotides in both the 5’ to 3’ and 3’ to 5’ directions. Which of the following structures would this cell not likely generate during DNA replication?
(A) RNA primers
(B) Okazaki fragments
(C) Replication fork
(D) Nicked DNA by topoisomerases
2. A eukaryotic gene, which does not normally undergo splicing, was exposed to benzopyrene, a known carcinogen and mutagen. Following exposure, the protein encoded by the gene was shorter than before exposure. Which of the following types of genetic rearrangements or mutations was likely introduced by the mutagen?
(A) Silent mutation
(B) Missense mutation
(C) Nonsense mutation
(D) Duplication
3. DNA replication occurs through a complex series of steps involving several enzymes. Which of the following represents the correct order beginning with the earliest activity of enzymes involved in DNA replication?
(A) Helicase, ligase, RNA primase, DNA polymerase
(B) DNA polymerase, RNA primase, helicase, ligase
(C) RNA primase, DNA polymerase, ligase, helicase
(D) Helicase, RNA primase, DNA polymerase, ligase
4. If a messenger RNA codon is UAC, which of the following would be the complementary anticodon triplet in the transfer RNA?
(A) ATG
(B) AUC
(C) AUG
(D) ATT
5. During post-translational modification, the polypeptide from a eukaryotic cell typically undergoes substantial alteration that results in
(A) excision of introns
(B) addition of a poly(A) tail
(C) formation of peptide bonds
(D) a change in the overall conformation of a polypeptide
6. Which of the following represents the maximum number of amino acids that could be incorporated into a polypeptide encoded by 21 nucleotides of messenger RNA?
(A) 3
(B) 7
(C) 21
(D) 42
7. A researcher uses molecular biology techniques to insert a human lysosomal membrane protein into bacterial cells to produce large quantities of this protein for later study. However, only small quantities of this protein result in these cells. What is a possible explanation for this result?
(A) The membrane protein requires processing in the ER and Golgi, which are missing in the bacterial cells.
(B) Bacteria do not make membrane proteins.
(C) Bacteria do not use different transcription factors than humans, so the gene was not expressed.
(D) Bacteria do not have enough tRNAs to make this protein sequence.
8. BamHI is a restriction enzyme derived from Bacillus amyloliquefaciens that recognizes short palindromic sequences in DNA. When the enzyme recognizes these sequences, it cleaves the DNA. What purpose would restriction enzymes have in a bacterium like Bacillus?
(A) They are enzymes that no longer have a purpose because evolution has produced better enzymes.
(B) They destroy extra DNA that results from errors in binary fission.
(C) They protect Bacillus from invading DNA due to viruses.
(D) They prevent, or restrict, DNA replication when the cell isn’t ready to copy its DNA.
9. Viruses and bacteria have which of the following in common?
(A) Ribosomes
(B) Nucleic acids
(C) Flagella
(D) Metabolism
10. Griffith was a researcher who coined the term transformation when he noticed that incubating nonpathogenic bacteria with heat-killed pathogenic bacteria produced bacteria that ultimately became pathogenic, or deadly, in mice. What caused the transformation in his experiment?
(A) DNA from the nonpathogenic bacteria revitalized the pathogenic bacteria.
(B) Protein from the pathogenic bacteria was taken up by the nonpathogenic bacteria.
(C) DNA from the pathogenic bacteria was taken up by the nonpathogenic bacteria.
(D) DNA in the nonpathogenic bacteria turned into pathogenic genes in the absence of pathogenic bacteria.
11. Meselson and Stahl performed an elegant experiment using bacteria grown in heavy nitrogen (15N) to show that DNA replication is semi-conservative. After growing the bacteria in heavy nitrogen, they switched the cultures to 14N for two rounds of replication. They then examined the cells by density centrifugation, comparing the double helices by density. What is a likely result?
(A) Half of the DNA was labeled with 15N and half was labeled with 14N.
(B) One strand of each new DNA molecule was labeled with 15N, and the other strand was labeled with 14N.
(C) One band averaging the two molecular masses of nitrogen is seen.
(D) Two bands are seen. One equal to the average of the two masses and one representing a lighter mass.
12. A biologist systematically removes each of the proteins involved in DNA replication to determine the effect each has on the process. In one experiment, she denatures the DNA to see the results, separating the strands. She sees many short DNA/RNA fragments as well as some long DNA pieces. Which of the following is most likely missing?
(A) Helicase
(B) DNA polymerase
(C) DNA ligase
(D) RNA primase
REFLECT
Respond to the following questions:
• Which topics from this chapter do you feel you have mastered?
• Which content topics from this chapter do you feel you need to study more before you can answer multiple-choice questions correctly?
• Which content topics from this chapter do you feel you need to study more before you can effectively compose a free response?
• Was there any content that you need to ask your teacher or another person about?
Chapter 8
Cell Reproduction
CELL DIVISION
Every second, thousands of cells are dying throughout our bodies. Fortunately, the body replaces them at an amazing rate. In fact, epidermal cells, or skin cells, die off and are replaced so quickly that the average 18-year-old grows an entirely new skin every few weeks. The body keeps up this unbelievable rate thanks to the mechanisms of cell division.
This chapter takes a closer look at how cells divide. But remember, cell division is only a small part of the life cycle of a cell. Most of the time, cells are busy carrying out their regular activities. There are also some types of cells that are nondividing. Often, these are highly specialized cells that are created from a population of less specialized cells. The body continually makes them as needed, but they do not directly replicate themselves. Red blood cells are nondividing cells. Other cells are just temporarily nondividing. They enter a phase called G0, where they hang out until they get a signal to re-enter the normal cell cycle.
THE CELL CYCLE
Every cell has a life cycle—the period from the beginning of one division to the beginning of the next. The cell’s life cycle is known as the cell cycle. The cell cycle is divided into two periods: interphase and mitosis. Take a look at the cell cycle of a typical cell.
INTERPHASE: THE GROWING PHASE
Interphase is the time span from one cell division to another. We call this stage interphase (inter- means between) because the cell has not yet started to divide. Although biologists sometimes refer to interphase as the “resting stage,” the cell is definitely not inactive. This phase is when the cell carries out its regular activities. All the proteins and enzymes it needs to grow are produced during interphase.
The Three Stages of Interphase
Interphase can be divided into three stages: G1, S, G2.
The most important phase is the S phase. That’s when the cell replicates its genetic material. The first thing a cell has to do before undergoing mitosis is to duplicate all of its chromosomes, which contain the organism’s DNA “blueprint.” During interphase, every single chromosome in the nucleus is duplicated.
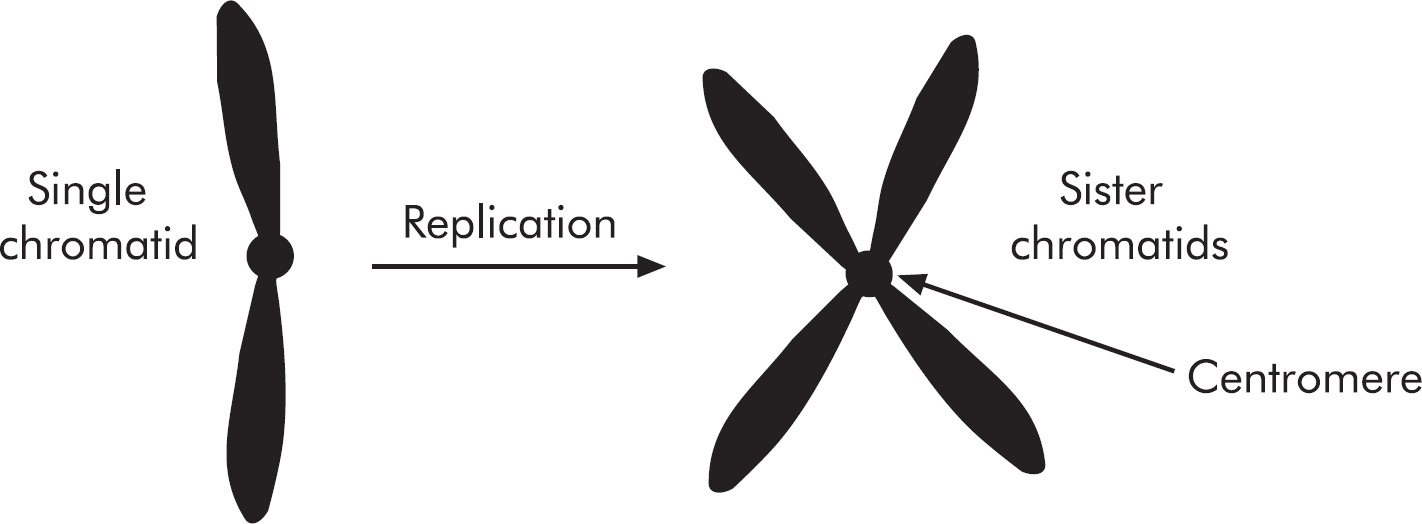
You’ll notice that the original chromosome and its duplicate are still linked, like conjoined twins. These identical strands of DNA are now called sister chromatids. The chromatids are held together by a structure called the centromere. You can think of each chromatid as a chromosome, but because they remain attached they are called chromatids instead. Chromosomes need to have their own centromere. Once the chromatids separate they will be full-fledged chromosomes.
Cell Cycle Regulation
We’ve already said that replication occurs during the S phase of interphase, so what happens during G1 and G2? During these stages, the cell produces proteins and enzymes. For example, during G1 the cell produces all of the enzymes required for DNA replication (DNA helicase, DNA polymerase, and DNA ligase, as we learned in Chapter 7). By the way, G stands for “gap,” but we can also associate it with “growth.” These three phases are highly regulated by checkpoints and special proteins called cyclins and cyclin-dependent kinases (CDKs).
Cell cycle checkpoints are control mechanisms that make sure cell division is happening properly in eukaryotic cells. They monitor the cell to make sure it is ready to progress through the cell cycle and stop progression if it is not. In eukaryotes, checkpoint pathways function mainly at phase boundaries (such as the G1/S transition and the G2/M transition). Checkpoint pathways are an example of an important cell signaling pathway inside of cells.
For example, if the DNA genome has been damaged in some way, the cell should not divide. If it does, damaged DNA will be passed onto daughter cells, which can be disastrous. When damaged DNA is found, checkpoints are activated and cell cycle progression stops. The cell uses the extra time to repair damage in DNA. If the DNA damage is so extensive that it cannot be repaired, the cell can undergo apoptosis, or programmed cell death. Apoptosis cannot stop once it has begun, so it is a highly regulated process. It is an important part of normal cell turnover in multicellular organisms.
Cell cycle checkpoints control cell cycle progression by regulating two families of proteins: cyclin-dependent kinases (CDKs) and cyclins. To induce cell cycle progression, an inactive CDK binds a regulatory cyclin. Once together, the complex is activated, can affect many proteins in the cell, and causes the cell cycle to continue. To inhibit cell cycle progression, CDKs and cyclins are kept separate. CDKs and cyclins were first studied in yeast, unicellular eukaryotic fungi. Budding yeast and fission yeast were good model organisms because they have only one CDK protein each and a few different cyclins. This relatively simple system allowed biologists to figure out how cell cycle progression is controlled. Their findings were then used to figure out how more complex eukaryotes (such as mammals) control the cell cycle.
Cancer
A cell losing control of the cell cycle can have disastrous consequences. For example, a mutation in a protein that normally controls progression through the cell cycle can result in unregulated cell division and cancer. Cancer means “crab,” as in the zodiac sign. The name comes from the observation that malignant tumors grow into surrounding tissue, embedding themselves like clawed crabs. Cancer occurs when normal cells start behaving and growing very abnormally and spread to other parts of the body.
Mutated genes that induce cancer are called oncogenes (“onco-” is a prefix denoting cancer). Normally, these genes are required for proper growth of the cell and regulation of the cell cycle. Oncogenes, then, are genes that can convert normal cells into cancerous cells. Often these are abnormal or mutated versions of normal genes. The normal healthy version is called a proto-oncogene.
Tumor suppressor genes produce proteins that prevent the conversion of normal cells into cancer cells. They can detect damage to the cell and work with CDK/cyclin complexes to stop cell growth until the damage can be repaired. They can also trigger apoptosis if the damage is too severe to be repaired.
Cell cycle checkpoints and apoptosis have been intimately linked with cancer. In order for a normal cell to become a cancer cell, it must override cell cycle checkpoints and grow in an unregulated way. It must also avoid cell death. Tumors often accumulate DNA damage. Cancer treatments target these changes, because they are what make a cancer cell different from a normal cell.
Let’s recap:
• The cell cycle consists of two things: interphase and mitosis.
• During the S phase of interphase, the chromosomes replicate.
• Growth and preparation for mitosis occur during the G1 and G2 stages of interphase.
• Cell cycle checkpoints make sure the cell is ready to continue through the cell cycle.
• CDK and cyclin proteins work together to promote cell cycle progression.
• Oncogenes promote cell growth and tumor suppressor genes inhibit cell growth.
MITOSIS: THE DANCE OF THE CHROMOSOMES
Once the chromosomes have replicated, the cell is ready to begin mitosis. Mitosis is the period when the cell divides. Mitosis consists of a sequence of four stages: prophase, metaphase, anaphase, and telophase. It is not important to memorize the name of each phase, but it is important that you know the basic order of operations for mitosis.
Stage 1: Prophase: Prep (the cell prepares to divide)
One of the first signs of prophase is the disappearance of the nucleolus (ribosome making area) and the nuclear envelope. In prophase, the chromosomes thicken, forming coils upon coils, and become visible, the dense chromosomes are called chromatin. (During interphase, the chromosomes are not visible.)
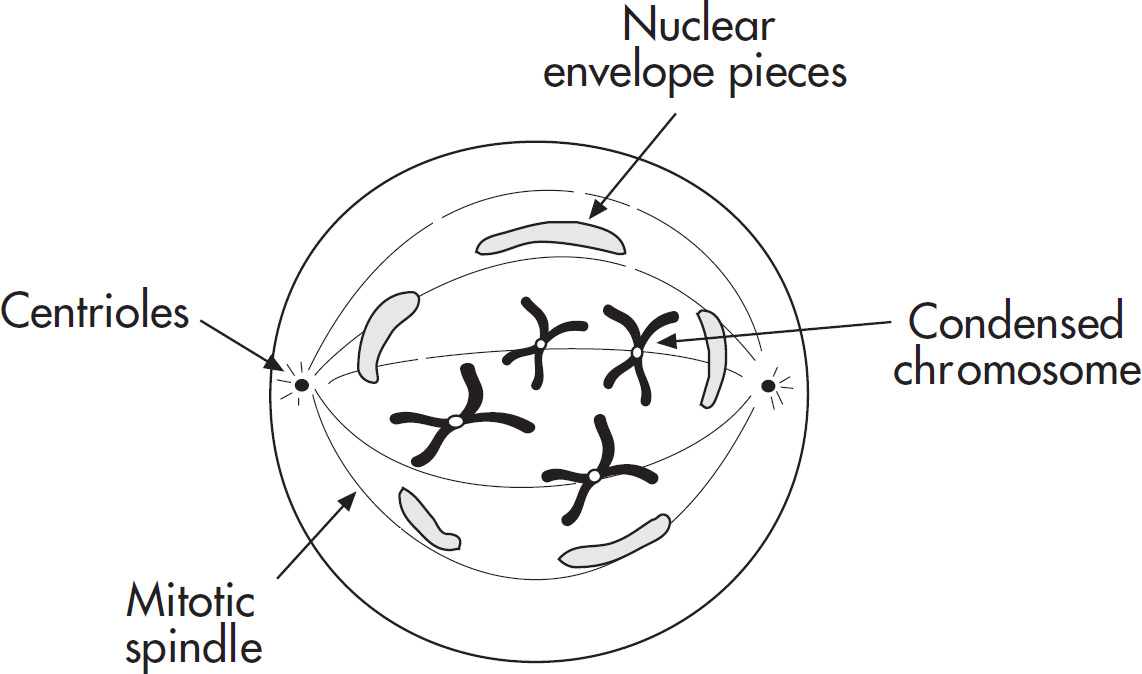
Now the cell has plenty of room to “sort out” the chromosomes. Remember centrioles? During prophase, these cylindrical bodies found within microtubule organizing centers (MTOCs) start to move away from each other, toward opposite ends of the cell. The centrioles will spin out a system of microtubules known as the spindle fibers. These spindle fibers will attach to a structure on each chromatid called a kinetochore. The kinetochores are part of the centromere.
Stage 2: Metaphase: Middle (chromosomes align in the middle)
The next stage is called metaphase. The chromosomes now begin to line up along the equatorial plane, or the metaphase plate, of the cell. That’s because the spindle fibers are attached to the kinetochore of each chromatid.
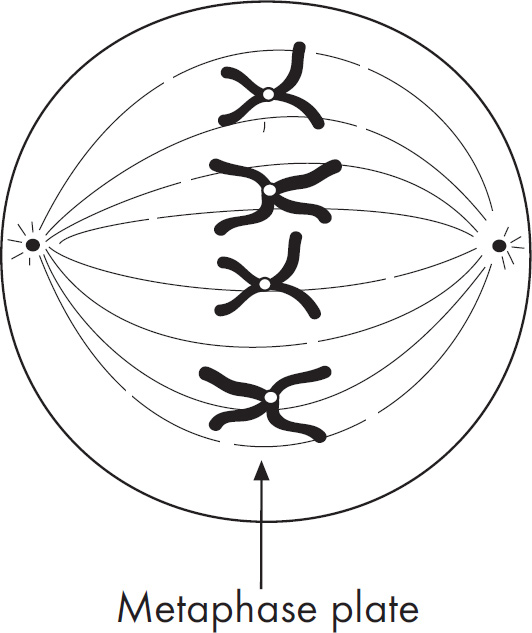
Stage 3: Anaphase: Apart (the chromatids are pulled apart)
During anaphase, the sister chromatids of each chromosome separate at the centromere and migrate to opposite poles. The chromatids are pulled apart by the microtubules, which begin to shorten. Each half of a pair of sister chromatids now moves to opposite poles of the cell. Non-kinetochore microtubules elongate the cell.
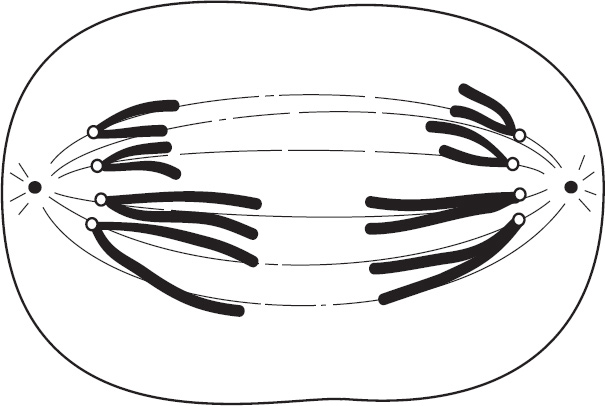
Stage 4: Telophase: Two (the cell completes splitting in two)
The final phase of mitosis is telophase. A nuclear membrane forms around each set of chromosomes and the nucleoli reappear.
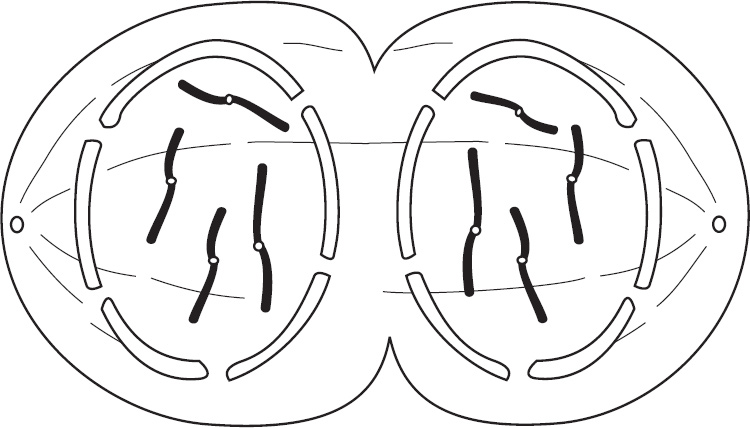
The nuclear membrane is ready to divide. Now it’s time to split the cytoplasm in a process known as cytokinesis. Look at the figure below and you’ll notice that the cell has begun to split along a cleavage furrow (which is produced by actin microfilaments).
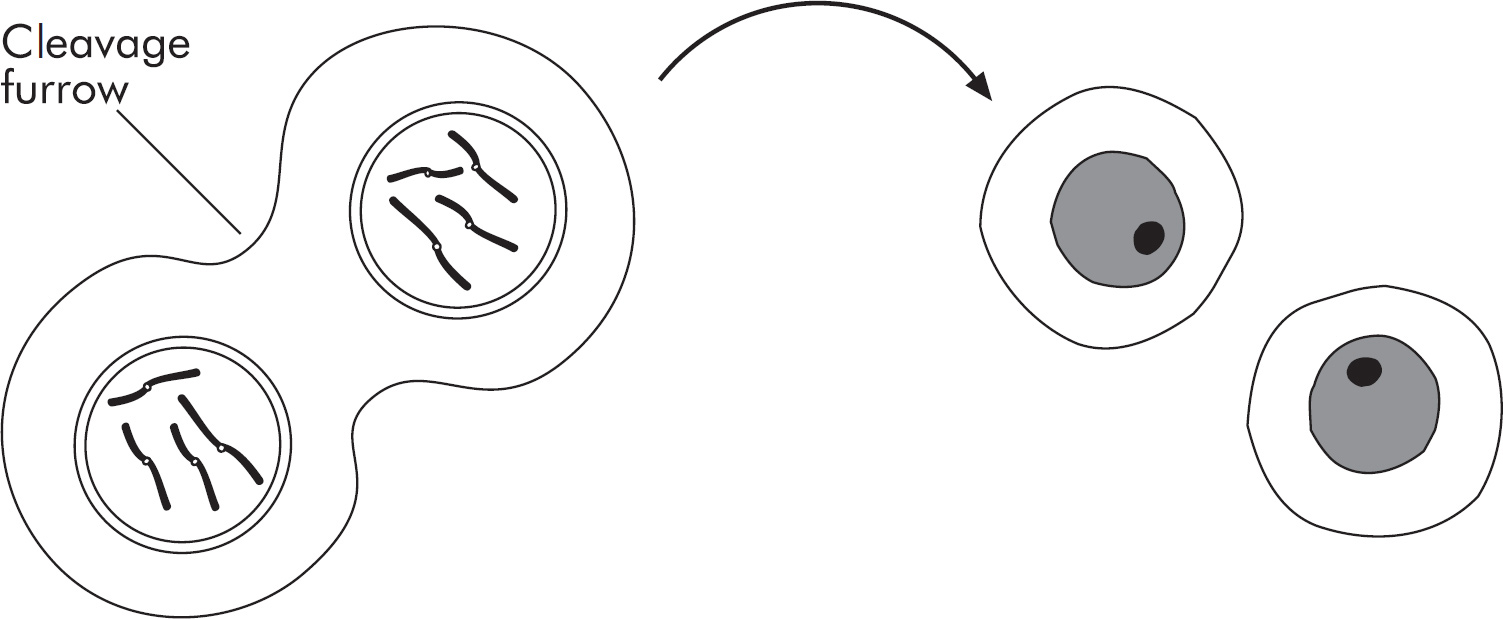
A cell membrane forms about each cell, and the cells split into two distinct daughter cells. The division of the cytoplasm yields two daughter cells.
Here’s one thing to remember: cytokinesis occurs differently in plant cells. The cell doesn’t form a cleavage furrow. Instead, a partition called a cell plate forms down the middle region.
Stage 5: Interphase
Once the daughter cells are produced, they reenter the initial phase—interphase—and the whole process starts over. The cell goes back to its original state. Once again, the chromosomes become invisible, and the genetic material is called chromatin again.
But How Will I Remember All That?
For mitosis, you may already have your own mnemonic. If not, here’s a table with a mnemonic we created for you.
| IPMAT | |
| Interphase | I is for Interlude |
| Prophase | P is for Prepare |
| Metaphase | M is for Middle |
| Anaphase | A is for Apart |
| Telophase | T is for Two |
Purpose of Mitosis
Mitosis has two purposes:
• to produce daughter cells that are identical copies of the parent cell
• to maintain the proper number of chromosomes from generation to generation
For our purposes, we can say that mitosis occurs in just about every cell except sex cells. When you think of mitosis, remember: “Like begets like.” Hair cells “beget” other hair cells, skin cells “beget” other skin cells, and so on. Mitosis is involved in growth, repair, and asexual reproduction.
HAPLOIDS VERSUS DIPLOIDS
Every organism has a certain number of unique chromosomes. For example, fruit flies have four chromosomes, humans have twenty-three chromosomes, and dogs have thirty-nine chromosomes. It turns out that most eukaryotic cells in fact have two full sets of chromosomes—one set from each parent. Humans, for example, have two sets of twenty-three chromosomes, giving us our grand total of forty-six.
A cell that has two sets of chromosomes is a diploid cell, and the zygotic chromosome number is given as “2n.” That means we have two copies of each chromosome. In humans, for example, the diploid number of chromosomes is forty-six.
If a cell has only one set of chromosomes, we call it a haploid cell. This kind of cell is given the symbol n. For example, we would say that the haploid number of chromosomes for humans is twenty-three.
Remember:
• Diploid refers to any cell that has two sets of chromosomes.
• Haploid refers to any cell that has one set of chromosomes.
Why do we need to know the terms haploid and diploid? Because they are extremely important when it comes to sexual reproduction. As we’ve seen, forty-six is the normal diploid number for human beings, but there are only twenty-three different chromosomes. The duplicate versions of each chromosome are called homologous chromosomes. The homologous chromosomes that make up each pair are similar in size and shape and express similar traits. This is the case in all sexually reproducing organisms. In fact, this is the essence of sexual reproduction: each parent donates half its chromosomes to its offspring.
Gametes
Although most cells in the human body are diploid, there are special cells that are haploid. These haploid cells are called sex cells, or gametes. Why do we have haploid cells?
As we’ve said, an offspring has one set of chromosomes from each of its parents. A parent, therefore, contributes a gamete with one set that will be paired with the set from the other parent to produce a new diploid cell, or zygote. This produces offspring that are a combination of their parents.
AN OVERVIEW OF MEIOSIS
Meiosis is the production of gametes. Since sexually reproducing organisms need only haploid cells for reproduction, meiosis is limited to sex cells in special sex organs called gonads. In males, the gonads are the testes, while in females they are the ovaries. The special cells in these organs—also known as germ cells—produce haploid cells (n), which then combine to restore the diploid (2n) number during fertilization.
female gamete (n) + male gamete (n) = zygote (2n)
When it comes to genetic variation, meiosis is a big plus. Variation, in fact, is the driving force of evolution. The more variation there is in a population, the more likely it is that some members of the population will survive extreme changes in the environment. Meiosis is far more likely to produce these sorts of variations than is mitosis, which therefore confers selective advantage on sexually reproducing organisms. We’ll come back to this theme in Chapter 10.
A Closer Look at Meiosis
Meiosis actually involves two rounds of cell division: meiosis I and meiosis II.
Before meiosis begins, the diploid cell goes through interphase. Just as in mitosis, double-stranded chromosomes are formed during this phase.
Meiosis I
Meiosis I consists of four stages: prophase I, metaphase I, anaphase I, and telophase I.
Prophase I
Prophase I is a little more complicated than regular prophase. As in mitosis, the nuclear membrane disappears, the chromosomes become visible, and the centrioles move to opposite poles of the nucleus. But that’s where the similarity ends.
The major difference involves the movement of the chromosomes. In meiosis, the chromosomes line up side-by-side with their counterparts (homologs). This event is known as synapsis.
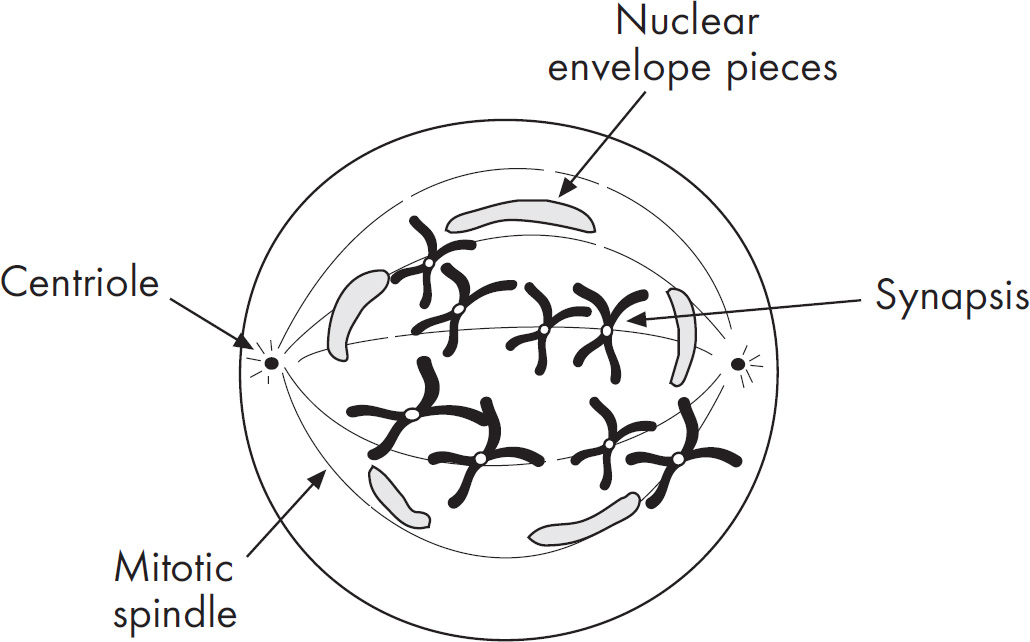
Synapsis involves two sets of chromosomes that come together to form a tetrad (or a bivalent). A tetrad consists of four chromatids. Synapsis is followed by crossing-over, the exchange of segments between homologous chromosomes.
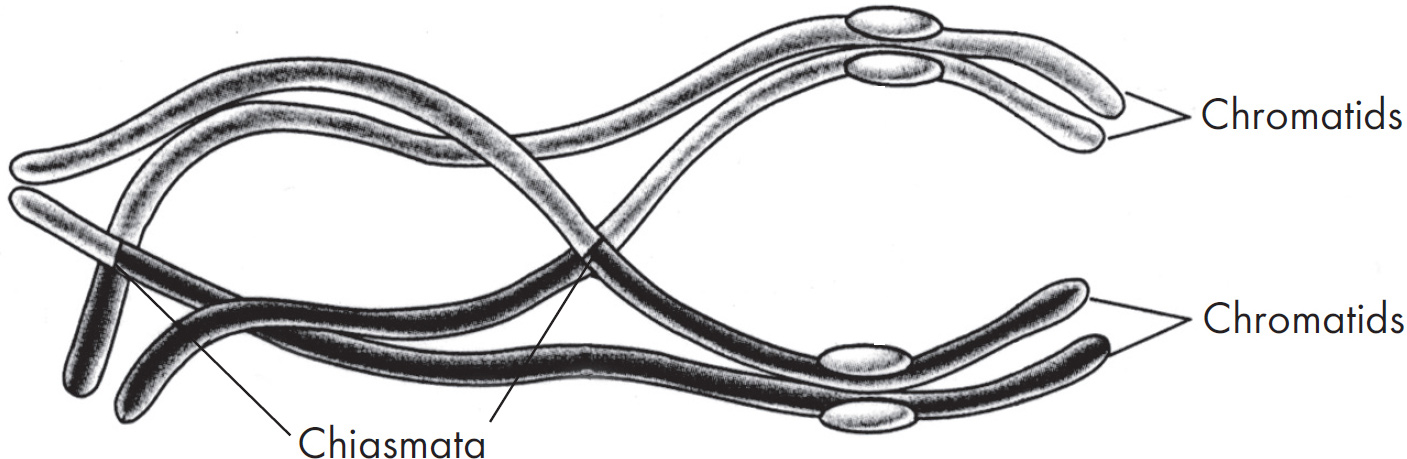
What’s unique in prophase I is that pieces of chromosomes are exchanged between the homologous partners. This is one of the ways organisms produce genetic variation. By the end of prophase I, the chromosomes will have exchanged regions containing several alleles, or different forms of the same gene. This means that each chromosome might have a different mix of alleles than it began with. Remember, the two homologous chromosomes are the versions that came from the person’s mother and father. So, a chromosome that has experienced crossing-over will now be a mix of alleles from the mother and father versions.
Metaphase I
As in mitosis, the chromosome pairs—now called tetrads—line up at the metaphase plate. By contrast, you’ll recall that in regular metaphase, the chromosomes line up individually. One important concept to note is that the alignment during metaphase is random, so the copy of each chromosome that ends up in a daughter cell is random. Therefore, the gamete will be created with a mixture of genetic information from the person’s father and mother. This means that the offspring created would be a combination of all four grandparents.
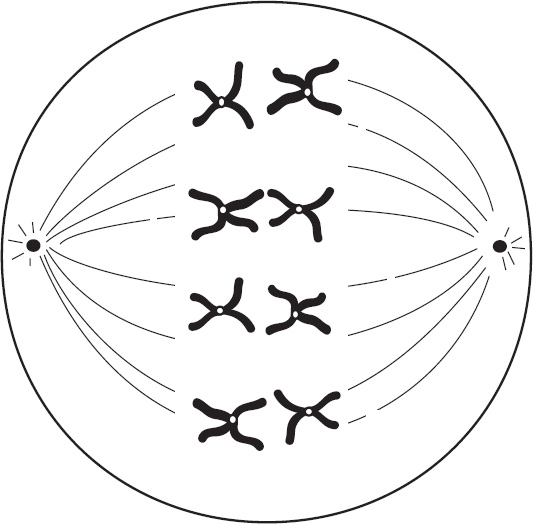
Anaphase I
During anaphase I, one of each pair of chromosomes within a tetrad separates and moves to opposite poles. Notice that the chromatids do not separate at the centromere. The homologs separate with their centromeres intact.
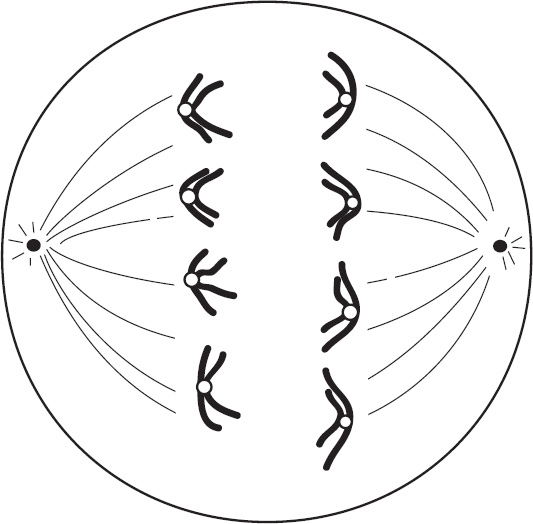
The chromosomes now go on to their respective poles.
Telophase I
During telophase I, the nuclear membrane forms around each set of chromosomes.
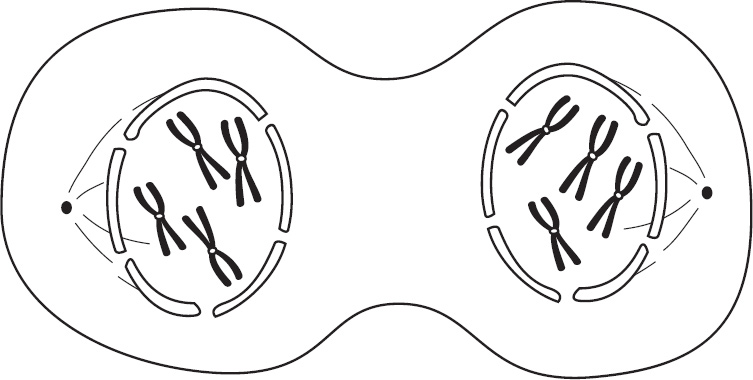
Finally, the cells undergo cytokinesis, leaving us with two daughter cells. Notice that at this point the nucleus contains the haploid number of chromosomes, but each chromosome is a duplicated chromosome.
Meiosis II
The purpose of the second meiotic division is to separate the sister chromatids, and the process is virtually identical to mitosis. Let’s run through the steps in meiosis II.
During prophase II, the chromosomes once again condense and become visible. In metaphase II, the chromosomes move toward the metaphase plate. This time they line up single file, not as pairs. During anaphase II, the chromatids of each chromosome split at the centromere, and each chromatid is pulled to opposite ends of the cell. At telophase II, a nuclear membrane forms around each set of chromosomes and a total of four haploid cells are produced.
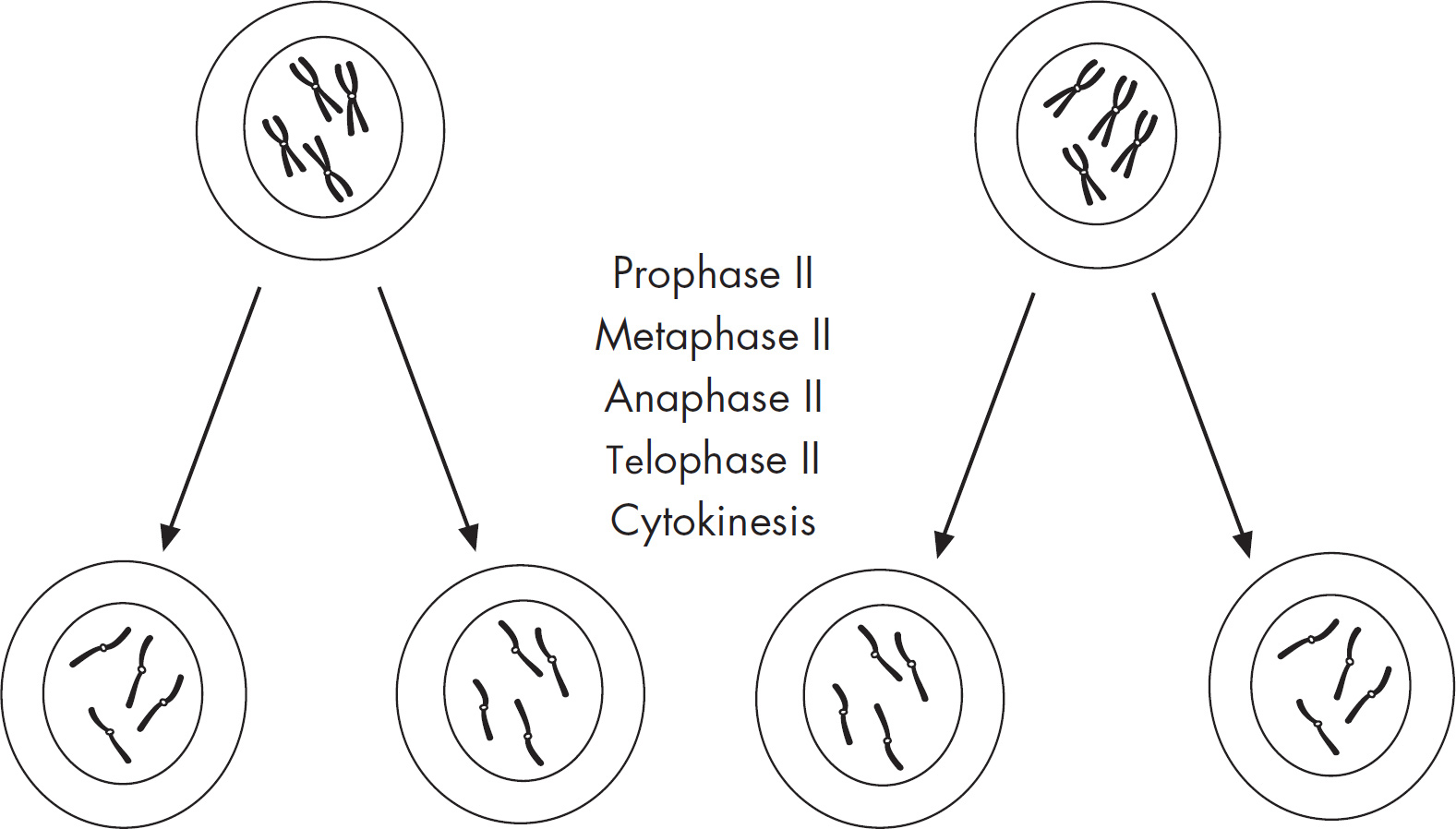
Gametogenesis
Meiosis is also known as gametogenesis. If sperm cells are produced, then meiosis is called spermatogenesis. During spermatogenesis, four sperm cells are produced for each diploid cell. If an egg cell or an ovum is produced, this process is called oogenesis.
Oogenesis is a little different from spermatogenesis. Oogenesis produces only one ovum, not four. The other three cells, called polar bodies, get only a tiny amount of cytoplasm and eventually degenerate. Why does oogenesis produce only one ovum? Because the female wants to conserve as much cytoplasm as possible for the surviving gamete, the ovum.
Here’s a summary of the major differences between mitosis and meiosis.
| MITOSIS | MEIOSIS |
| • Occurs in somatic (body) cells | • Occurs in germ (sex) cells |
| • Produces identical cells | • Produces gametes |
| • Diploid cell → diploid cells | • Diploid cell → haploid cells |
| • 1 cell becomes 2 cells | • 1 cell becomes 4 cells |
| • Number of divisions: 1 | • Number of divisions: 2 |
Meiotic Errors
Sometimes, a set of chromosomes has an extra or a missing chromosome. This occurs because of nondisjunction—the chromosomes failed to separate properly during meiosis. This error, which produces the wrong number of chromosomes in a cell, usually results in miscarriage or severe genetic defects. For example, humans typically have two copies of each chromosome, but individuals with Down syndrome have three—instead of two—copies of the twenty-first chromosome.
Chromosomal abnormalities also occur if one or more segments of a chromosome break and are either lost or reattach to another chromosome. The most common example is translocation (a segment of a chromosome moves to another chromosome).
Here’s an example of a translocation.
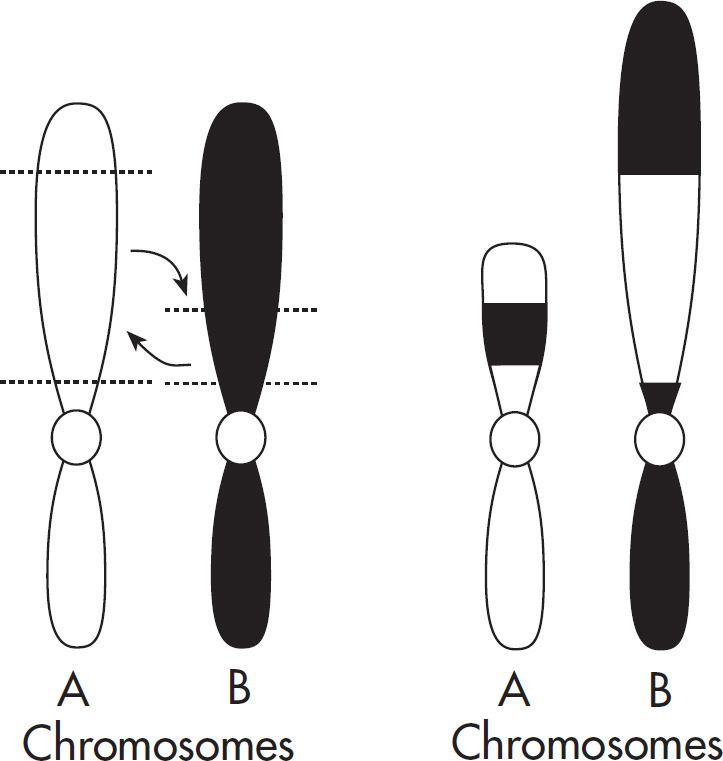
Fortunately, in most cases, damaged DNA can usually be repaired with special repair enzymes.
KEY TERMS
cell division
cell cycle
interphase
mitosis
G0 phase
G1 phase
G2 phase
S phase
sister chromatids
centromere
cyclins
cyclin-dependent kinases (CDKs)
cell cycle checkpoints
apoptosis
cancer
oncogene
tumor suppressor gene
prophase
metaphase
anaphase
telophase
chromatin
spindle fibers
kinetochores
metaphase plate
cytokinesis
cleavage furrow
cell plate
diploid cell
haploid cell
homologous chromosomes
sex cells (or gametes)
meiosis
gonads
testes
ovaries
germ cells
meiosis I
meiosis II
synapsis
tetrad (or bivalent)
crossing-over (or recombination)
alleles
gametogenesis
spermatogenesis
oogenesis
polar bodies
ovum
nondisjunction
Down syndrome
translocation
Summary
 The cell cycle is divided into interphase and mitosis, or cellular division.
The cell cycle is divided into interphase and mitosis, or cellular division.
 The three stages of interphase are G1, G2, and S phase.
The three stages of interphase are G1, G2, and S phase.
• S phase is the “synthesis” phase, when chromosomes replicate.
• Growth and preparation for mitosis occur in G1 and G2.
 Cell cycle progression is controlled by checkpoint pathways and CDK/cyclin complexes.
Cell cycle progression is controlled by checkpoint pathways and CDK/cyclin complexes.
 Cancer occurs when cells grow abnormally and spread to other parts of the body. Tumor-suppressor genes are genes that prevent the cell from dividing when it shouldn’t. Proto-oncogenes are genes that help the cell divide. If either type is mutated (a mutated proto-oncogene is called an oncogene) it can lead to cell growth that is out of control.
Cancer occurs when cells grow abnormally and spread to other parts of the body. Tumor-suppressor genes are genes that prevent the cell from dividing when it shouldn’t. Proto-oncogenes are genes that help the cell divide. If either type is mutated (a mutated proto-oncogene is called an oncogene) it can lead to cell growth that is out of control.
 Mitosis is cellular division and occurs in four stages: prophase, metaphase, anaphase, and telophase.
Mitosis is cellular division and occurs in four stages: prophase, metaphase, anaphase, and telophase.
• Prophase is when the nuclear envelope disappears and chromosomes condense.
• Next is metaphase, when chromosomes align at the metaphase plate and mitotic spindles attach to kinetochores.
• Anaphase pulls the chromosomes away from the center.
• Telophase terminates mitosis, and the two new nuclei form.
• The process of cytokinesis, which occurs during telophse, ends mitosis, as the cytoplasm and plasma membranes pinch to form two distinct, identical daughter cells.
 Meiosis produces four genetically distinct haploid gametes (sperm or egg cells). Meiosis involves two rounds of cell division: meiosis I and meiosis II.
Meiosis produces four genetically distinct haploid gametes (sperm or egg cells). Meiosis involves two rounds of cell division: meiosis I and meiosis II.
• Meiosis I is when the homologous chromosome pairs separate, but before they can they undergo crossing-over and swap some DNA.
• Meiosis II is when the sister chromatids separate.
• Mutations, such as nondisjunction events or whole chromosome translocations, can occur as a result of crossing-over.
Chapter 8 Drill
Answers and explanations can be found in Chapter 16.
1. A scientist is testing new chemicals designed to stop the cell cycle at various stages of mitosis. Upon applying one of the chemicals, she notices that all of the cells appear as shown below. Which of the following best explains how the chemical is likely acting on the cells?
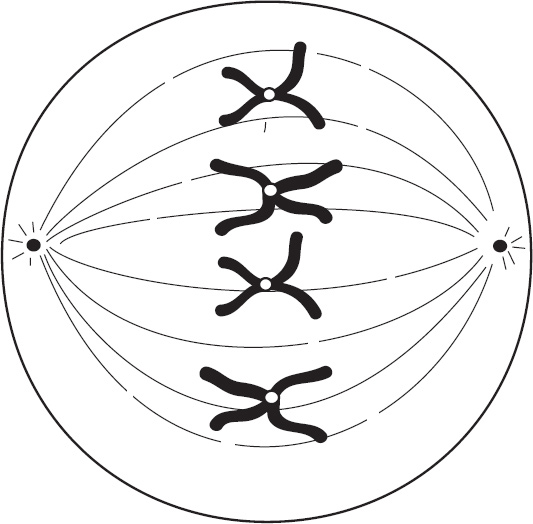
(A) The chemical has arrested the cells in prophase and has prevented attachment of the spindle fibers to the kinetochore.
(B) The chemical has arrested the cells in metaphase and has prevented dissociation of the spindle fibers from the centromere.
(C) The chemical has arrested the cells in metaphase and is preventing the shortening of the spindle fibers.
(D) The chemical has arrested the cells in anaphase and is preventing the formation of a cleavage furrow.
Questions 2 and 3 refer to the following graph and paragraph.
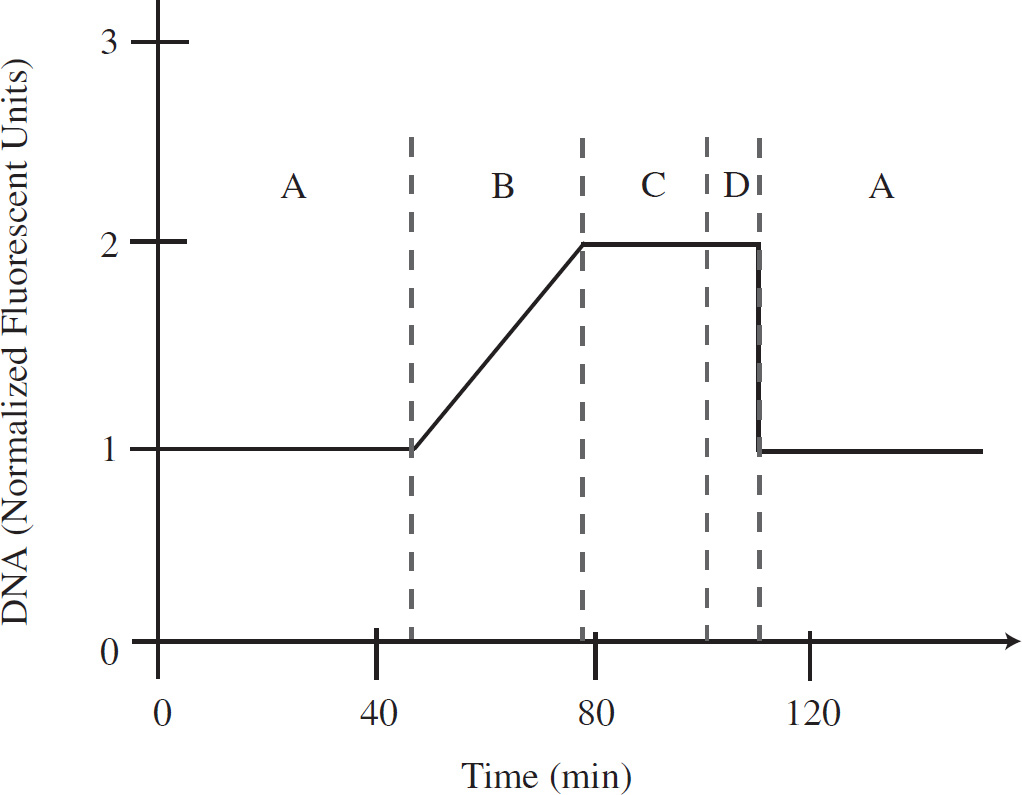
An experiment is performed to evaluate the amount of DNA present during a complete cell cycle. All of the cells were synced prior to the start of the experiment. During the experiment, a fluorescent chemical was applied to cells, which would only fluoresce when bound to DNA. The results of the experiment are shown above. Differences in cell appearance by microscopy or changes in detected DNA were determined to be phases of the cell cycle and are labeled with the letters A–D.
2. Approximately how long does S phase take to occur in these cells?
(A) 15 min
(B) 20 min
(C) 30 min
(D) 40 min
3. During which of the labeled phases of the experiment would the cell undergo anaphase?
(A) Phase A
(B) Phase B
(C) Phase C
(D) Phase D
Question 4 represents a question requiring a numeric answer. Calculate the correct answer for each question.
4. A new mammal has recently been discovered in the Amazonian jungle. A karyotype was performed on gametic cells and revealed that the animal had 13 completely unique chromosomes. How many homologous pairs would you expect to find in a diploid cell of the organism following completion of S phase? Give your answer to the nearest whole number.

5. Trisomy 21, which results in Down syndrome, results from nondisjunction of chromosome 21 in humans. Nondisjunction occurs when two homologous chromosomes, or two sister chromatids, do not separate. Which of the following describes the mechanism of this defect?
(A) During DNA replication in S phase of the cell cycle, the two new strands do not separate.
(B) During mitosis, at the metaphase plate, non-sister chromatids do not separate.
(C) The mitotic spindle attaches to chiasmata rather than kinetochores.
(D) The same microtubule in the spindle attaches to both sister chromatids during meiosis II.
6. A researcher isolates DNA from different types of cells and determines the amount of DNA for each type of cell. The samples may contain cells in various stages of the cell cycle. In order of increasing DNA content, which of the following would have the least amount of DNA to the greatest amount of DNA?
(A) Sperm, neuron, liver
(B) Ovum, muscle, taste bud
(C) Liver, heart, intestine
(D) Skin, neuron, ovum
7. Which of the following mutations would be least likely to have any discernable phenotype on the individual?
(A) Translocation of the last 1,000 base pairs of chromosome 1 onto chromosome 2
(B) Nondisjunction of chromosome 7 to produce trisomy-7
(C) Nondisjunction of chromosome 8 to produce monosomy-8
(D) Deletion of 400 base pairs, resulting in the loss of an enhancer
REFLECT
Respond to the following questions:
• Which topics from this chapter do you feel you have mastered?
• Which content topics from this chapter do you feel you need to study more before you can answer multiple-choice questions correctly?
• Which content topics from this chapter do you feel you need to study more before you can effectively compose a free response?
• Was there any content that you need to ask your teacher or another person about?
Chapter 9
Heredity
GREGOR MENDEL: THE FATHER OF GENETICS
What is genetics? In its simplest form, genetics is the study of heredity. It explains how certain characteristics are passed on from parents to children. Much of what we know about genetics was discovered by the monk Gregor Mendel in the 19th century. Since then, the field of genetics has vastly expanded. But before we get ahead of ourselves, let’s study the basic rules of genetics.
Let’s begin then with some of the fundamental points of genetics:
• Traits—or expressed characteristics—are influenced by one or more of your genes. Remember, a gene is simply a chunk of DNA that codes for a particular recipe. Different recipes affect different traits. DNA is passed from generation to generation, and from this process the genes and the traits associated with those genes are inherited. Within a chromosome, there are many genes, each controlling the inheritance of a particular trait. For example, in pea plants, there’s a gene on the chromosome that codes for seed coat. The position of a gene on a chromosome is called a locus.
• Diploid organisms (organisms that have two sets of chromosomes) usually have two copies of a gene, one on each homologous chromosome. Homologous chromosomes are the two copies/versions of a chromosome that diploids have. Humans have 23 pairs of homologous chromosomes.
These copies may be different from one another—that is, they may be alleles, or alternate versions/flavors of the same gene. For example, if we’re talking about the height of a pea plant, there’s an allele for tall and an allele for short. In other words, both alleles are alternate forms of the gene for height.
• When an organism has two identical alleles for a given trait, the organism is homozygous. If an organism has two different alleles for a given trait, the organism is heterozygous.
• When discussing the physical appearance of an organism, we refer to its phenotype. The phenotype tells us what the organism looks like. When talking about the genetic makeup of an organism, we refer to its genotype. The genotype tells us which alleles the organism possesses.
• An allele can be dominant or recessive. This is determined by which allele “wins out” over the other in a heterozygote. The convention is to assign one of two letters for the two different alleles. The dominant allele receives a capital letter and the recessive allele receives a lowercase of the same letter. For instance, we might give the dominant allele for height in pea plants a T for tall. This means that the recessive allele would be t.
| Name | Genotype | Phenotype |
| homozygous dominant | TT | Tall |
| homozygous recessive | tt | Short |
| heterozygous | Tt | Tall |
One of the major ways the testing board likes to test genetic information is by having you do crosses. Crosses involve the mating of hypothetical organisms with specific phenotypes and genotypes. We’ll look at some examples in a moment, but for now, keep these test-taking tips in mind:
• Label each generation in the cross. The first generation in an experiment is always called the parent, or P1 generation. The offspring of the P1 generation are called the filial, or F1 generation. The next generation, the grandchildren, is called the F2 generation.
• Always write down the symbols you’re using for the alleles, along with a key to remind yourself what the symbols refer to. Use uppercase for dominant alleles and lowercase for recessive alleles.
Now let’s look at some basic genetic principles.
MENDELIAN GENETICS
One of Mendel’s hobbies was to study the effects of cross-breeding on different strains of pea plants. Mendel worked exclusively with true-breeder pea plants. This means the plants he used were genetically pure and consistently produced the same traits. For example, tall plants always produced tall plants; short plants always produced short plants. Through his work, he came up with three principles of genetics: the law of dominance, the law of segregation, and the law of independent assortment.
The Law of Dominance
Mendel crossed two true-breeding plants with contrasting traits: tall pea plants and short pea plants.
To his surprise, when Mendel mated these plants, the characteristics didn’t blend to produce plants of average height. Instead, all the offspring were tall.
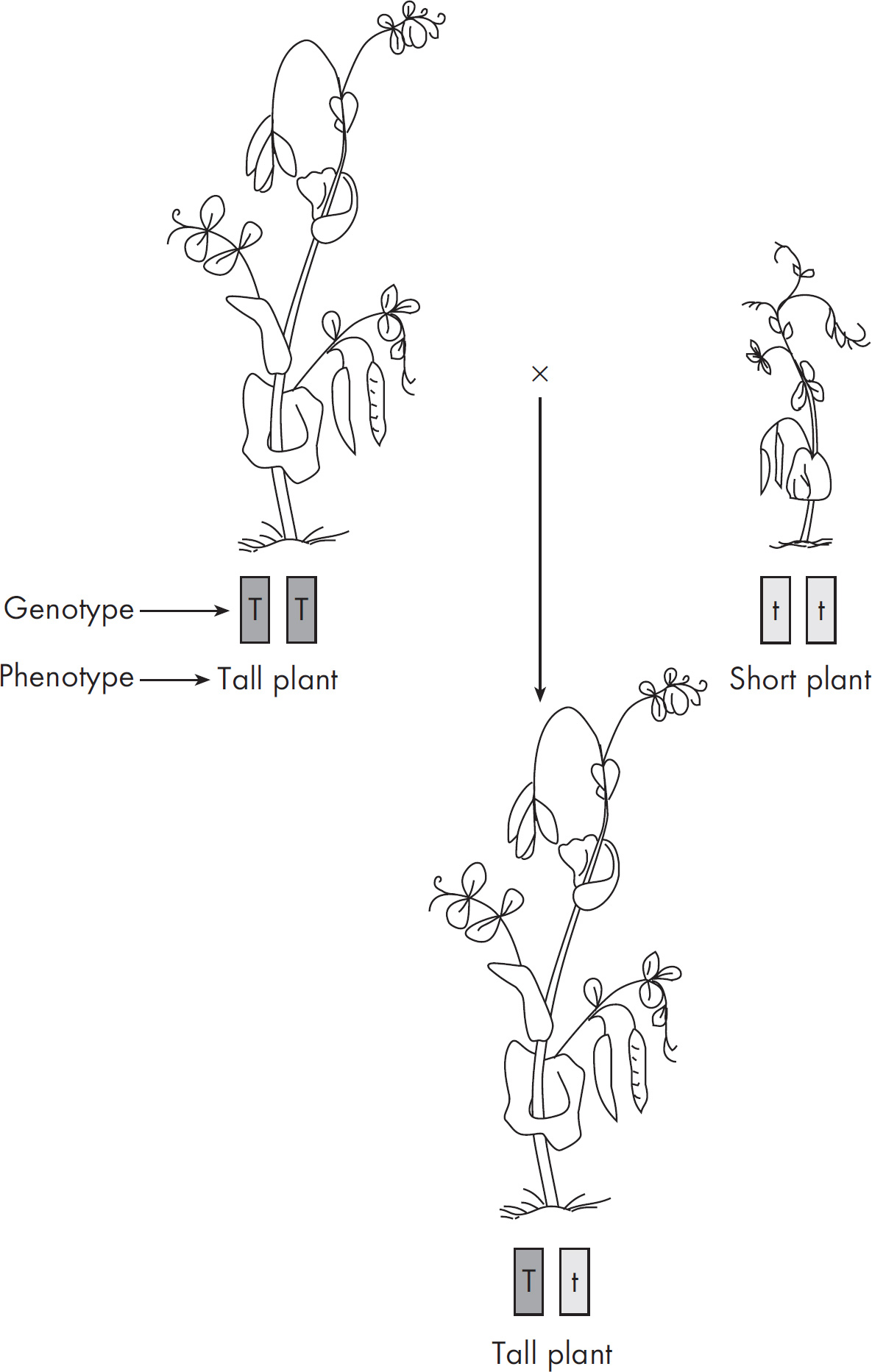
Mendel recognized that one trait must be masking the effect of the other trait. This is called the law of dominance. The dominant tall allele, T, somehow masked the presence of the recessive short allele, t. Consequently, all a plant needs is one tall allele to make it tall.
Monohybrid Cross
A monohybrid cross is when two individuals that are heterozygous for two traits are crossed. However, in order to be sure that you have heterozygotes, the first step is to cross true-breeding plants of allele #1 with true-breeding plants of allele #2. Follow along as these plants are crossed.
A simple way to represent a cross is to set up a Punnett square. Punnett squares are used to predict the results of a cross. Let’s construct a Punnett square for the cross between Mendel’s tall and short pea plants. Let’s first designate the alleles for each plant. As we saw earlier, we can use the letter “T” for the tall dominant allele and “t” for the recessive short allele.
Since one parent was a pure, tall pea plant, we’ll give it two dominant alleles (TT homozygous dominant). The other parent was a pure, short pea plant, so we’ll give it two recessive alleles (tt homozygous recessive). Let’s put the alleles for one of the parents across the top of the box and the alleles for the other parent along the side of the box.
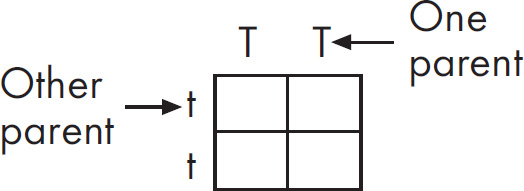
Now we can fill in the four boxes by matching the letters. What are the results for the F1 generation?
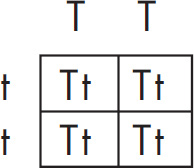
Each offspring received one allele from each parent. They all received one T and one t. They’re all Tt! Our parents had duplicate copies of single alleles, TT and tt, respectively. We could therefore refer to them as homozygous. The offspring, on the other hand, are heterozygous: they possess one copy of each allele.
Let’s compare the results of this cross with what we already know about meiosis. From meiosis, we know that when gametes are formed, the chromosomes separate so that each cell gets one copy of each chromosome. We now know that chromosomes are made up of genes, and genes consist of alleles. We’ve just seen that alleles also separate and recombine. We can say, therefore, that each allele on the edge of a Punnett square also “represents” a gamete.
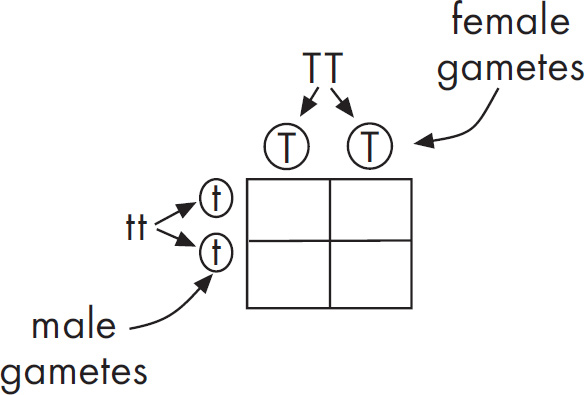
When fertilization occurs, chromosomes—along with the alleles they carry—get paired up in a new combination.
The Law of Segregation
Next, Mendel took the offspring and self-pollinated them. Let’s use a Punnett square to spell out the results. This time we’re mating the offspring of the first generation—F1. Take a look at the results:
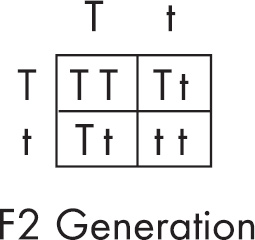
One of the offspring could be a short pea plant! The short-stemmed trait reappeared in the F2 generation. How could that happen? Once again, the alleles separated and recombined, producing a new combination for this offspring. The cross resulted in one offspring with a pair of recessive alleles, tt. Because there is no T (dominant) allele around to mask the expression of the recessive short allele, our new plant could wind up short.
Although all of the F1 plants appear to be tall, the alleles separate and recombine during the cross. This is an example of the Law of Segregation. It just means that each gamete only gets one of the two copies of a gene. Gametes are haploid.
What about the genotype and phenotype for this cross? Remember, genotype refers to the genetic makeup of an organism, whereas phenotype refers to the appearance of the organism. Using the results of our Punnett square, what is the ratio of phenotypes and genotypes in the offspring?
Let’s sum up the results. We have four offspring with two different phenotypes: three of the offspring are tall, whereas one of them is short. On the other hand, we have three genotypes: 1 TT, 2 Tt, and 1 tt.
Here’s a summary of the results:
• The ratio of phenotypes is 3:1 (three tall:one short).
• The ratio of genotypes is 1:2:1 (one TT:two Tt:one tt).
The Law of Independent Assortment
So far, we have looked at only one trait: tall versus short. What happens when we study two traits at the same time? The two traits also segregate randomly. This is an example of independent assortment. For example, let’s look at two traits in pea plants: height and color. When it comes to height, a pea plant can be either tall or short. As for color, the plant can be either green or yellow, with green being dominant. Each gene comes in two alleles, so this gives us four alleles total. By the Law of Independent Assortment, these four alleles can combine to give us four different gametes.
TG Tg tG tg
In other words, each allele of the height gene can be paired with either allele of the color gene. The reason alleles segregate independently is because chromosomes segregate independently. During meiosis I, each pair of homologous chromosomes is split, and which one aligns left or right during metaphase I is different for each pair.
Dihybrid Cross
Now that we know that different genes assort independently, let’s look at a cross of two traits. The dihybrid cross is just like the monohybrid, but it is when two heterozygotes for two genes are crossed. Keep in mind that the uppercase letter refers to the dominant allele. Therefore, T refers to tall and G refers to green, whereas t refers to short and g refers to yellow. Now let’s set up a cross between plants differing in two characteristics—called a dihybrid cross—using these four gametes and see what happens.
Each trait will act independently, meaning that a plant that is tall can be either green or yellow. Similarly, a green plant can be either tall or short.
Here is the Punnett Square for a cross between two double heterozygotes (Tt Gg).

This is an example of the Law of Independent Assortment. Each of the traits segregated independently. Don’t worry about the different combinations in the cross—you’ll make yourself dizzy with all those letters. Simply memorize the phenotype ratio of the pea plants. For the 16 offspring there are:
• 9 tall and green
• 3 tall and yellow
• 3 short and green
• 1 short and yellow
That’s 9:3:3:1, but don’t forget the circumstances that this applies. It is only when two heterozygotes for two genes are crossed. It is not just a magic ratio that always works.
The Punnett square method works well for monohybrid crosses and helps us visualize the possible combinations. However, a better method for predicting the likelihood of certain results from a dihybrid cross is to apply the law of probability. For dihybrid ratios, the law states that the probability that two or more independent events will occur simultaneously is equal to the product of the probability that each will occur independently. To illustrate the product rule, let’s consider again the cross between two identical dihybrid tall, green plants with the genotype TtGg. To find the probability of having a tall, yellow plant, simply multiply the probabilities of each event. If the probability of being tall is  and the probability of being yellow is
and the probability of being yellow is  , then the probability of being tall and yellow is
, then the probability of being tall and yellow is  ×
×  =
= 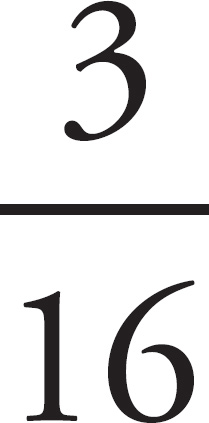 .
.
Let’s summarize Mendel’s three laws.
| SUMMARY OF MENDEL’S LAWS | |
| Laws | Definition |
| Law of dominance | One trait masks the effects of another trait. |
| Law of segregation | Each gamete only gets one of the copies of each gene. |
| Law of independent assortment | Each pair of homologous chromosomes splits independently, so the alleles of different genes can mix and match |
Test Cross
Suppose we want to know if a tall plant is homozygous (TT) or heterozygous (Tt). Its physical appearance doesn’t necessarily tell us about its genetic makeup. The only way to determine its genotype is to cross the plant with a recessive, short plant, tt. This is known as a test cross, or back cross. Using the recessive plant, there are only two possibilities: (1) TT × tt or (2) Tt × tt. Let’s take a look.
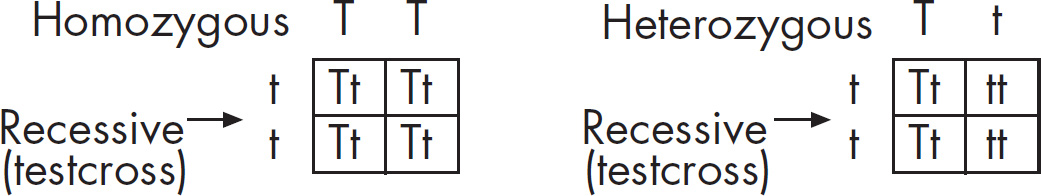
If none of the offspring is short, our original plant must have been homozygous, TT. If, however, even one short plant appears in the bunch, we know that our original pea plant was heterozygous, Tt. In other words, it wasn’t a pure-breeding plant. A test cross uses a recessive organism to determine the genotype of an organism of unknown genotype.
BEYOND MENDELIAN GENETICS
Not all patterns of inheritance obey the principles of Mendelian genetics. In fact, many traits we observe are due to a combined expression of alleles. Here are a couple of examples of non-Mendelian forms of inheritance:
• Incomplete dominance (blending inheritance): In some cases, the traits will blend. For example, if you cross a white snapdragon plant (genotype WW) with a red snapdragon plant (RR), the resulting progeny will be pink (RW). In other words, neither color is dominant over the other.
• Codominance: Sometimes you’ll see an equal expression of both alleles. For example, an individual can have an AB blood type. In this case, each allele is equally expressed. That is, both the A allele and the B allele are expressed (IAIB). That’s why the person is said to have AB blood. The expression of one allele does not prevent the expression of the other.
• Polygenic inheritance: In some cases, a trait results from the interaction of many genes. Each gene will have a small effect on a particular trait. Height, skin color, and weight are all examples of polygenic traits.
• Multiple alleles: Some traits are the product of many different alleles that occupy a specific gene locus. The best example is the ABO blood group system in which three alleles (IA, IB, and i) determine blood type.
• Non-nuclear inheritance: Apart from the genetic material held in the nucleus, there is also genetic material that is contained in the mitochondria. The mitochondria are always provided by the egg during sexual reproduction, so mitochondrial inheritance is always through the maternal line, not the male line.
• Linked genes: Sometimes genes on the same chromosome stay together during assortment and move as a group. The group of genes is considered linked and tends to be inherited together. For example, the genes for flower color and pollen shape are linked on the same chromosomes and show up together.
Since linked genes are found on the same chromosome, they cannot segregate independently. This violates the Law of Independent Assortment.
Let’s pretend that the height and color genes we looked at in the dihybrid cross are linked. A heterozygote for both traits still has two alleles for height (T and t) and two alleles for color (G and g). However, because height and color are located on the same chromosome, the allele for height and the allele for color are physically linked. For example, maybe the heterozygote has one chromsome with Tg and one chromosome with tG. When gametes are formed, the T and g will travel together, and the t and G will travel together and be packaged into a gamete together. So, in the unlinked dihybrid shown earlier there were four possible gamete combinations (TG, Tg, tG, tg), but now there are only two (Tg and tG). The only way to physically separate linked alleles is by crossing over. If a crossover event occurs between the linked genes then recombinant gametes can occur.
If the genes were unlinked then the the four gametes (TG, Tg, tG, tg) would be equally likely. If certain combinations of alleles are found more often in offspring (are likely appearing more often in gametes), then this is a sign of possible linkage.
Look at the example below from a cross of TtGg and ttgg. If you draw out the punnett square you should get equal numbers of each phenotype occurring if the genes are assumed to be unlinked.
Green and Tall: 6 offspring
Green and short: 37 offspring
yellow and Tall: 45 offspring
yellow and short: 3 offspring
There are more Green/short and yellow/Tall offspring. This is a sign that these alleles are linked on a chromosome.
The percentage of recombination (or recombination frequency) can be determined by adding up the recombinants and dividing by the total number of offspring.
9/91 = 9.9%
This percentage can also be used as a measure of how far apart the genes are. The distance on a chromosome is measured in map units or centimorgans.
The frequency of crossing-over between any two linked alleles is proportional to the distance between them. That is, the farther apart two linked alleles are on a chromosome, the more often the chromosome will cross over between them. This finding led to recombination mapping—mapping of linkage groups with each map unit being equal to 1 percent recombination. For example, if two linked genes, A and B, recombine with a frequency of 15 percent, and B and C recombine with a frequency of 9 percent, and A and C recombine with a frequency of 24 percent, what is the sequence and the distance between them?
The sequence and the distance of A-B-C is

If the recombination frequency between A and C had been 6 percent instead of 24 percent, the sequence and distance of A-B-C would instead be
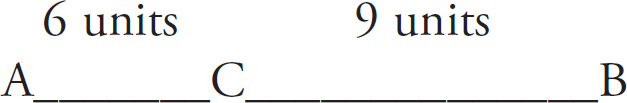
SEX-LINKED TRAITS
We already know that humans contain twenty-three pairs of chromosomes. Twenty-two of the pairs of chromosomes are called autosomes. They code for many different traits. The other pair contains the sex chromosomes. This pair determines the sex of an individual. A female has two X chromosomes. A male has one X and one Y chromosome—an X from his mother and a Y from his father. Some traits, such as color blindness and hemophilia, are carried on sex chromosomes. These are called sex-linked traits. Most sex-linked traits are found on the X chromosome and are more properly referred to as “X-linked.”
Since males have one X and one Y chromosome, what happens if a male has a defective X chromosome? Unfortunately, he’ll express the sex-linked trait, even if it is recessive. Why? Because his one and only X chromosome is defective. He doesn’t have another X to mask the effect of the bad X. However, if a female has only one defective X chromosome, she won’t express a recessive sex-linked trait. For her to express the trait, she has to inherit two defective X chromosomes. A female with one defective X is called a carrier. Although she does not exhibit the trait, she can still pass it on to her children.
You can also use the Punnett square to figure out the results of sex-linked traits. Here’s a classic example: a male who has normal color vision and a woman who is a carrier for color blindness have children. How many of the children will be color-blind? To figure out the answer, let’s set up a Punnett square.
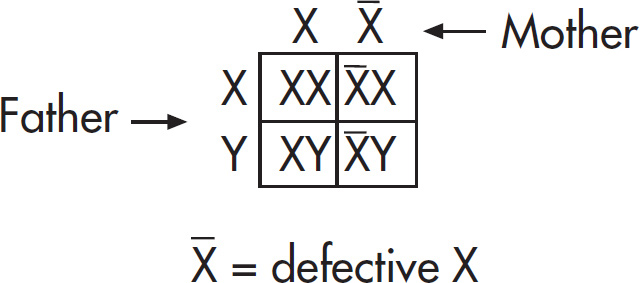
Notice that we placed a bar above any defective X to indicate the presence of a defective allele. And now for the results. The couple have one son who is color-blind, a son who sees color normally, a daughter who is a carrier, and a daughter who sees color normally. The color-blind child is a son.
Barr Bodies
A look at the cell nucleus of normal females will reveal a dark-staining body known as a Barr body. A Barr body is an X chromosome that is condensed and visible. In every female cell, one X chromosome is activated and the other X chromosome is deactivated during embryonic development. Surprisingly, the X chromosome destined to be inactivated is randomly chosen in each cell. Therefore, in every tissue in the adult female one X chromosome remains condensed and inactive. However, this X chromosome is replicated and passed on to a daughter cell. X-inactivation is why it is okay that females have two X chromosomes and males only have one. After X-inactivation, it is like everyone has one copy.
Other Sex-Linked Diseases
• Duchenne muscular dystrophy
• Vitamin D resistant rickets
• Hereditary nephritis
• Juvenile gout
KEY TERMS
Gregor Mendel
trait
genes
locus
homologous chromosomes
alleles
homozygous
heterozygous
phenotype
genotype
dominant
recessive
parent, or P1 generation
filial, or F1 generation
F2 generation
law of dominance
law of segregation
law of independent assortment
monohybrid cross
Punnett square
dihybrid cross
test cross
incomplete dominance
codominance
polygenetic inheritance
multiple alleles
non-nuclear inheritance
linked genes
map units
autosomes
sex chromosomes
color blindness
hemophilia
sex-linked traits
carrier
Barr body
Summary
 Genetics is the study of heredity. The foundation of genetics is that traits are produced by genes, which are found on chromosomes.
Genetics is the study of heredity. The foundation of genetics is that traits are produced by genes, which are found on chromosomes.
 Diploid organisms carry two copies of every gene, one on each of two homologous chromosomes, one that came from their mother and one that came from their father.
Diploid organisms carry two copies of every gene, one on each of two homologous chromosomes, one that came from their mother and one that came from their father.
 When the two alleles are the same, the organism is homozygous for that trait. When the two alleles are different, the organism is heterozygous for the trait.
When the two alleles are the same, the organism is homozygous for that trait. When the two alleles are different, the organism is heterozygous for the trait.
 The genes are described as the genotype of the organism. The physical expression of the traits, or what you see in the organism, is the phenotype.
The genes are described as the genotype of the organism. The physical expression of the traits, or what you see in the organism, is the phenotype.
 Homozygous individuals only have one allele that will determine the phenotype. However, if the individual is heterozygous they have two different alleles that could contribute to the phenotype. The allele that “wins out” and directs the phenotype is the dominant allele. The one that doesn’t show up is the recessive allele.
Homozygous individuals only have one allele that will determine the phenotype. However, if the individual is heterozygous they have two different alleles that could contribute to the phenotype. The allele that “wins out” and directs the phenotype is the dominant allele. The one that doesn’t show up is the recessive allele.
 Simple genetics with dominants and recessives was illuminated by Gregor Mendel. In Mendelian genetics, there are three important laws: The law of dominance, the law of segregation, and the law of independent assortment.
Simple genetics with dominants and recessives was illuminated by Gregor Mendel. In Mendelian genetics, there are three important laws: The law of dominance, the law of segregation, and the law of independent assortment.
 Non-Mendelian genetics is when traits do not follow the Mendelian laws. Examples include the following:
Non-Mendelian genetics is when traits do not follow the Mendelian laws. Examples include the following:
• incomplete dominance
• codominance
• polygenic inheritance
• multiple alleles
• linked genes
 Humans have twenty-three pairs of homologous chromosomes for a total of forty-six. Of these, twenty-two pairs are autosomes and two are designated as sex chromosomes:
Humans have twenty-three pairs of homologous chromosomes for a total of forty-six. Of these, twenty-two pairs are autosomes and two are designated as sex chromosomes:
• Females have two X chromosomes; males have one X and one Y chromosome.
• Because females have two X chromosomes, one is inactivated in each cell and condenses into a Barr body.
Chapter 9 Drill
Answers and explanations can be found in Chapter 16.
1. Two expecting parents wish to determine all possible blood types of their unborn child. If both parents have an AB blood type, which of the following blood types will their child NOT possess?
(A) A
(B) B
(C) AB
(D) O
2. A new species of tulip was recently discovered. A population of pure red tulips was crossed with a population of pure blue tulips. The resulting F1 generation was all purple. This result is an example of which of the following?
(A) Complete dominance
(B) Incomplete dominance
(C) Codominance
(D) Linkage
3. In pea plants, tall (T) is dominant over short (t) and green (G) is dominant over yellow (g). If a pea plant heterozygous for both traits is crossed with a plant that is recessive for both traits, approximately what percentage of the progeny plants will be tall and yellow?
(A) 0%
(B) 25%
(C) 66%
(D) 75%
Questions 4-6 refer to the following pedigree tree and paragraph.
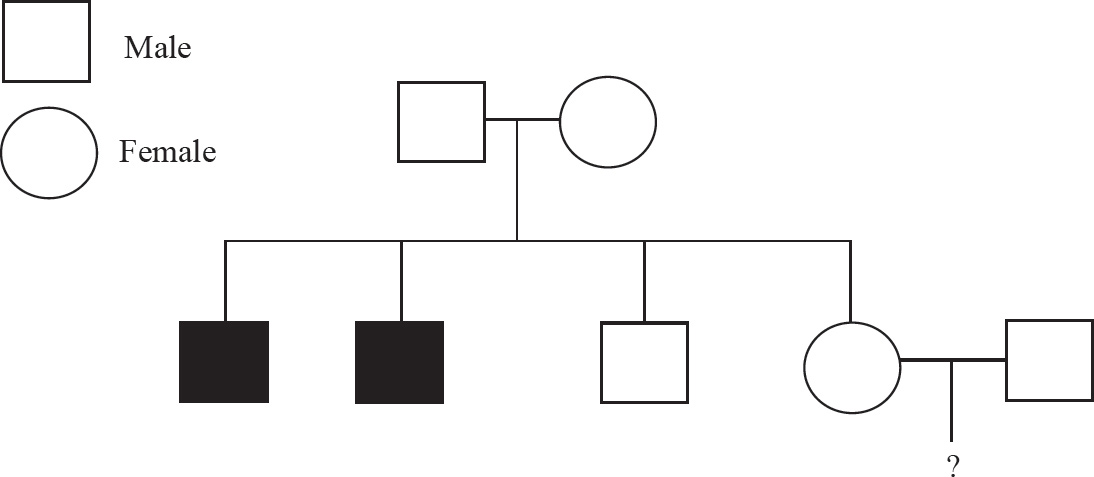
Hemophilia is an X-linked disease associated with the inability to produce specific proteins in the blood-clotting pathway. Shown above is a family pedigree tree in which family members afflicted with the disease are shown with filled-in squares (male) or circles (females). A couple is trying to determine the likelihood of passing on the disease to their future children (represented by the ? symbol above) because the hemophilia runs in the woman’s family.
4. Assuming that the woman in the couple is a carrier, what is the probability that the couple’s first son will have hemophilia?
(A) 0%
(B) 25%
(C) 50%
(D) 100%
5. Why were both women in the family tree above free of the disease?
(A) They were lucky because they didn’t receive an X chromosome with the diseased allele.
(B) They cannot get hemophilia because it is only associated with a diseased Y chromosome.
(C) They are carriers and will only get the disease if they have children.
(D) They have at least one normal (unaffected) copy of the X chromosome.
6. Turner syndrome is a disease in which an individual is born with only a single X chromosome. Suppose the woman in the couple is a carrier for hemophilia and has a child with Turner Syndrome, would this child have the disease?
(A) Yes, because the child would only have one copy of the X chromosome and it would be affected.
(B) No, because women cannot be affected by hemophilia.
(C) No, because the child would have to receive a normal copy of the X chromosome from its mother.
(D) Maybe, it depends on which X chromosome she receives from her mother.
7. Neuronal ceroid lipofuscinosis (NCL) is a group of autosomal recessive diseases characterized by blindness, loss of cognitive and motor function, and early death. One of the genes that are mutated in this disease is CLN3. When functional CLN3 protein is absent, neurons die because of increased storage material in the cells, presumably because the lysosomes aren’t working properly. What is a valid explanation as to why parents of children with this devastating disease can be completely normal?
(A) Parents of the autosomal recessive disease must both be carriers of the mutation on the CLN3 gene.
(B) Carriers of the CLN3 mutation genotype do not show the phenotype because one normal CLN3 allele is present to provide a functioning CLN3 protein.
(C) Redundant proteins take over the function of the mutant CLN3 protein in the parents.
(D) X inactivation prevents expression of the CLN3 mutated protein in the parents.
8. A breeder of black Labrador puppies notices that in his line a gene predisposing his dogs for white spots has arisen. He believes the white spot allele is autosomal recessive, and he wants to prevent it from continuing in his dogs. He sees that one of his female dogs, Speckle, has white spots, and he therefore assumes she is homozygous for the gene. But he must determine which males in his pack are heterozygous to avoid breeding them in the future. What is a reasonable plan?
(A) Stop breeding Speckle.
(B) Pair Speckle with a male with white spots.
(C) Perform a testcross of Speckle with a black Labrador male. If any pups are born with white spots, stop breeding that male because he is heterozygous.
(D) Breed Speckle with a chocolate Labrador.
9. Which of the following is a reason why certain traits do not follow Mendel’s Law of Independent Assortment?
(A) The genes are linked on the same chromosome.
(B) It only applies to eukaryotes.
(C) Certain traits are not completely dominant.
(D) Nondisjunction
10. The gene for coat color in some breeds of cats is found on the X chromosome. Calico cats are mottled with orange and black colorings. What is a possible explanation for the fact that true calico cats are only female?
(A) The allele for coat color is randomly chosen by X-inactivation.
(B) The coat color is linked with genes that are lethal to male cats, so male calico cats never live past birth.
(C) Coat orange and black color is expressed only in female cats; males have only one color.
(D) Male cats have Y chromosomes.
REFLECT
Respond to the following questions:
• Which topics from this chapter do you feel you have mastered?
• Which content topics from this chapter do you feel you need to study more before you can answer multiple-choice questions correctly?
• Which content topics from this chapter do you feel you need to study more before you can effectively compose a free response?
• Was there any content that you need to ask your teacher or another person about?
Chapter 10
Evolutionary Biology
All of the organisms we see today arose from earlier organisms. This process, known as evolution, can be described as a change in a population over time. Interestingly, however, the driving force of evolution, natural selection, operates on the level of the individual. In other words, evolution is defined in terms of populations but occurs in terms of individuals.
NATURAL SELECTION
What is the basis of our knowledge of evolution? Much of what we now know about evolution is based on the work of Charles Darwin. Darwin was a nineteenth-century British naturalist who sailed the world in a ship named the HMS Beagle. Darwin developed his theory of evolution based on natural selection after studying animals in the Galápagos Islands and other places.
He observed that there were similar animals on the various islands, but the beaks varied in length on the finches and the necks varied in length on the tortoises. There must be a reason why animals in different areas had different traits. Darwin concluded that it was impossible for the finches and tortoises of the Galápagos simply to “grow” longer beaks or necks as needed. Rather, those traits were inherited and passed on from generation to generation. So, he decided there must be a way for populations to evolve and change their traits (i.e., a population of finches developing longer beaks).
He decided that there must have been a variety of beak lengths originally, but only the longest one was particularly helpful. Since those finches could eat better, survive better, and reproduce better, they were more likely to contribute offspring to the next generation. Over many years, the long-beak finches became more and more plentiful with each generation until long beaks were the “norm.” In another example, on the first island Darwin studied, there must once have been short-necked tortoises. Unable to reach the higher vegetation, these tortoises eventually died off, leaving only those tortoises with longer necks. Consequently, evolution has come to be thought of as “the survival of the fittest”: Only those organisms most fit to survive will survive and reproduce.
Darwin elaborated his theory in a book entitled On the Origin of Species. In a nutshell, here’s what Darwin observed:
• Each species produces more offspring than can survive.
• These offspring compete with one another for the limited resources available to them.
• Organisms in every population vary.
• The fittest offspring, or those with the most favorable traits, are the most likely to survive and therefore produce a second generation.
Lamarck and the Long Necks
Darwin was not the first to propose a theory explaining the variety of life on Earth. One of the most widely accepted theories of evolution in Darwin’s day was that proposed by Jean-Baptiste de Lamarck.
In the eighteenth century, Lamarck had proposed that acquired traits were inherited and passed on to offspring. For example, in the case of our giraffes, Lamarck’s theory said that the giraffes had long necks because they were constantly reaching for higher leaves while feeding. This theory is referred to as the “law of use and disuse,” or, as we might say now, “use it or lose it.” According to Lamarck, giraffes have long necks because they constantly use them.
We know now that Lamarck’s theory was wrong: acquired changes—that is, changes at a “macro” level in somatic (body) cells—cannot be passed on to germ cells. For example, if you were to lose one of your fingers, your children would not inherit this trait.
Evidence for Evolution
In essence, nature “selects” which living things survive and reproduce. Today, we find support for the theory of evolution in several areas:
• Paleontology, or the study of fossils: Paleontology has revealed to us both the great variety of organisms (most of which, including trilobites, dinosaurs, and the woolly mammoth, have died off) and the major lines of evolution.
• Biogeography, or the study of the distribution of flora (plants) and fauna (animals) in the environment: scientists have found related species in widely separated regions of the world. For example, Darwin observed that animals in the Galápagos have traits similar to those of animals on the mainland of South America. One possible explanation for these similarities is a common ancestor. As we’ll see below, there are other explanations for similar traits. However, when organisms share multiple traits, it’s pretty safe to say that they also shared a common ancestor.
• Embryology, or the study of the development of an organism: if you look at the early stages in vertebrate development, all the embryos look alike! For example, all vertebrates—including fish, amphibians, birds, and even humans—show fishlike features called gill slits.
• Comparative anatomy, or the study of the anatomy of various animals: scientists have discovered that some animals have similar structures that serve different functions. For example, a human’s arm, a dog’s leg, a bird’s wing, and a whale’s fin are all the same appendages, though they have evolved to serve different purposes. These structures, called homologous structures, also point to a common ancestor.
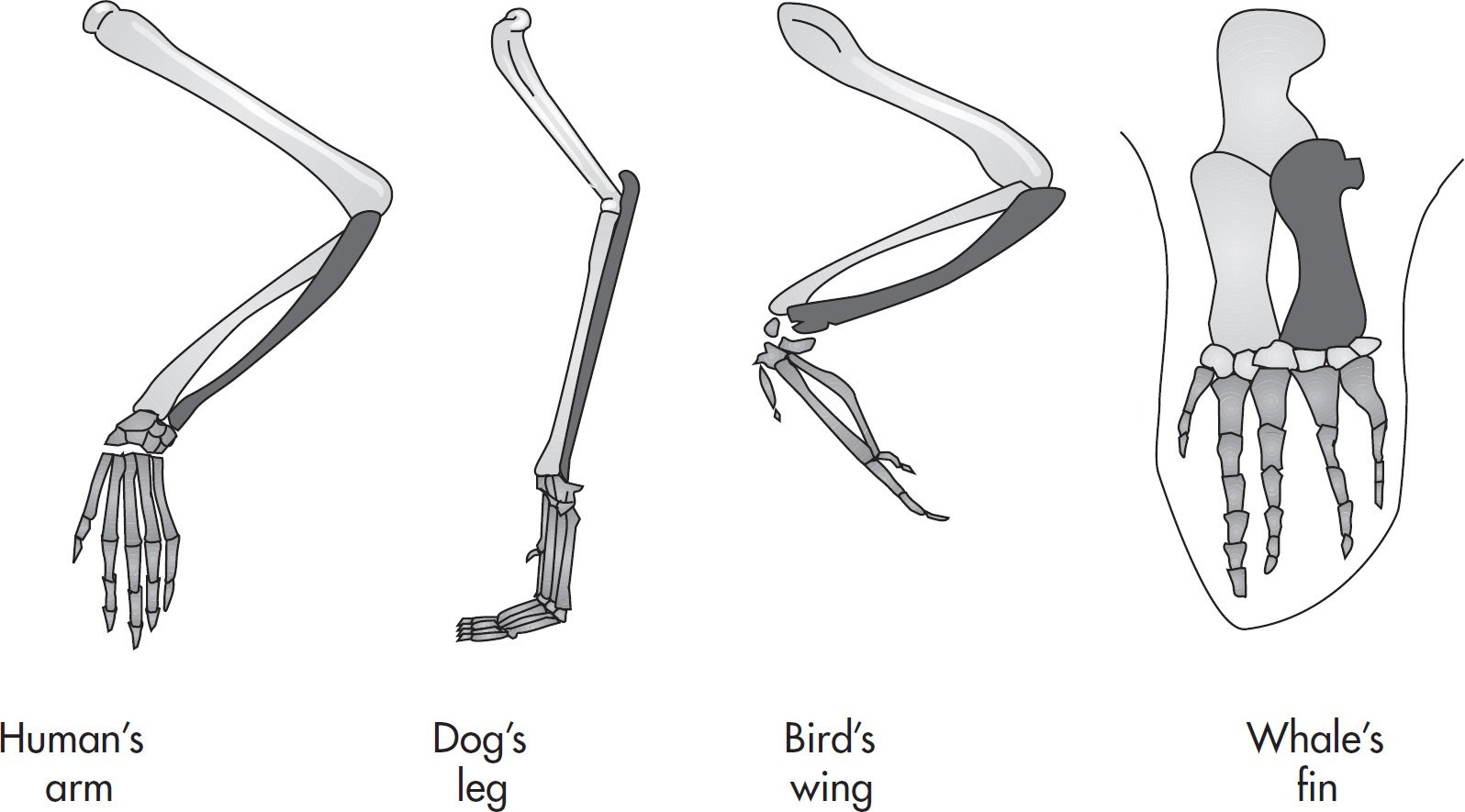
In contrast, sometimes animals have features with the same function but that are structurally different. A bat’s wing and an insect’s wing, for example, are both used to fly. They therefore have the same function, but they have evolved totally independently of one another. These are called analogous structures. Another classic example of an analogous structure is the eye. Though scallops, insects, and humans all have eyes, these three different types of eyes are thought to have evolved entirely independently of one another. They are therefore analogous structures.
• Molecular biology: Perhaps the most compelling proof of all is the similarity at the molecular level. Today, scientists can examine the nucleotide and amino acid sequences of different organisms. From these analyses, we’ve discovered that organisms that are closely related have a greater proportion of sequences in common than distantly related species. For example, most of us don’t look much like chimpanzees. However, by some estimates, as much as 99 percent of our genetic code is identical to that of a chimp.
COMMON ANCESTRY
By using the evidence mentioned above, scientists can get a good handle on how evolution of certain species occurred. It all comes down to who has common ancestors. Life started somewhere. Some original life-form is the common ancestor to all life, but where things went from there can become quite convoluted.
Scientists use charts called phylogenetic trees, or cladograms, to study the relationships between organisms. They always begin with the common ancestor and then branch out. Anytime there is a fork in the road, it is called a common ancestor node. These common ancestors likely do not exist anymore, but they are the point at which evolution went in two directions. One direction eventually led to one species, and the other eventually led to the other species.
For example, we have a common ancestor with chimps. This doesn’t mean that our great-great…great-grandfather was a chimp. It means that chimps and we have the same great-great…great-grandfather, who was neither a human nor a chimp. Instead, he was some other species completely, and a speciation event must have occurred where his species was split and evolved in different directions. One lineage became chimps, and the other linage became humans.
This is an example of a phylogenetic tree.
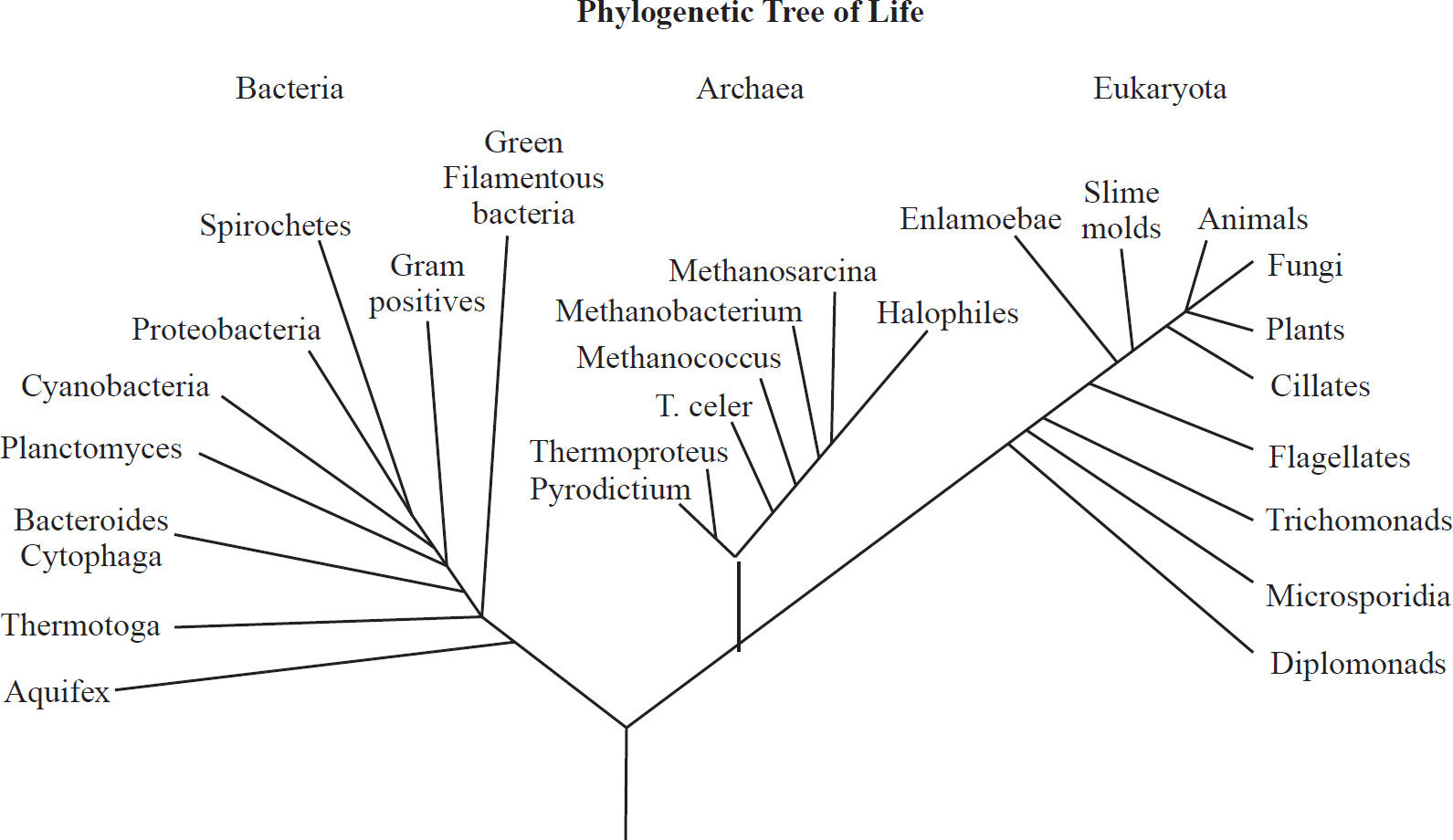
In this tree, the three main domains of life are seen. You can see that plants, animals, and fungi are much more closely related to each other than they are to bacteria. The archaea and the eukarya have a common ancestor with bacteria, and then they also have another common ancestor that bacteria doesn’t have.
GENETIC VARIABILITY
As you know, no two individuals are identical. The differences in each person are known as genetic variability. All this means is that no two individuals in a population have identical sets of alleles (except, of course, identical twins). In fact, the survival of a species is dependent on this genetic variation; it allows a species to survive in a changing environment. If a population were all the same then they would have the same weaknesses and the same strengths. Natural selection only occurs if some individuals have more evolutionary fitness and can be selected. The more variations there are among a population, the more likely that a trait will exist that might be the perfect lifesaver. The most famous example is Rudolph the Red-Nosed Reindeer. His red nose was a random mutation, but under the right conditions, it became the best nose to have. A more diverse population is more likely to survive and evolve when things are constantly changing around them.
How did all this wonderful variation come about? Well, the simple answer is that random mutations are occurring all the time. This can be because of errors by the DNA polymerase, changes to the DNA caused by transposons, or other types of DNA damage. Either way, mutations create new variations and alleles.
However, just the mutations occurring does not account for the great amount of genetic variation among a population. The mixing of genes through sexual reproduction contributes to genetic variation. During meiosis, crossing-over mixes the alleles among homologous chromosomes, and the independent assortment when chromosomes are packaged further adds to the genetic uniqueness of each gamete.
In bacteria, a process called conjugation increases the genetic variation, even though they are asexual reproducers. Additionally, viruses can pass around chunks of genome during infection. This process is called transduction. Each of these processes will be covered more in Chapter 11.
It might be hard to think of it in this way, but genetic variation is the very foundation of evolution, as we’ll soon see. Now that we’ve reintroduced genes, we can refine our definition of evolution. More specifically:
Evolution is the change in the gene pool of a population over time.
The Peppered Moths
Let’s take an example. During the 1850s in England, there was a large population of peppered moths. In most areas, exactly half of them were dark or carried alleles for dark coloring. The other half carried the alleles for light coloring. This 1:1 ratio of phenotypes was observed until air pollution, due primarily to the burning of coal, changed the environment. What happened?
Imagine two different cities: City 1 (in the south) and City 2 (in the north). Prior to the Industrial Revolution, both of these cities had unpolluted environments. In both of these environments, dark moths and light moths lived comfortably side by side. For simplicity’s sake, let’s say our proportions were a perfect fifty-fifty, half dark and half light. However, at the height of the Industrial Revolution, City 2, our northern city, was heavily polluted, whereas City 1, our southern city, was unchanged. In the north, where all the trees and buildings were covered with soot, the light moths didn’t stand a chance. They were impossible for a predator to miss! As a result, the predators gobbled up light-colored moths just as fast as they could reproduce, sometimes even before they reached an age in which they could reproduce. However, the dark moths were just fine. With all the soot around, the predators couldn’t even see them; the dark moths continued doing their thing—above all, reproducing. And when they reproduced, they had more and more offspring carrying the dark allele.
After a few generations, the peppered moth gene pool in City 2 changed. Although our original moth gene pool was 50 percent light and 50 percent dark, excessive predation changed the population’s genetic makeup. By about 1950, the gene pool reached 90 percent dark and only 10 percent light. This occurred because the light moth didn’t stand a chance in an environment where it was so easy to spot. The dark moths, on the other hand, multiplied just as fast as they could.
In the southern city, you’ll remember, there was very little pollution. What happened there? Things remained pretty much the same. The gene pool was unchanged, and the population continued to have roughly equal proportions of light moths and dark moths.
CAUSES OF EVOLUTION
Natural selection, the evolutionary mechanism that “selects” which members of a population are best suited to survive and which are not, works both “internally” and “externally”: internally through random mutations and externally through environmental pressures.
Natural selection requires genetic variation and an environmental pressure that gives some individuals an advantage.
To see how this process unfolds in nature, let’s return to the moth case. Why did the dark moths in the north survive? Because they were dark-colored. But how did they become dark-colored? The answer is, through random mutation. One day, a moth was born with dark-colored wings. As long as a mutation does not kill an organism before it reproduces (most mutations, in fact, do), it may be passed on to the next generation. Over time, this one moth had offspring. These, too, were dark. The dark- and the light-colored moths lived happily side by side until something from the outside—in our example, the environment—changed all that.
The initial variation came about by chance. This variation gave the dark moths an edge. However, that advantage did not become apparent until something made it apparent. In our case, that something was the intensive pollution from coal burning. The abundance of soot made it easier for predators to spot the light-colored moths, thus effectively removing them from the population. Therefore, dark color is an adaptation, a variation favored by natural selection.
Eventually, over long stretches of time, these two different populations might change so much that they could no longer reproduce together. At that point, we would have two different species, and we could say, definitively, that the moths had evolved. Evolution occured as a consequence of random mutation and the pressure put on the population by an environmental change. Catastrophic events can speed up natural selection and adaptation. When the rules change, sometimes the odd variations are selected for. Earth has undergone several mass extinction events. The tree of life would look much different if these events had not occurred.
Types of Selection
The situation with our moths is an example of directional selection. One of the phenotypes was favored at one of the extremes of the normal distribution.
In other words, directional selection “weeds out” one of the phenotypes. In our case, dark moths were favored, and light moths were practically eliminated. Here’s one more thing to remember: directional selection can happen only if the appropriate allele—the one that is favored under the new circumstances—is already present in the population. Two other types of selection are stabilizing selection and disruptive selection.
Stabilizing selection means that organisms in a population with extreme traits are eliminated. This type of selection favors organisms with common traits. It “weeds out” the phenotypes that are less adaptive to the environment. A good example is birth weight in human babies. An abnormally small baby has a higher chance of having birth defects; conversely an abnormally large baby will have a challenge in terms of a safe birth delivery.
Disruptive selection, on the other hand, does the reverse. It favors both the extremes and selects against common traits. For example, females are “selected” to be small and males are “selected” to be large in elephant seals. You’ll rarely find a female or male of intermediate size.
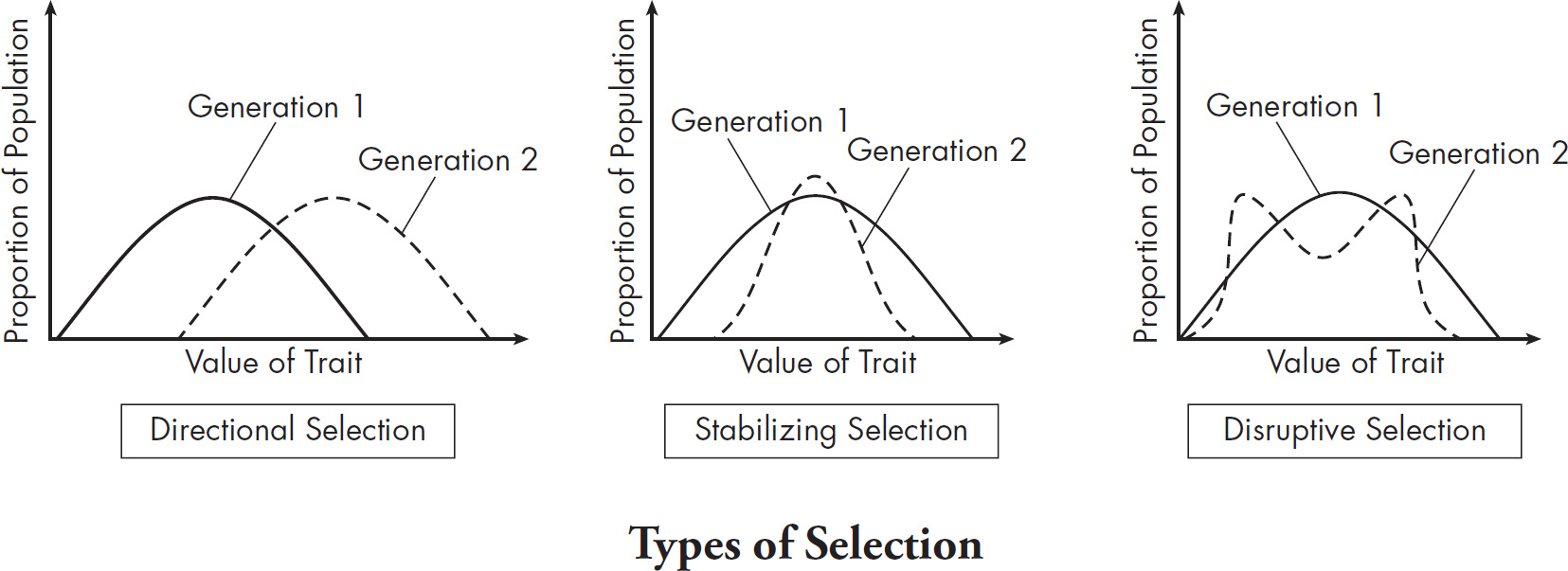
SPECIES
A dog and a bumblebee obviously cannot produce offspring. Therefore, they are different species. However, a poodle and a Great Dane can reproduce (at least in theory). We would not say that they are different species; they are merely different breeds. In order for these to become different species of dogs, they would have to become reproductively isolated from each other. This would then allow the two groups to undergo natural selection and evolve differently. With different variation and different environmental pressures, they could each change in different ways and no longer be able to mate. This is called divergent evolution.
Species often originate from a common ancestor. More often than not, the “engine” of evolution is cataclysmic environmental change, such as pollution in the case of the moths. Geographic barriers, new stresses, disease, and dwindling resources are all factors in the process of evolution. Pre- and post-zygotic barriers also prevent organisms of two different species from mating to produce viable offspring. Pre-zygotic barriers prevent fertilization. Examples of this kind of barrier include temporal isolation, or when two species reproduce at different times of the year. A post-zygotic barrier is related to the inability of the hybrid to produce offspring. For example, a horse and a donkey can mate to produce a mule, but mules are sterile and therefore cannot produce a second generation. Often plants undergo speciation due to polyploidy, which occurs when they get doubles of their chromosomes.
Convergent evolution is the process by which two unrelated and dissimilar species come to have similar (analogous) traits, often because they have been exposed to similar selective pressures. Examples of convergent evolution include aardvarks, anteaters, and pangolins. They all have strong, sharp claws and long snouts with sticky tongues to catch insects, yet they evolved from three completely different mammals.
There are two types of speciation: allopatric speciation and sympatric speciation. Allopatric speciation simply means that a population becomes separated from the rest of the species by a geographic barrier so that the two populations can’t interbreed. An example would be a mountain that separates two populations of ants. In time, the two populations might evolve into different species. If, however, new species form without any geographic barrier, it is called sympatric speciation. This type of speciation is common in plants. Two species of plants may evolve in the same area without any geographic barrier.
POPULATION GENETICS
Mendel’s laws can also extend to the population level. Suppose you caught a bunch of fruit flies—about 1,000. Let’s say that 910 of them were red-eyed and 90 were green-eyed. If you allowed the fruit flies to mate and counted the next generation, we’d see that the ratio of red-eyed to green-eyed fruit flies would remain the same: 91 percent red-eyed and 9 percent green-eyed. That is, the allele frequency would remain constant. At first glance you may ask, how could that happen?
The Hardy-Weinberg law states that even with all the shuffling of genes that goes on, the relative frequencies of genotypes in a population still prevail over time. The alleles don’t get lost in the shuffle. The dominant gene doesn’t become more prevalent, and the recessive gene doesn’t disappear.
Hardy-Weinberg Equilibrium
The Hardy-Weinberg law says that a population will be in genetic equilibrium only if it meets these five conditions: (1) a large population, (2) no mutations, (3) no immigration or emigration, (4) random mating, and (5) no natural selection.
When these five conditions are met, the gene pool in a population is pretty stable. Any departure from them results in changes in allele frequencies in a population. For example, if a small group of your fruit flies moved to a new location, the allele frequency may be altered and result in evolutionary changes. That’s an example of genetic drift called the founder effect. In other words, the gene frequency may differ from the original gene pool. Genetic drift often occurs in new colonies with small populations. You can think of genetic drift as changes to the alleles in a population due to random luck rather than natural selection. Things that really change the population size are called bottleneck events. If most of a species is killed by some crazy weather event, whichever individuals are left now make up the new gene pool, even if they were not previously being selected for.
Hardy-Weinberg Equations
Let’s say that the allele for red eyes, R, is dominant over the allele for green eyes, r. Red-eyed fruit flies include homozygous dominants, RR, and heterozygous, Rr. The green-eyed fruit flies are recessive, rr. The frequency of each allele is described in the equation below. The allele must be either R or r. Let “p” represent the frequency of the R allele and “q” represent the frequency of the other allele in the population.
p + q = 1
This sum of the frequencies must add up to one. If you know the value of one of the alleles, then you’ll also know the value of the other allele. This makes sense because Hardy-Weinberg ONLY works for populations with 2 alleles that have normal dominant-recessive behavior. So, if you know there are only 2 alleles possible and you know that 70% are the dominant allele, then the remaining 30% must be the recessive allele.
We can also determine the frequency of the genotypes in a population using the following equation.
p2 + 2pq + q2 = 1
In this equation, p2 represents the homozygous dominants, 2pq represents the heterozygotes, and q2 represents the homozygous recessives.
Both of these equations are listed on the AP Biology Equations and Formulas sheet. But how do you use them? Use the proportions in the population to figure out both the allele and genotype frequencies. Let’s calculate the frequency of the genotype for green-eyed fruit flies. If 9 percent of the fruit flies are green-eyed, then the genotype frequency, q2, is 0.09. You can now use this value to figure out the frequency of the recessive allele in the population. The allele frequency for green eyes is equal to the square root of 0.09—that’s 0.3. If the recessive allele is 0.3, the dominant allele must be 0.7. That’s because 0.3 + 0.7 equals 1.
Using the second equation, you can calculate the genotypes of the homozygous dominants and the heterozygotes. The frequency for the homozygous dominants, p2, is 0.7 × 0.7, which equals 0.49. The frequency for the heterozygotes, 2pq, is 2 × 0.3 × 0.7, which equals 0.42. If you include the frequency of the recessive genotype—0.09—the numbers once again add up to 1.
KEY TERMS
evolution
natural selection
Charles Darwin
Jean-Baptiste de Lamarck
paleontology
biogeography
flora
fauna
embryology
comparative anatomy
homologous structures
analogous structures
molecular biology
common ancestor
phylogenetic tree
genetic variability
conjugation
transduction
Peppered moths
environmental pressure
random mutation
adaptation
directional selection
stabilizing selection
disruptive selection
species
speciation
reproductively isolated
divergent evolution
pre-zygotic barriers
post-zygotic barriers
convergent evolution
allopatric speciation
sympatric speciation
Hardy-Weinberg law
genetic drift
Summary
 Charles Darwin made the following key observations that led to his theory of natural selection:
Charles Darwin made the following key observations that led to his theory of natural selection:
• Each species produces more offspring than can survive.
• Offspring compete with each other for limited resources.
• Organisms in every population vary.
• The offspring with the most favorable traits are most likely to survive and reproduce.
 Evidence for evolution includes:
Evidence for evolution includes:
• fossils
• biogeography
• comparison of developmental embryology
• comparative anatomy, including homologous and analogous structures
• molecular biology (sequences of genes are conserved across many types of species)
 Members of a species are defined by the ability to reproduce to produce fertile offspring. Evolution and speciation occur when environmental pressures favor traits that permit survival and reproduction of selected individuals in a varied population. The favored traits may evolve to cause divergent or convergent evolution.
Members of a species are defined by the ability to reproduce to produce fertile offspring. Evolution and speciation occur when environmental pressures favor traits that permit survival and reproduction of selected individuals in a varied population. The favored traits may evolve to cause divergent or convergent evolution.
 The Hardy-Weinberg equations can be used to determine genetic variation within a population. These equations are as follows:
The Hardy-Weinberg equations can be used to determine genetic variation within a population. These equations are as follows:
| • p + q = 1 | Frequency of the dominant (p) and recessive (q) alleles |
| • p2 + 2pq + q2 = 1 | Frequency of the homozygous dominants (p2), heterozygotes (2pq), and homozygous recessives (q2) |
 The Hardy-Weinberg equations describe a population that is not evolving but is instead said to be in genetic equilibrium. Such a population will only be so if it is not violating any of the following five conditions (almost all populations are evolving due to a violation of one or more of these factors):
The Hardy-Weinberg equations describe a population that is not evolving but is instead said to be in genetic equilibrium. Such a population will only be so if it is not violating any of the following five conditions (almost all populations are evolving due to a violation of one or more of these factors):
• large population
• no mutations
• no immigration or emigration
• random mating
• no natural selection
Chapter 10 Drill
Answers and explanations can be found in Chapter 16.
1. The eye structures of mammals and cephalopods such as squid evolved independently to perform very similar functions and have similar structures. This evolution is an example of which of the following?
(A) Allopatric speciation
(B) Sympatric association
(C) Divergent evolution
(D) Convergent evolution
Questions 2-5 refer to the following graph and paragraph.
During the industrial revolution, a major change was observed in many insect species due to the mass production and deposition of ash and soot around cities and factories. One of the most famous instances was within the spotted moth population. An ecological survey was performed where the number of spotted moths and longtail moths were counted in 8 different urban settings over a square kilometer in 1802. A repeat experiment was performed 100 years later in 1902. The results of the experiment are shown below.
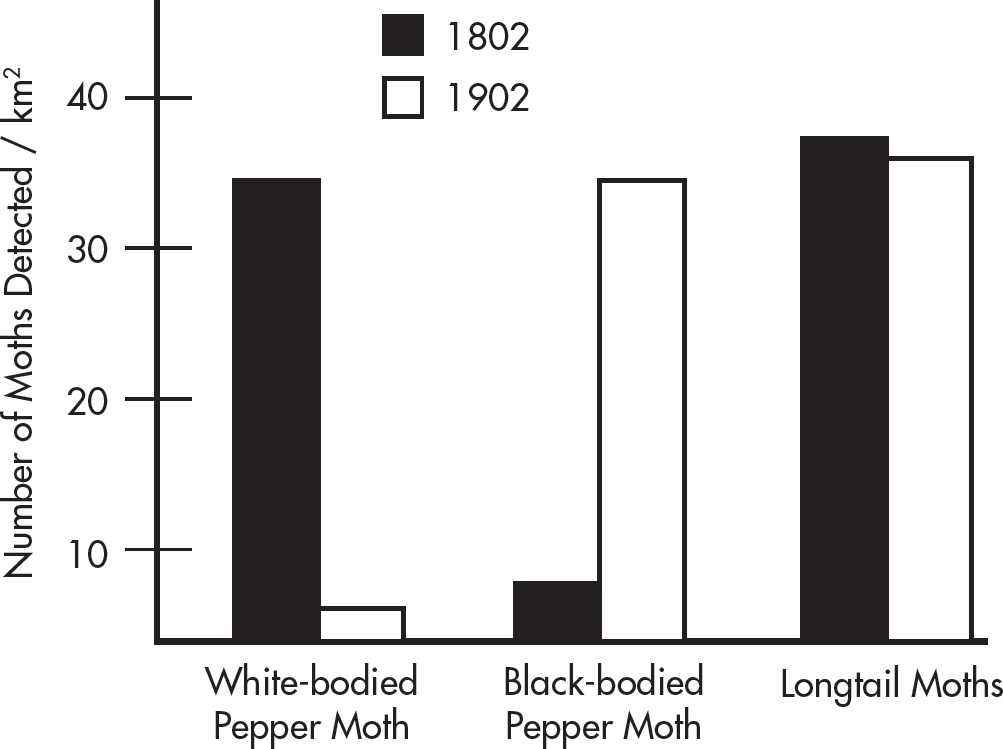
2. What type of selection is represented by the results of this study?
(A) Stabilizing selection
(B) Directional selection
(C) Disruptive selection
(D) Divergent selection
3. Which of the following statements best explains the data?
(A) As time passed from 1802 to 1902, the frequency of white-bodied pepper moths increased and black-bodied pepper moths decreased.
(B) As time passed from 1802 to 1902, the frequency of white-bodied pepper moths decreased and black-bodied pepper moths increased.
(C) As time passed from 1802 to 1902, the frequency of white-bodied pepper moths and black-bodied pepper moths both increased.
(D) As time passed from 1802 to 1902, the frequency of white-bodied pepper moths and black-bodied pepper moths both decreased.
4. Why was the population of longtail moths also surveyed in this study?
(A) Variations in the environment were expected to alter the population of longtail moths.
(B) Longtail moths were included as a control because they were not expected to change appreciably due to changes associated with the Industrial Revolution.
(C) As time passed from 1802 to 1902, the frequency of white-bodied pepper moths and black-bodied pepper moths both increased.
(D) As time passed from 1802 to 1902, the frequency of white-bodied pepper moths and black-bodied pepper moths both decreased.
5. How would the results of this study have been different if factories produced white or light gray ash and soot rather than black?
(A) There would be no change to the results of the experiment.
(B) There would have been added selection pressure for more white-bodied spotted moths and against black-bodied spotted moths.
(C) There would have been added selection pressure for more black-bodied spotted moths and against white-bodied spotted moths.
(D) There would have been an increase in the frequency of both black-bodied and white-bodied spotted moths.
Question 6 represents a question requiring a numeric answer.
6. A recessive allele of a gene has a calculated frequency of 0.3 in a population. Assuming the population is in Hardy-Weinberg equilibrium, what percentage of the population is expected to be heterozygous for the gene? Give your answer to the nearest hundredths place.

7. The Middle East blind mole rat (Nannosplalax ehrenbergi) lives in the Upper Galilee Mountains of Israel. Two groups of these mole rats live in the same region, but scientists discovered that there is a 40% difference in mitochondrial DNA between these two groups. These rats do not seem to interbreed in the wild. This is an example of
(A) allopatric speciation
(B) sympatric speciation
(C) prezygotic barrier
(D) hybrid zone
Questions 8 and 9 refer to the following scenario.
A cruise ship with 500 people on board crashes on a deserted island. They like the island so much that they decide to stay. There is a small number of people on board with red hair, an autosomal recessive trait.
8. What is most likely to occur with respect to the number of red-haired individuals on the island?
(A) There will be no change in the overall number of red-haired people in the population over time.
(B) Convergent evolution will occur, with all people eventually becoming red-haired.
(C) It is likely that over many generations, the proportion of red-haired people in the population will increase.
(D) Red-haired people will probably decrease in the population over many generations.
9. Evolution does not occur when a population is said to be in equilibrium by the Hardy-Weinberg definition. In the situation above, which of the following does not support the claim that evolution will occur in this population?
(A) The situation describes a small population.
(B) Mutations occur in this population.
(C) People do not usually mate randomly.
(D) Natural selection pressures do not exist on this island.
10. Peacock males have evolved to have huge, beautiful tails with numerous large eye spots. Long, colorful tails are difficult to carry when you are running from a predator. However, peahens (female peacocks) have been shown by researchers to mate preferably with males with more eye spots and longer tails, despite these traits making the males more susceptible to predation. Over time, this preference has resulted in peacocks with huge tails due to
(A) sexual selection
(B) disrupted selection
(C) divergent evolution
(D) sympatric speciation
REFLECT
Respond to the following questions:
• Which topics from this chapter do you feel you have mastered?
• Which content topics from this chapter do you feel you need to study more before you can answer multiple-choice questions correctly?
• Which content topics from this chapter do you feel you need to study more before you can effectively compose a free response?
• Was there any content that you need to ask your teacher or another person about?
Chapter 11
Animal Structure and Function
THE STRUCTURE AND FUNCTION OF ORGANISMS
To carry on with the business of life, higher organisms must all contend with the same basic challenges: obtaining nutrients, distributing them throughout their bodies, voiding wastes, responding to their environments, and reproducing. To accomplish these basic tasks, nature has come up with solutions.
Unicellular and multicellular organisms alike meet these challenges through organization and specialization. We already learned about the organelles within a cell allowing compartmentalization. Each organelle does it own task, and they work together to keep the cell alive. In many animals and humans, there are higher levels of organization.
Homeostasis
All living things have a “steady-state” that they try to maintain. This is helpful because the structures and processes within living things are sensitive to things like temperature, pH, pressure, salinity, osmotic pressure, and many other things. There is a set range of conditions that living things can live in successfully, called homeostasis. The body is constantly working to maintain this state by taking measurements and then responding appropriately. That might mean shivering if it is cold or releasing the hormone insulin if blood glucose is high. Depending on the condition, the body will respond accordingly.
Many of these responses are controlled by negative or positive feedback pathways. A negative feedback pathway (also called feedback inhibition) works by turning itself off using the end product of the pathway. The end-product inhibits the process beginning again, shutting down the pathway. This is a common strategy to conserve energy. If plenty of X is made, then the pathway to make X can be turned off. Sometimes, a very distant end product will turn off a process occuring several steps back. For example, ATP turns off the beginning stages of glycolysis, although ATP is not made until much later in cellular respiration.
A positive feedback pathway also involves an end-product playing a role, but instead of inhibiting the pathway, it further stimulates it. For example, once X is made, it tells the pathway to make even more X. This is less common, but it occurs during fruit ripening and labor and delivery, which ramps up and up as it proceeds.
DEVELOPMENT
For the AP test, it is important to have a general idea of how the body works. Let’s start at the beginning.
Embryonic Development
How does a tiny, single-celled egg develop into a complex, multicellular organism? By dividing, of course. The cell changes shape and organization many times by going through a succession of stages. This process is called morphogenesis.
When an egg is fertilized by a sperm, it forms a diploid cell called a zygote.
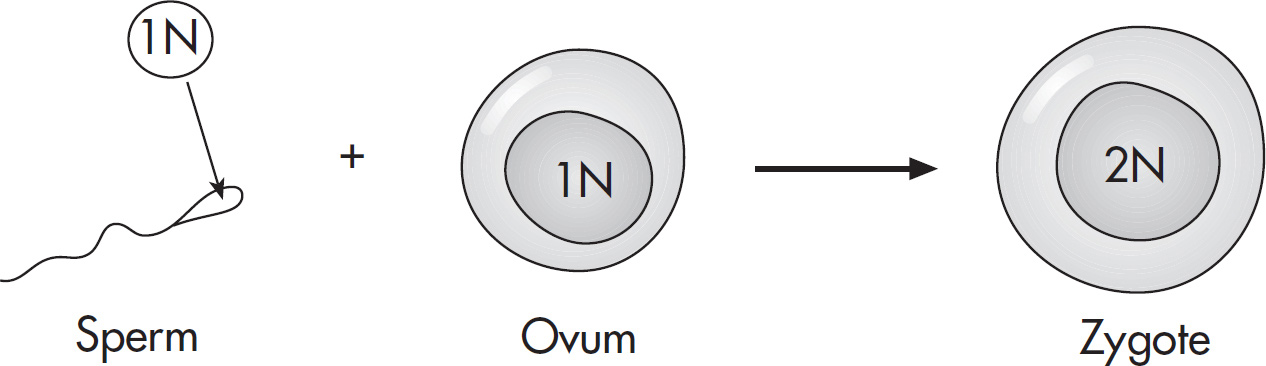
Fertilization triggers the zygote to go through a series of divisions. As these occur, the embryo becomes increasingly differentiated, or specialized. An undifferentiated cell is like a blank slate—it can become any type of cell. Once a cell begins to specialize, it is limited in its future options.
In order for a cell to differentiate, it must change. Certain genes will be expressed, and other genes might be turned off. Cells called organizers release signals that let each cell know how they should develop. As an over-simplified example, a cell destined to become a muscle might have an increase in things that make it flexible, while something destined to become bone might have a decrease in those things. Once the change has been made, the future muscle cell can’t change its mind and become a bone cell.
The early genes that turn certain cells in the early embryo into future-this or future-that are called homeotic genes. A subset of homeotic genes are called Hox genes. The timing is essential for the induction of these genes. Exactly the right bit of the embryo must be modified at exactly the right time, or the embryo may develop with the brain in the wrong place or too many limbs or only limbs on one side of the body. Severely damaged embryos will stop developing.
An interesting additional tool that the developing embryo uses is apoptosis, or programmed cell death. Certain bits of the developing embryo are used as a scaffold for development (like those between your fingers and toes), and when they are no longer necessary, those cells commit suicide. Think of it like an eraser coming along and erasing the webbing tissue between fingers and toes.
BODY SYSTEMS
Once the embryo develops into a fully formed organism, it is highly specialized. Groups of cells that all perform the same function are called a tissue, and several tissues can come together to form specialized structures called organs. Several organs then come together to form body systems.
Fortunately, you only need to know details about the immune system, the nervous system, and the endocrine system. Here is a brief list of other major systems so you get the gist of their function:
Circulatory: Blood vessels (arteries, capillaries, and veins) carry blood around the body to transport chemical signals and to bring supplies to cells and carry waste away. The blood flow is controlled by the heart, which pumps the blood through the blood vessels.
Respiratory: The lungs are responsible for gas exchange (O2 and CO2), also helps maintain pH levels in the blood.
Digestive: The esophagus, stomach, small intestine, large intestine, pancreas, liver, and gall bladder work together to break food down and absorb nutrients. The stomach does mixing and breakdown. The small intestine does most of the absorption. The pancreas makes lots of enzymes.
Excretory: The kidneys filter the blood and reabsorb things that the body wants to keep and get rid of the rest in urine.
Reproductive: The male and female systems are largely controlled by hormones and allow the production of gametes and the ability to reproduce.
Muscular and Skeletal: The body has skeletal muscle and smooth muscles that each contract by action potential signals from the nervous system. The skeletal system provides structure, protection, and calcium storage.
THE IMMUNE SYSTEM
The immune system is the body’s defense system. It is a carefully and closely coordinated system of specialized cells, each of which plays a specific role in the war against bodily invaders. Let’s take a brief sidetrack and talk a bit about these invaders. Pathogens are disease-causing biological agents that can generally be divided into bacteria, viruses, fungi, and parasites.
Bacteria
We have been talking on and off about bacteria already. They are prokaryotes that come in many shapes and sizes. They can infect many things, and sometimes they cause harm and sometimes they do not. You may have heard of “gut bacteria” before; this is a special colony of bacteria that lives inside each one of us. It helps us with some digestion and makes some things we need. We have a mutualistic relationship with our gut bacteria.
Bacteria divide by fission; however, this does not increase their genetic diversity. Instead, they can perform conjugation with other bacterial cells and swap some of their DNA. Genetic variety among bacteria is leading to increased antibiotic resistance. As we mentioned in Chapter 10, an increase in genetic variation increases the likelihood that a population will survive a catastrophic event.
VIRUSES
Viruses are nonliving agents capable of infecting cells. Why are viruses considered nonliving? They require a host cell’s machinery in order to replicate. A virus consists of two main components: a protein capsid and genetic material made of DNA or RNA, depending on the virus. Viruses are all very specific in which type of cells they infect, and the thing infected by a virus is called a host.
Viruses have one goal: replicate/spread. In order to do this, a virus needs to make more genome and make more capsid. They then assemble together into new viral particles.
The viral genome carries the genes for building the capsid and anything else the virus needs that the host cannot provide. Sometimes, if two viruses infect the same cell, there will be mixing of he genomes, especially if the viruses that have genomes split between several chromosome-like segments. A new virus particle might emerge that is a blend of the two viruses.
A commonly studied virus is a bacteriophage (a virus that infects bacteria). Bacteriophages undergo two different types of replication cycles, the lytic cycle and the lysogenic cycle. In the lytic cycle, the virus immediately starts using the host cell’s machinery to replicate the genetic material and create more protein capsids. These spontaneously assemble into mature viruses and cause the cell to lyse, or break open, releasing new viruses into the environment. In the lysogenic cycle, the virus incorporates itself into the host genome and remains dormant until it is triggered to switch into the lytic cycle. A virus can hide in the genome of a bacteria cell for a very long time. During this time, the cell may divide and replicate the virus as well. By the time the lytic cycle is triggered the virus may have been replicated many many times as the cell hosting it divides.
When a virus excises from a host genome (becomes unintegrated), it sometimes accidentally takes some of the bacterial cell’s DNA with it. Then, that gets replicated and packaged into new viral particles with the viral genome. The next cell that gets infected is not only getting infected with the viral genome, but also with that chunk of bacterial DNA. If that chunk held a gene for something like antibiotic resistance, the next cell that gets infected will gain that trait. The transfer of DNA from a virus to a bacterial cell is called transduction.
Viruses that infect animals do not have to break their way out of the cell the same way that a bacteriophage does. Since animals cells don’t have a cell wall, the viruses often just “bud” out of the membrane like exocytosis. When a virus does this, it becomes enveloped by a chunk of cell membrane that it takes with it. Viruses with a lipid envelope are called enveloped viruses.
Retroviruses like the HIV virus are RNA viruses that use an enzyme called reverse transcriptase to convert their RNA genomes into DNA so that they can be inserted into a host genome. RNA viruses have extremely high rates of mutation because they lack proofreading mechanisms when they replicate their genomes of mutation. This high rate of mutation will create lots of variety, which makes these viruses difficult to treat. As was discussed in Chapter 10, they evolve quickly as drug-resistant mutations become naturally selected. New drugs must constantly be identified to treat the resistance.
Foreign molecules—be they viral, bacterial, or chemical—that can trigger an immune response are called antigens. Humans and other vertebrates have two types of immune responses: the innate immune response and the adaptive/specific immune response. The innate is more of a general anti-invader response. The body recognizes certain things that are common of foreign things and destroys them. The adaptive response carefully catalogs and handles each antigen in a particular way. It also has a memory component that helps it fight repeat attackers very efficiently.
THE TWO IMMUNE RESPONSES
The Innate Immune System
The body’s first line of defense against foreign substances is the skin and the mucous lining of the respiratory and digestive tracts. If these defenses are not sufficient, other nonspecific defense mechanisms are activated. These include phagocytes such as macrophages (which engulf antigens), complement proteins (which lyse the cell wall of the antigen), interferons (which inhibit viral replication and activate surrounding cells that have antiviral actions), and inflammatory response (a series of events in response to antigen invasion or physical injury). Activation of these defenses requires immune cells to recognize the foreign substance (usually via receptor binding) and activate intracellular signaling pathways. This is another example of how important cell communication is to survival. These first line defenses are called the innate immune system, and many organisms have this level of defense. The specifics vary with organism structure, but there are several common themes. For example, a thick outer surface usually protects the organism, such as the skin of a human, the cell wall of a bacteria, or the cuticle of a plant. In all organisms, cell communication pathways are essential to recognize “self” cells, recognize foreign substances, and initiate immune responses when under attack by a pathogen (a disease-causing agent) or chemical.
The Adaptive/Specific Immune Response
Lymphocytes are the primary cells of the immune system. They are found mostly in the blood and lymph nodes and are a type of white blood cell (or leukocyte). There are two types of lymphocytes: B-cells and T-cells. When an individual becomes infected by a pathogen, B- and T-lymphocytes get activated.
B-lymphocytes mature in bone marrow and are involved in the humoral response, which defends the body against pathogens present in extracellular fluids, such as lymphatic fluid or blood. Each B-cell has a special receptor on its surface that can only bind to foreign things. If a pathogen arrives that fits the receptor, the B-cell becomes activated (with the help of a T-cell). The B-cell will begin to replicate and seek out more of the pathogens. Some B-cells become memory B-cells that remain in circulation, allowing the body to mount a quicker response if a second exposure to the same pathogen should occur. Other B-cells become plasma cells that produce antibodies, which are specific proteins that bind to antigens on the surface of pathogens that originally activated them. The antibody that is made by each B-cell is identical to the the surface receptor that caught the antigen. Each B-cell has a unique receptor/antibody that it makes.
Antibodies all have the same basic monomer structure that is shaped like the letter Y. The stem of the Y is always the same and can interact with other cells in the immune response. The arms of the Y are always unique because this is where the antibody binds antigen. On each antibody, both arms bind the same shape so that it can hold two antigens at once. As mentioned, this Y shape is just the monomer of an antibody. Sometimes antibodies can have one, two, or five Y-monomers with the stem parts attaching and the arms facing outward. Antibodies are typically made only when the appearance of antigens in the body stimulates a defense mechanism, but they will linger around after an infection. When an antibody binds to the antigen it is made for, it marks the antigen for destruction. Antibodies can also combat the antigen just by binding to it because this alone can wreak havoc for the pathogen.
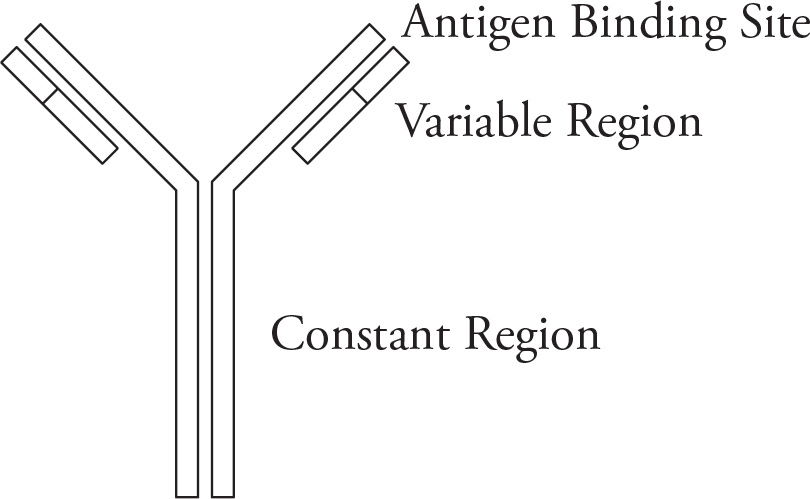
T-lymphocytes, maturing in the thymus, are involved in cell-mediated immunity; this branch of the immune system is responsible for monitoring “self” cells to make sure they are still healthy. The plasma membrane of cells has major histocompatibility complex (MHC) markers that allow T-cells to get a glimpse of what is happening inside each cell. T-cells have special antigen-recognizing receptors just like B-cells, and if the receptors see something foreign, the T-cells jump into action.
MHC I are on all nucleated cells. They take peptides that have been found inside the cell and hold them up on the surface so that a type of circulating T-cell called a cytotoxic T-cell can inspect them. If they decide the cell is infected or is a tumor cell, they signal for it to be destroyed by apoptosis, which is a careful self-destruct program.
MHC II are on special immune cells, like B-cells and macrophages, that identify and engulf antigens. These types of cells are called antigen-presenting cells. They hold up a chunk of whatever antigen the immune cell has picked-up and let a type of T-cell called a helper T-cell inspect it. If the T-cell agrees that it is something foreign, it helps activate the immune cell. Think of the helper T-cell as a double-checker. This interaction between different branches of the immune system requires B-cells and T-cells to recognize each other and coordinate an immune attack—another process tightly controlled by cell signaling and communication pathways.
A lymph node is a mass of tissue found along a lymph vessel. A lymph node contains a large number of lymphocytes. Because lymphocytes are important in fighting infection, they multiply rapidly when they come in contact with an antigen. As a consequence, lymph nodes swell when they’re fighting an infection. That’s why when you have a sore throat, one of the first things a doctor does is touch the sides of your throat to see if your lymph nodes are swollen, a probable sign of infection.
One thing to remember about immune cells and blood cells: all blood cells, white and red, are produced in the bone marrow. However, red blood cells (or erythrocytes) play no role in the immune system; instead, they help transport oxygen throughout the body. They are full of hemoglobin.
Vaccines
After an infection, the adaptive immune response keeps around groups of cells called memory B-cell and memory T-cells. They act as a militia and keep their eyes peeled for the same antigens they have seen before. The second time that you are infected with a pathogen, you will usually fight it off very quickly. You might not even know you were even infected because your body destroys it so fast. Vaccines are tiny doses of antigen that have been modified so they are not dangerous. Your immune system is introduced to them and prepares a militia, but you don’t ever get the “real” infection. If you do come into contact with the real version, your body will fight it off ASAP. In a nutshell, getting vaccinated is getting a tiny harmless dose now so you don’t get knocked off your feet by the big, bad version later.
AIDS
AIDS, or “acquired immunodeficiency syndrome,” is a devastating disease that interferes with the body’s immune system. AIDS is caused by the specific infection of helper T-cells by human immunodeficiency virus (HIV). The virus essentially wipes out helper T-cells, preventing the body from defending itself. Those afflicted with AIDS do not die of AIDS itself, but rather of infections that they can no longer fight off, due to their compromised immune systems.
Auto-Immunity
The B-cells and T-cells are carefully created so that they do not react to “self” and only react to foreign things. However, sometimes mistakes happen and the immune system begins to think that “self” is dangerous. These attacks cause auto-immune diseases such as Type I diabetes, rheumatoid arthritis, lupus, and celiac disease.
THE NERVOUS SYSTEM
All organisms must be able to react to changes in their environment. As a result, organisms have evolved systems that collect and process information from the outside world. The task of coordinating this information falls to the nervous system. The simplest nervous system is found in the hydra. It has a nerve net made up of a network of nerve cells, which sends impulses in both directions. As animals became more complex, they developed clumps of nerve cells called ganglia. These cells are like primitive brains. More complex organisms, like humans, have a brain with specialized cells called neurons.
Neurons
The functional unit in the nervous system is a neuron. Neurons receive and send the neural impulses that trigger organisms’ responses to their environments. Let’s talk about the parts of a neuron. A neuron consists of a cell body, dendrites, and an axon.
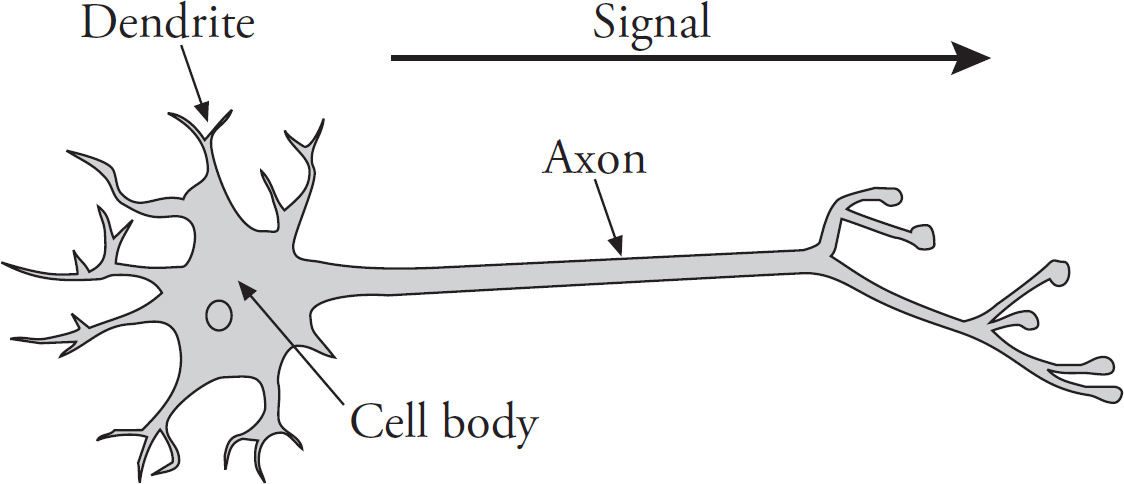
The cell body contains the nucleus and all the usual organelles found in the cytoplasm. Dendrites are short extensions of the cell body that receive stimuli. The axon is a long, slender extension that transmits an impulse from the cell body to another neuron or to an organ. A nerve impulse begins at the top of the dendrites, passes through the dendrites to the cell body, and moves down the axon.
How Neurons Communicate
Within a neuron, the signal is called an action potential, which is a wave of positive charge that sweeps down the axon. In order for the signal to be clear, the cell must have a “normal,” non-positive state. Nearly all cells in the body have a natural resting state that is negative. The natural charge of a cell is called the resting membrane potential, which represents the difference in charge from inside the cell to outside the cell.
The resting potential arises from two activities:
• The Na+K+-ATPase pump: This pump pushes two potassium ions (K+) into the cell for every three sodium ions (Na+) it pumps out of the cell, which leads to a net loss of positive charges within the cell.
• Leaky K+ channels: Some potassium channels in the plasma membrane are “leaky,” allowing a slow diffusion of K+ out of the cell.
Both the Na+K+-ATPase pump and the leaky channels cause a potential difference between the inside of the neuron and the surrounding interstitial fluid. The resting membrane potential is always negative inside the cell, and the neuronal membrane is said to be polarized. In humans, the negative charge is –70mV.
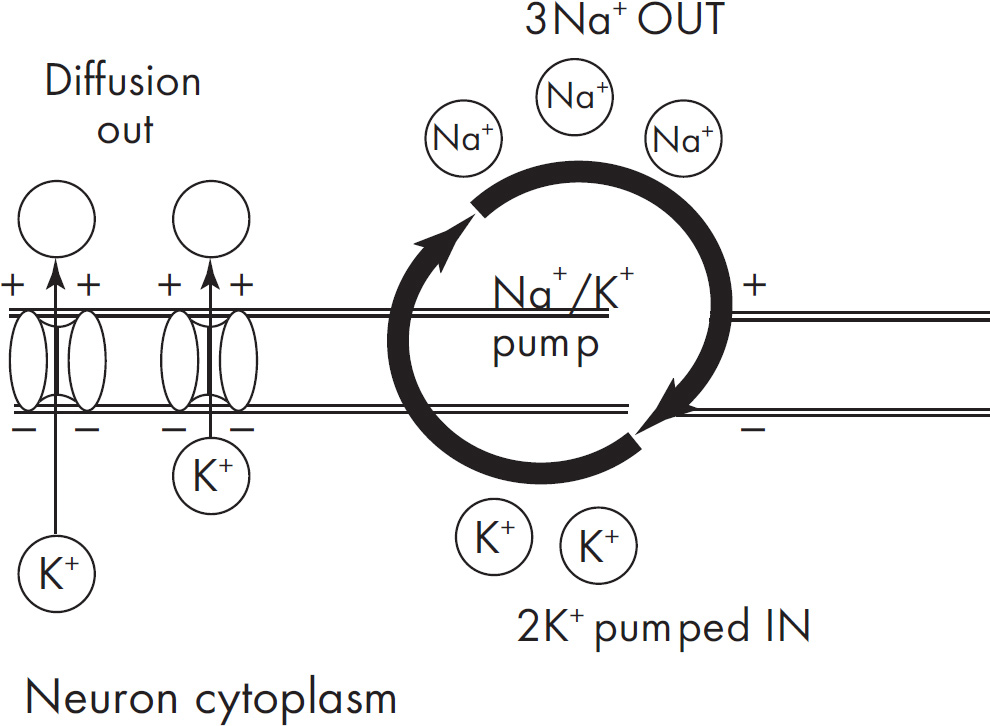
Action Potential
Here’s what a neuron does in response to a stimulus. If a stimulus has enough intensity to excite a neuron, the cell reaches its threshold—the minimum amount of stimulus a neuron needs to respond. When the threshold is reached, the cell “fires” an action potential. The action potential is an all-or-none response—it doesn’t fire “part way.”
(1) Reaching threshold: An outside stimulus causes a slight influx of positive charge in the neuron. When this influx causes the cell body of the neuron to reach −50mV, threshold is reached.
(2) Sodium channels open: The −50mV voltage causes the opening of many voltage-gated sodium channels near where the axon meets the cell body. Remember, the sodium potassium pump is actively pumping sodium out of the cell. This means that it is pumping sodium against the concentration gradient, meaning there is already a lot of sodium outside the cell. Therefore, the sodium wants to flow into the cell. When these channels are opened at threshold, a ton of sodium rushes into the cell. This causes the inside of the cell to become very positive (since sodium is positively charged). This is called depolarization. When the cell reaches +35mV inside, these sodium channels close again.
(3) Potassium channels open: As the sodium channels close, the potassium channels open. Potassium flows the opposite direction of sodium, out of the cell. This outflux of positives causes the cell to become negative again. This is called repolarization. When the voltage reaches arond −90mV, the potassium channels close, and the cell returns to resting membrane potential.
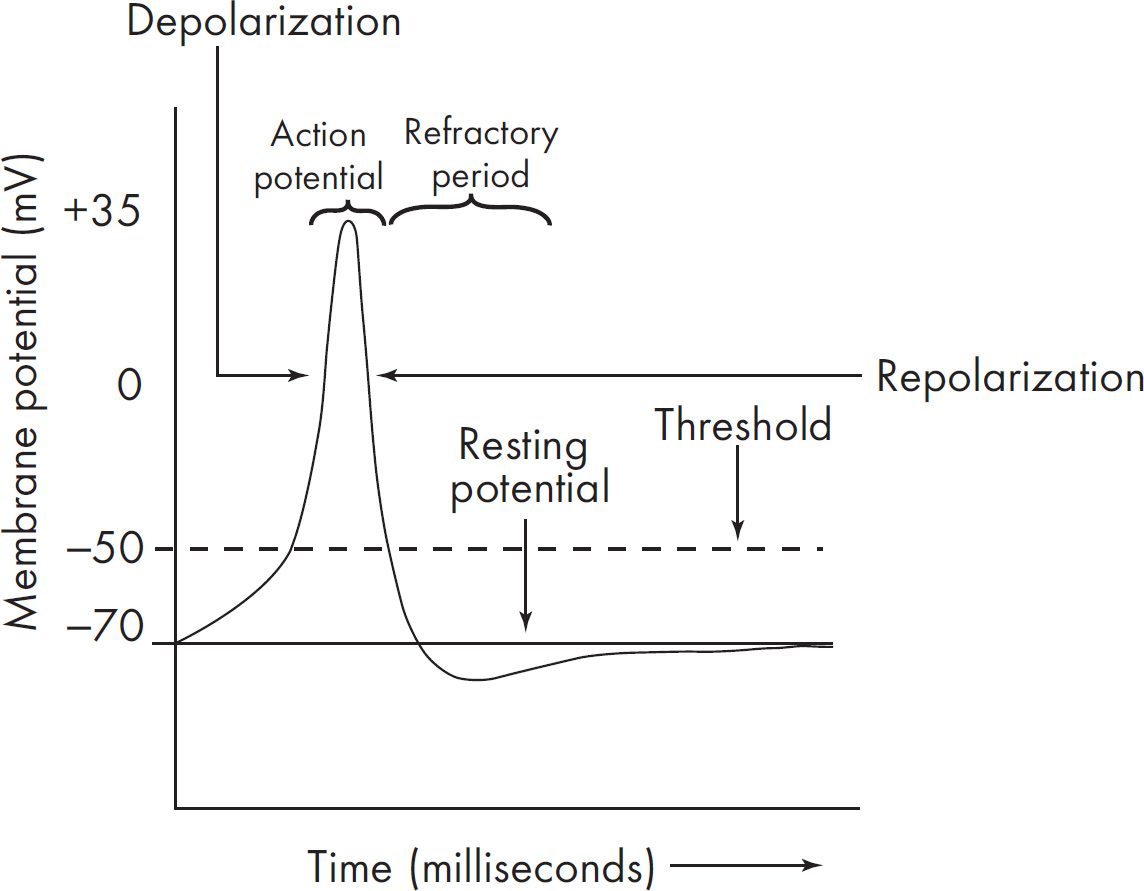
The Refractory Period
The period after an action potential is known as the refractory period. There are two different refractory periods. The first is caused because the sodium channels are unable to open again right away. There are in lockdown mode for a short time. The second is caused because the cell dips down below the resting membrane potential and is super negative. A greater stimulus is required to reach the threshold, so it is more difficult to initiate another action potential.
To summarize:
• A “resting” neuron is polarized; that is, the charge is more negative on the inside of the cell than on the outside.
• When an action potential comes along, the neuron transmits an impulse down its axon.
• First, voltage-gated sodium channels open, allowing sodium ions to rush in. This is known as depolarization: the neuron becomes more positive on the inside and more negative on the outside.
• Sodium channels close, and potassium channels open, which restores its negative charge. This is known as repolarization.
• The neuron enters a refractory period.
• The neuron reestablishes the ion distribution thanks to the sodium-potassium pump.
Passing Along the Signal
When one small area is depolarized, it causes a “domino effect.” The action potential spreads to the rest of the axon. The impulse is transmitted down the axonal membrane until it reaches the end of the axon called the axon bulb. Let’s discuss how the neuron manages to pass the impulse to the next neuron.
When an impulse reaches the end of an axon, the axon releases a chemical called a neurotransmitter into the space between the two neurons. This space is called a synapse. The cell before the synapse is called the presynaptic cell, and the cell after the synapse is called the postsynaptic cell. The neurotransmitter diffuses across the synaptic cleft and binds to receptors on the dendrites of the next neuron.
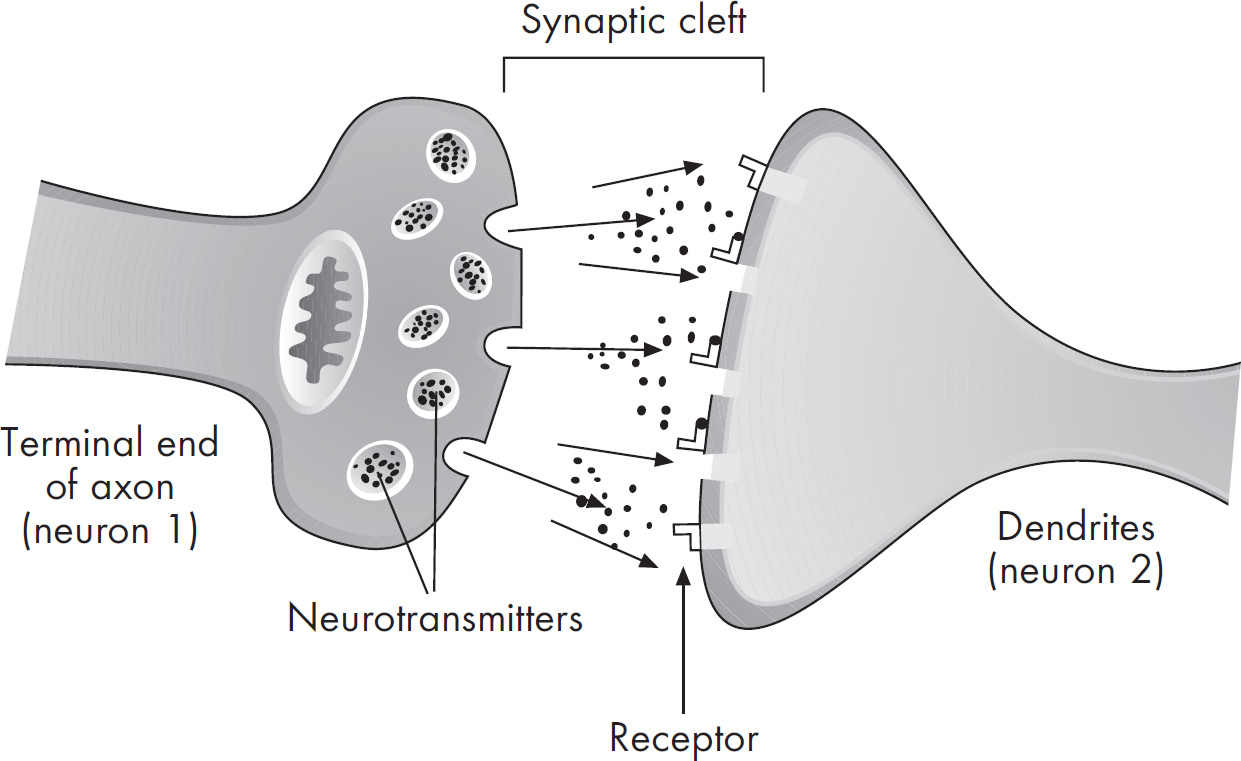
This will trigger an action potential in the second neuron if the postsynaptic membrane is excited. However, sometimes the signal will cause the postsynaptic membrane to become negatively charged (inhibited). Now the impulse moves along the second neuron from dendrites to axon.
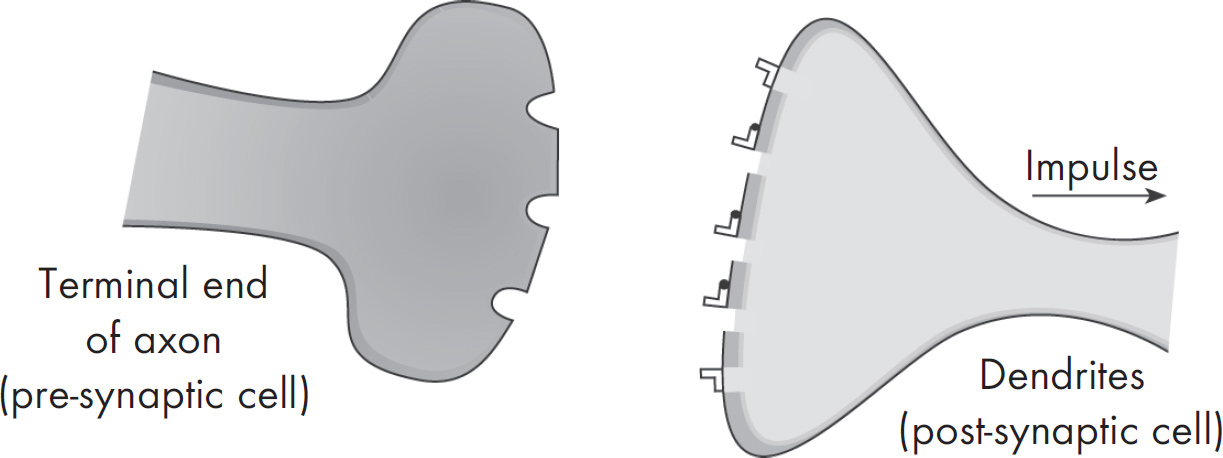
There are many neurotransmitters, but the most important one for the AP Biology Exam is called acetylcholine. Acetylcholine is a neurotransmitter that:
• is released from the end of an axon when Ca2+ moves into the terminal end of the axon
• is picked up almost instantly by the dendrites of the next neuron
• can stimulate muscles to contract
• is released between neurons in the parasympathetic system, which we will discuss shortly
The extra acetylcholine in the synaptic cleft is broken down by the enzyme acetylcholinesterase.
Other important neurotransmitters include norepinephrine and GABA. Norepinephrine is a peptide neurotransmitter that is released between neurons within the central nervous system. GABA is secreted in the central nervous system and acts as an inhibitor.
Speed of an Impulse
Sometimes a neuron has supporting cells that wrap around its axon. These cells are called Schwann cells. Schwann cells produce a substance called the myelin sheath, which insulates the axon.
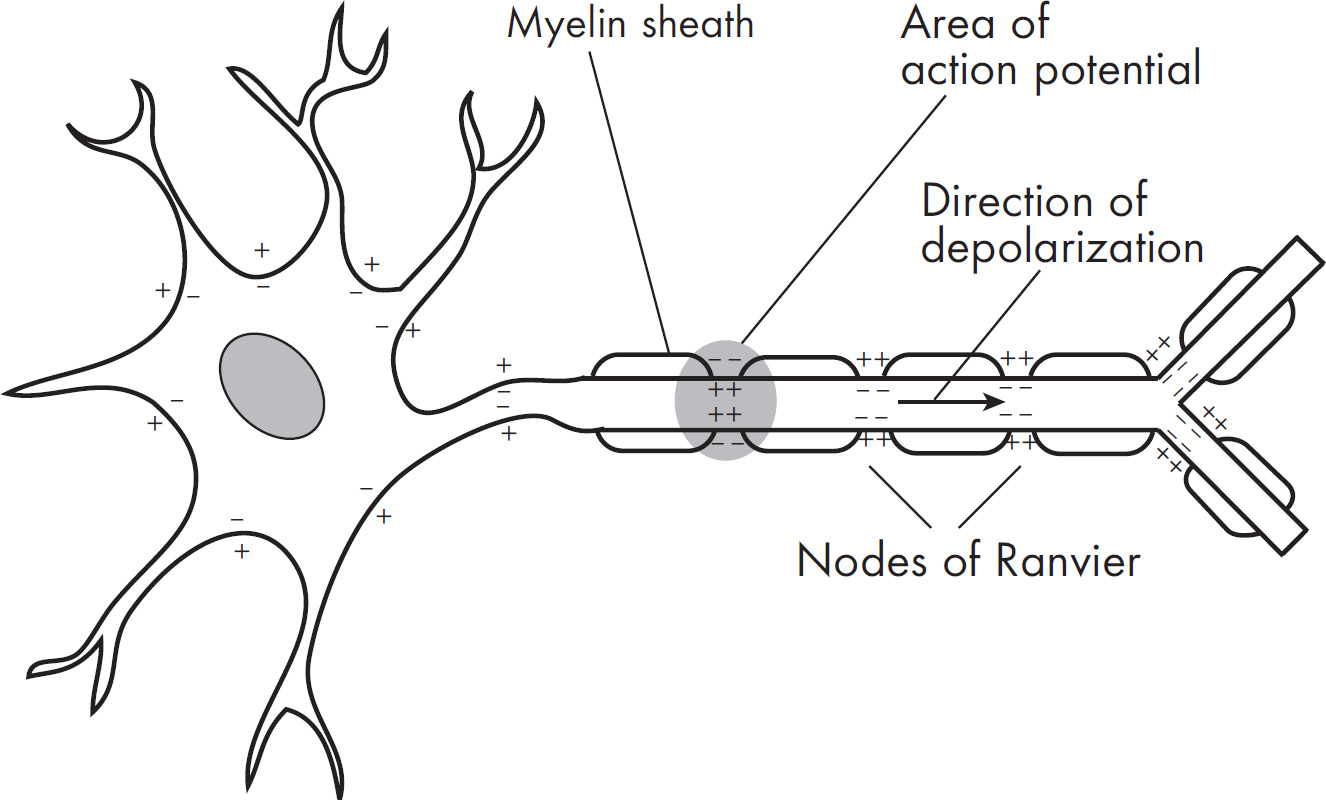
As you can see from the illustration above, the whole axon isn’t covered with myelin sheaths. The spaces between myelin sheaths—the exposed regions of the axon—are called the nodes of Ranvier. Myelin sheaths speed up the propagation of an impulse. Instead of the standard “domino effect” that occurs during an action potential, the impulse can now jump from node to node. It is almost as if there are giant dominoes which can skip sections of the neuron. This form of conduction is called saltatory conduction.
Thanks to the myelin sheath, the neuron can transmit an impulse down the axon far more rapidly than it could without its help.
Parts of the Nervous System
Central Nervous System
All of the neurons within the brain and spinal cord make up the central nervous system. All of the other neurons lying outside the brain and the spinal cord—in our skin, our organs, and our blood vessels—are collectively part of the peripheral nervous system. Although both of these systems are really part of one system, we still use the terms central and peripheral. The brain has many regions and parts that each have specific functions, but you do not need to know the details of the different sections.
So keep the following in mind:
• The central nervous system includes the neurons in the brain and spinal cord.
• The peripheral nervous system includes all the rest.
The Stimulus-Decision-Response Pathway
Neurons can be classified into three groups: sensory (affector) neurons, interneurons, and motor (effector) neurons. Sensory neurons receive impulses from the environment and bring them to the body. For example, sensory neurons in your hand are stimulated by touch. They transmit the message about the touch stimulus to the brain and the spinal cord. Once there, interneurons will make the decision about what to do about the stimulus. Then, motor neurons transmit the decision from the brain to muscles or glands to produce a response. The muscle will respond by contracting, or the gland will respond by secreting a substance (such as a hormone).
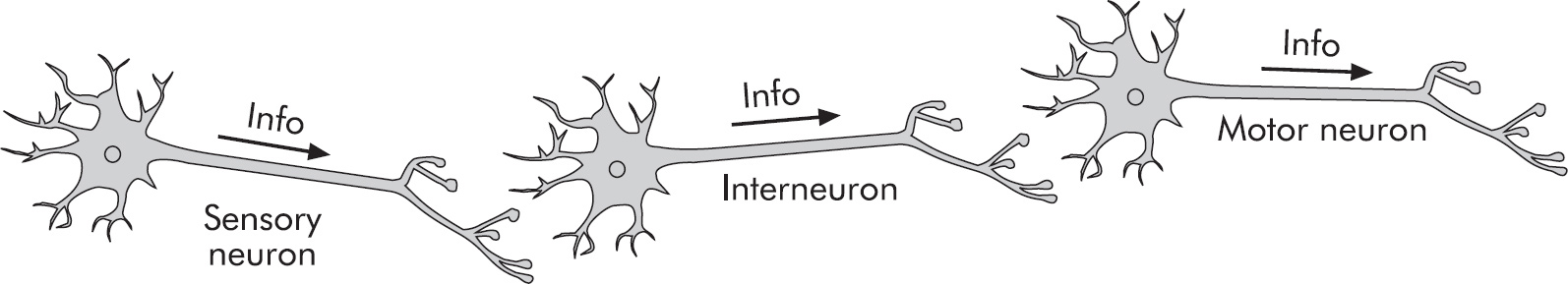
So, the basic function of the nervous system is to detect a stimulus and transmit it to the central nervous system, which makes a decision and then sends that decision out to the body to respond to the stimulus.
Reflexes
As mentioned, the central nervous system is composed of the brain and the spinal cord. Usually, the brain is the part that makes the decisions, so most stimuli are sent to the brain. However, in certain instances, the spinal cord is capable of making the decision. This is what happens in reflexes. A quick decision is required to prevent injury, and reflex arcs are pre-determined decisions that the spinal cord is set up to execute. The spinal cord may make a decision to withdraw a hand because of pain or to relax a muscle when its opposite contracts to prevent injury.
THE ENDOCRINE SYSTEM
Like the nervous system, the endocrine system is an excellent example of signaling. It is responsible for maintaining homeostasis and for coordinating responses to many stimuli in the body. Chemical messengers can be produced in one region of the body to act on target cells in another region. These chemicals, known as hormones, are produced in specialized organs called endocrine glands. Hormones have a number of functions including regulating growth, behavior, development, and reproduction. Other chemical messengers are used for communication. For example, pheromones help animals to communicate with members of their species and attract the opposite sex.
Endocrine glands release hormones directly into the bloodstream, which carries them throughout the body. Take a look at the endocrine glands in the human body.
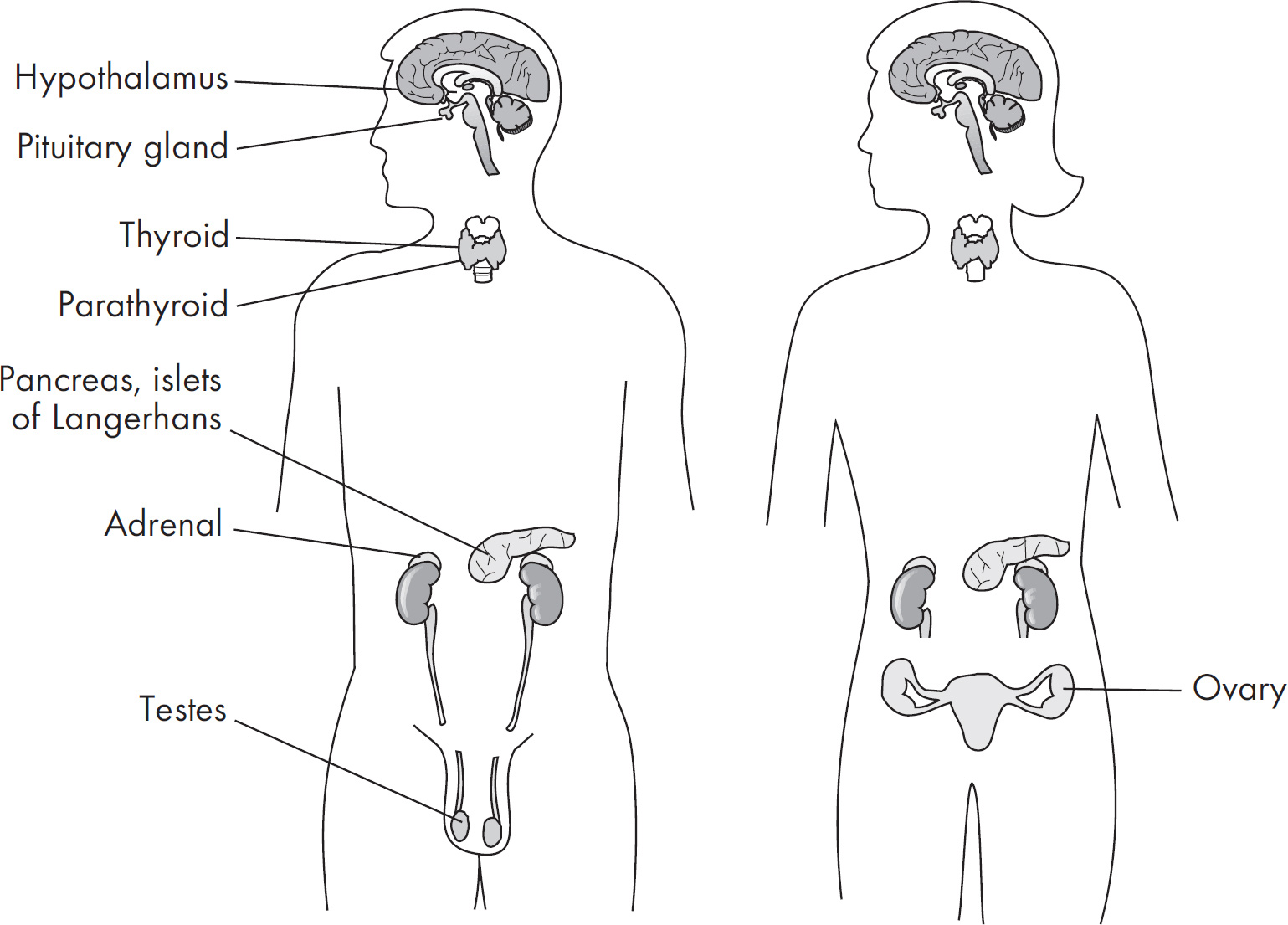
Although hormones flow in your blood, they affect only specific cells. The cells that a hormone affects are known as the target cells. Suppose, for example, that gland X makes hormone Y. Hormone Y, in turn, has some effect on organ Z. We would then say that organ Z is the target organ of hormone Y.
Hormones also operate by a negative feedback system. That is, an excess of the hormone will signal the endocrine gland to temporarily shut down production. For example, when hormone Y reaches a peak level in the bloodstream, the organ secreting the hormone, gland X, will get a signal to stop producing hormone Y. Once the levels of hormone Y decline, the gland can resume production of the hormone. Now we will go through a few examples of hormone regulation.
The Pancreas
The pancreas secretes two well-known hormones, glucagon and insulin, both of which are produced in clusters of cells called the islets of Langerhans. The target organs for these hormones are the liver and muscle cells. Glucagon, produced by α cells, stimulates the liver to convert glycogen into glucose and to release that glucose into the blood. Glucagon therefore increases the levels of glucose in the blood. Insulin has precisely the opposite effect that glucagon does.
When the blood has too much glucose floating around, insulin, produced by β cells, allows body cells to remove glucose from the blood. Consequently, insulin decreases the level of glucose in the blood. Insulin is particularly effective on muscle and liver cells. In short:
• insulin lowers the blood sugar level
• glucagon raises the blood sugar level
Sex Hormones
There are three key hormones involved in the reproductive system: estrogen, progesterone, and testosterone. Estrogen and progesterone are hormones released by the ovaries that regulate the menstrual cycle. Testosterone is the male hormone responsible for promoting spermatogenesis, the production of sperm. In addition, these hormones maintain secondary sex characteristics.
How Hormones Work
How do hormones trigger the activities of their target cells? That all depends on whether the hormone is a steroid (lipid soluble) or a protein, peptide, or amine (not lipid soluble). If the hormone is a steroid, then the hormone can diffuse across the membrane of the target cell. It then binds to a receptor protein in the nucleus and regulates transcription of the DNA, which in turn effects production of proteins.
However, if the hormone is a protein, peptide, or amine, it can’t get into the target cell by means of simple diffusion. Remember: only hydrophobic things can pass through the membrane. The hormone binds to a receptor protein on the cell membrane of the target cell. This often causes a signal cascade whereby the signal is transduced into the cell without the actual hormone itself ever entering the cell.
Here’s a summary of the hormones and their effects on the body.
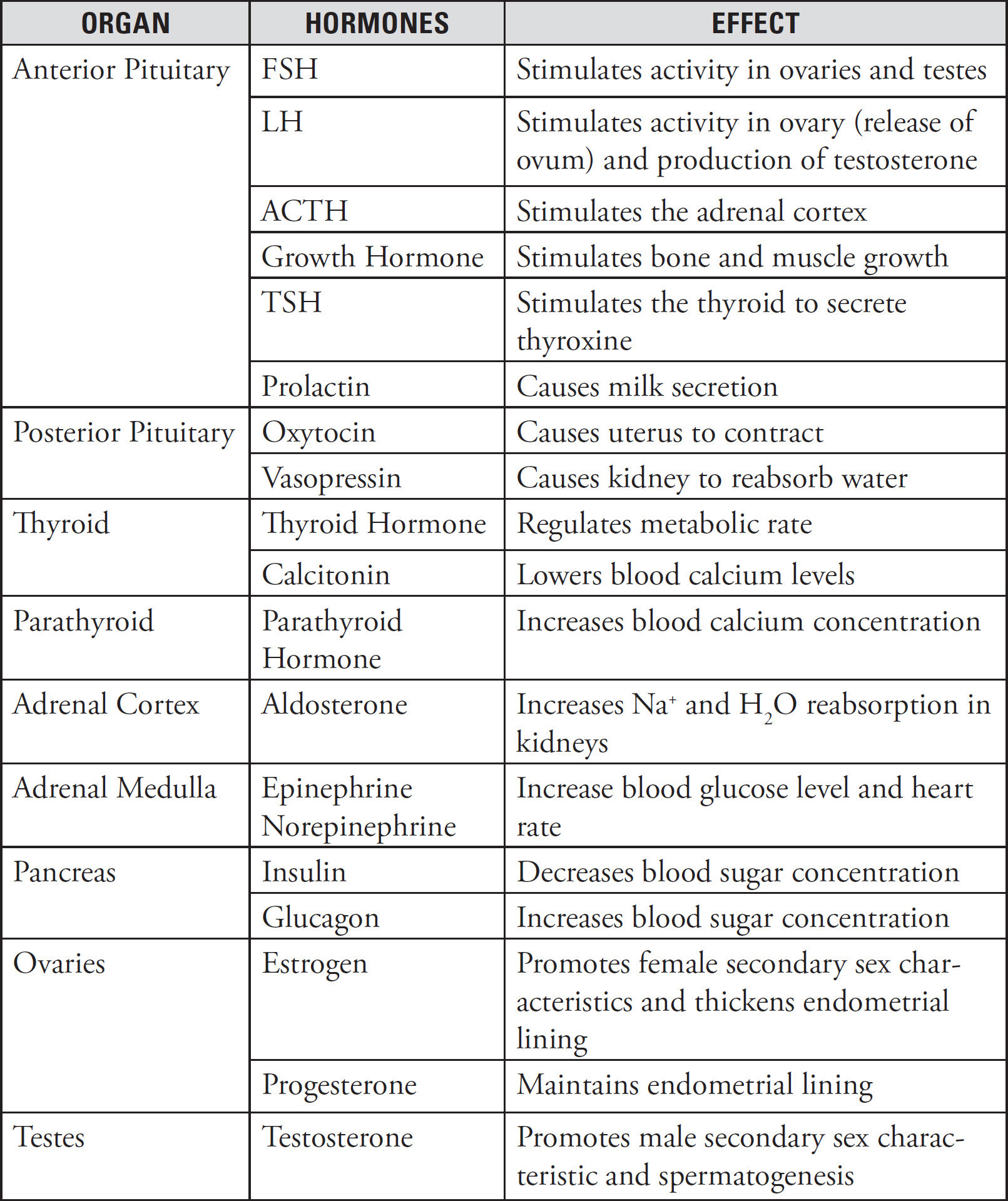
While the nervous system and the endocrine system work in close coordination, there are significant differences between the two:
• The nervous system sends nerve impulses using neurons, whereas the endocrine system secretes hormones.
• Nerve impulses control rapidly changing activities, such as muscle contractions, whereas hormones deal with long-term adjustments.
KEY TERMS
homeostasis
negative feedback pathway (feedback inhibition)
positive feedback pathway
embryonic development
morphogenesis
zygote
fertilization
homeotic genes/Hox genes
tissue
organ
body systems
Immune System
pathogen
bacteria
conjugation
viruses
host
bacteriophage
lytic cycle
lysogenic cycle
enveloped virus
retrovirus
reverse transcriptase
antigen
innate immune system
adaptive and specific immune response
phagocytes
macrophages
complement proteins
interferons
inflammatory response
lymphocytes
B-lymphocytes
humoral response
memory B-cell
plasma cell
antibody
T-lymphocytes
cell-mediated immunity
major histocompatibility complex (MHC) markers
cytotoxic T-cell
apoptosis
helper T-cell
lymph node
AIDS
Nervous System
nerve net
ganglia
neurons
cell body
dendrites
axon
action potential
resting membrane potential
Na+/K+ ATPase pump
polarized
threshold
all-or-none response
depolarization
refractory period
axon bulb
neurotransmitter
synapse
acetylcholine
aceytlcholinesterase
norepinephrine
GABA
Schwann cells
myelin sheath
nodes of Ranvier
saltatory conduction
central nervous system
peripheral nervous system
sensory (effector) neurons
interneurons
motor (affector) neurons
reflexes
Endocrine System
hormones
endocrine glands
pheromones
target cells
glucagon (alpha cell)
insulin
islets of Langerhans
glycogen
estrogen
progesterone
testosterone
Summary
 The AP Biology Exam requires specific knowledge of only three systems: immune system, nervous system, and endocrine system. Other body systems might appear on the test, but only in the context of the kinds of themes covered in this chapter: structure and function relationships, communication between cells, and so on. Understanding the following key points about the three systems highlighted in this chapter will help you with the thematic questions about various body systems on the test.
The AP Biology Exam requires specific knowledge of only three systems: immune system, nervous system, and endocrine system. Other body systems might appear on the test, but only in the context of the kinds of themes covered in this chapter: structure and function relationships, communication between cells, and so on. Understanding the following key points about the three systems highlighted in this chapter will help you with the thematic questions about various body systems on the test.
Immune System
 The immune system is the defense system against pathogens that enter the organism. Bacteria and viruses are common pathogens. Bacteriophages are viruses that infect bacteria. Viruses require a host to replicate and sometimes lyse the host cell during infection. There are two sides to the immune system:
The immune system is the defense system against pathogens that enter the organism. Bacteria and viruses are common pathogens. Bacteriophages are viruses that infect bacteria. Viruses require a host to replicate and sometimes lyse the host cell during infection. There are two sides to the immune system:
• The innate immune system is the first response, the initial protection, and it includes the skin and mucous linings. Phagocytes and complement proteins also target this initial response to neutralize invaders.
• Acquired immunity can be cell mediated or antibody based. T-lymphocytes include cytotoxic killer T-cells and helper T-cells. B-cells include plasma and memory B-cells.
Nervous System
 Neurons are highly specialized, terminally differentiated cells. They are composed of dendrites, cell bodies, axons, and synaptic terminals:
Neurons are highly specialized, terminally differentiated cells. They are composed of dendrites, cell bodies, axons, and synaptic terminals:
• Neurons are classified into three groups: sensory neurons, motor neurons, and interneurons.
• Neuronal transmission is based on action potentials, which are temporary disruptions of the ion concentrations set up by the resting membrane potential.
• The junction between neurons is called a synapse. The presynaptic axon releases neurotransmitters to the postsynaptic dendrite to perpetuate the transmission.
• Myelin speeds the transmission along axons.
 The central nervous system is composed of the brain and spinal cord.
The central nervous system is composed of the brain and spinal cord.
 The peripheral nervous system senses and responds to stimuli.
The peripheral nervous system senses and responds to stimuli.
Endocrine System
 Endocrine glands are tissues or organs that excrete hormones. Hormones mediate growth, reproduction, waste disposal, nutrient absorption, and behavior.
Endocrine glands are tissues or organs that excrete hormones. Hormones mediate growth, reproduction, waste disposal, nutrient absorption, and behavior.
 Target cells receive the specific hormone via receptors either on the surface of the cell or internally and react through signal transduction and response.
Target cells receive the specific hormone via receptors either on the surface of the cell or internally and react through signal transduction and response.
Chapter 11 Drill
Answers and explanations can be found in Chapter 16.
1. Lymph nodes are often evaluated when a patient is believed to have an active infection or cancer. Why?
(A) Active infections or cancer often generate large quantities of fluids as cells rupture; the lymph nodes often swell as a result of the buildup of these fluids.
(B) During an active infection, lymphocytes expand rapidly in lymph nodes, causing swelling of the structures.
(C) Lymph nodes are the most common tissue site of active infections or cancer.
(D) Cancer often metastasizes to lymph nodes, causing them to expand as new cancerous cells multiply.
Questions 2-4 refer to the following chart and paragraph.
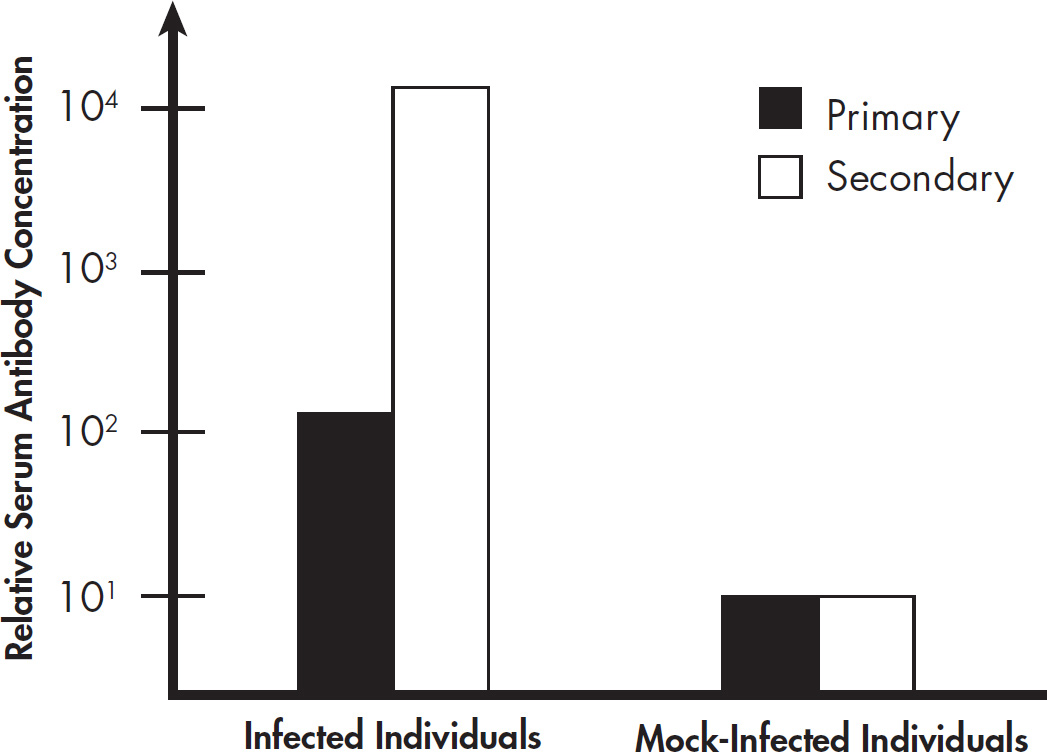
A scientist wishes to test the efficacy and immunogenicity of a newly developed, live-attenuated virus vaccine strain. To test for immunogenicity, the scientist acquires 100 healthy volunteers and exposes them to either the vaccine strain or a placebo strain (consisting of isotonic saline solution). Twelve days following the initial primary exposure, the serum antibody concentration is determined by an ELISA analysis. A follow-up exposure to test for existing immunity is performed on day 60 using the same 100 individuals. Serum antibody titers are determined for the secondary challenge on day 72. These data are shown in the chart above.
2. The placebo strain was included in the experimental design in order to
(A) evaluate the immune response to isotonic saline solution
(B) determine primary and secondary immune responses to the virus
(C) compare antibody titers of infected and mock-infected individuals as a control
(D) practice before challenging the same individuals with the vaccine strain
3. What type of immune cells are responsible for generating the antibodies generated in this experiment?
(A) Erythrocytes
(B) B-cells
(C) T-cells
(D) Phagocytes
4. How can the scientist increase the statistical significance of these data?
(A) She could repeat the experiment with a greater sample size.
(B) She could use more control variables.
(C) She could perform a third challenge on the same 100 individuals.
(D) She could use a different statistical test, which provides the necessary statistical significance.
5. Myasthenia gravis is an autoimmune disorder in which patients experience fatigue, sluggishness, and weakness. Rogue autoantibodies in the bloodstream target the body’s own acetylcholine receptors at the neuromuscular junctions, causing the patients’ muscles to exhibit delayed responses. Which treatment would have the most immediate positive effect for patients of this disorder?
(A) Broad-spectrum immunosuppression drugs to reduce the effectiveness of the immune system
(B) Removal of the thymus because it is responsible for maturation of macrophages
(C) Blood transfusion to remove or dilute the acetylcholine autoantibodies
(D) Additional acetylcholine
6. Innate immune system defenses in animals include skin, mucous membranes, and defensive enzymes present in saliva, tears, stomach acid, and skin. Analogous structures in plants could include
(A) long stems
(B) thorns and spines
(C) attractive scents
(D) stomata
7. When lymphocytes first encounter a pathogen, receptors on their surface first bind to the outside of the pathogen. Next, this activated lymphocyte multiplies and becomes numerous, allowing them to bind to many more pathogens of the same type in the bloodstream. What are other results from this initial recognition of the pathogen?
(A) Antibodies specific to the pathogen are secreted by B-cells.
(B) The pathogen is destroyed by helper T-cells.
(C) Only plasma B-cells remain.
(D) The pathogen is permitted to infect more cells prior to its destruction.
8. After a macrophage engulfs a pathogen through phagocytosis, it destroys the pathogen, and then the macrophage matures to become
(A) T-cells that secrete specific antibodies for the pathogen
(B) MHC antigen-presenting cells
(C) activated plasma cells
(D) memory B-cells
9. The image below depicts an action potential occurring in an active neuron. Which of the following best explains why the membrane potential depolarizes at –50 mV?
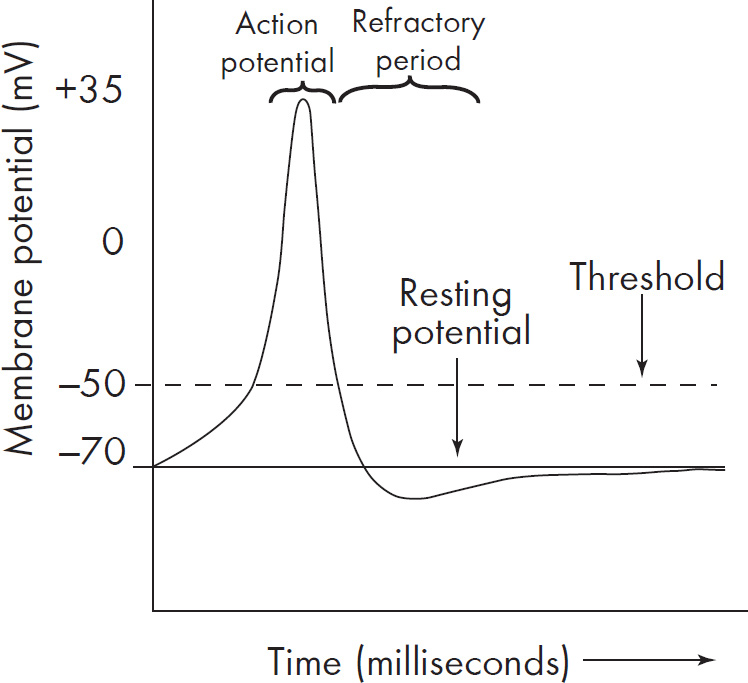
(A) At –50 mV, voltage-gated potassium channels open, allowing potassium ions (K+) to rush into the cell.
(B) At –50 mV, voltage-gated sodium channels open, allowing sodium ions (Na+) to rush into the cell.
(C) At –50 mV, voltage-gated potassium channels open, allowing potassium ions (K+) to rush out of the cell.
(D) At –50 mV, voltage-gated sodium channels open, allowing sodium ions (Na+) to rush out of the cell.
10. Fugu is a Japanese delicacy made from the preparation of pufferfish. Pufferfish produce a powerful toxin called tetrodotoxin, which binds to and blocks voltage-gated sodium channels. Specially trained Japanese chefs must carefully remove the toxin during preparation of fugu to prevent inadvertent poisoning by ingestion. Which of the following would likely occur to a neuron exposed to tetrodotoxin?
(A) The neuron would be able to send action potentials but would not be able to release neurotransmitters to neighboring neurons.
(B) The neuron would be able to send action potentials but would not be able to receive neurotransmitters from a presynaptic neuron.
(C) The neuron would not be able to send action potentials and would be unable to continue a neural response.
(D) The neuron would not be able to send action potentials but will still be depolarized by the influx of sodium ions.
11. Ouabain is historically used by Somali tribes on poisoned arrows. Ouabain inhibits the activity of the sodium-potassium pump and is therefore used as a powerful neurotoxin. What immediate effect would ouabain have on motor and sensory neurons in a victim of a poisoned arrow?
(A) The muscles would spasm because of an excess amount of calcium released from the sarcoplasmic reticulum.
(B) Action potentials would not be capable of firing because the pumps are blocked and the cell would be very positively-charged.
(C) Brain damage would result as the poison spread up the neurons to the central nervous system.
(D) Action potentials would fire in the affected neurons, and then repolarization would be impossible without functioning sodium-potassium pumps. Paralysis would result.
12. In a neuron, the majority of the protein manufacturing machinery will be found in the
(A) dendrites
(B) cell body
(C) axon
(D) axon terminal
13. Acetylcholinesterase is an enzyme hydrolase that destroys the neurotransmitter acetylcholine in the neuromuscular junction. Patients with defective acetylcholinesterase will have
(A) more easily relaxed musculature
(B) extreme weakness
(C) overactive and hyper-contraction of muscles
(D) no effect
14. In an action potential, the “threshold” is reached when
(A) enough voltage-gated channels open to produce a change in membrane polarity
(B) the synaptic vesicles release neurotransmitters into the synaptic cleft
(C) the peak depolarization occurs
(D) sodium ions enter the cell
15. Oligodendricytes exist in the central nervous system, acting analogously to the Schwann cells of the peripheral nervous system. Therefore, their main objective is to
(A) act as immune cells of the brain
(B) produce a myelin sheath that insulates the axonal transmission
(C) direct depolarization of the membrane and perpetuate the nerve impulse
(D) transmit nerve impulses
16. Testosterone and estradiol are hormones that are derivatives of steroid molecules. Therefore, these hormones reach the interior of many different types of cells easily. Which of the following accurately describes this type of hormone reception?
(A) Many types of cells have receptors for testosterone and estradiol on their plasma membrane.
(B) These hormones are lipid-soluble, so they reach the interior of cells by crossing the plasma membrane.
(C) Transport proteins on the surface of cells shuttle these hormones into the cell.
(D) Hormones never enter cells, so this question must be inaccurate.
17. Only certain cells in the body are target cells for the hormone vasopressin. Which of the following best explains the specificity of this hormone?
(A) Target cells are the only cells exposed to vasopressin.
(B) Target cells are the only cells with receptors for vasopressin.
(C) Vasopressin enters all cells because it is nonpolar.
(D) Cells without vasopressin receptors destroy vasopressin before it can have an effect.
REFLECT
Respond to the following questions:
• Which topics from this chapter do you feel you have mastered?
• Which content topics from this chapter do you feel you need to study more before you can answer multiple-choice questions correctly?
• Which content topics from this chapter do you feel you need to study more before you can effectively compose a free response?
• Was there any content that you need to ask your teacher or another person about?
Chapter 12
Behavior and Ecology
In the preceding chapters, we looked at individual organisms and the ways they solve life’s many problems: acquiring nutrition, reproducing, and so on. Now let’s turn to how organisms deal with their environments. We can divide the discussion of organisms and their environments into two general categories: behavior and ecology.
BEHAVIOR
Some animals behave in a programmed way to specific stimuli, while others behave according to some type of learning. We humans can do both. For the AP Biology Exam, you’ll have to know a little about the different types of behavior.
All behavior means, basically, is how organisms cope with their environments. This chapter looks at the general types of "Ahead1" id="calibre_link-108">INSTINCT
Instinct is an inborn, unlearned behavior. Sometimes the instinctive behavior is triggered by environmental signals called releasers. The releaser is usually a small part of the environment that is perceived. For example, when a male European robin sees another male robin, the sight of a tuft of red feathers on the male is a releaser that triggers fighting behavior. In fact, because instinct underlies all other behavior, it can be thought of as the circuitry that guides behavior.
For example, hive insects, such as bees and termites, never learn their roles; they are born knowing them. On the basis of this inborn knowledge, or instinct, they carry out all their other behaviors: a worker carries out “worker tasks,” a drone “drone tasks,” and a queen “queen tasks.” Another example is the “dance” of the honey bee, which is used to communicate the location of food to other members of the beehive.
There are other types of instinct that last for only a part of an animal’s life and are gradually replaced by “learned” behavior. For example, human infants are born with an ability to suck from a nipple. If it were not for this instinctual behavior, the infant would starve. Ultimately, however, the infant will move beyond this instinct and learn to feed itself. What exactly, then, is instinct?
For our purposes:
Instinct is the inherited “circuitry” that directs and guides behavior.
A particular type of innate behavior is a fixed action pattern. These behaviors are not simple reflexes, and yet they are not conscious decisions. An example is the egg-rolling behavior exhibited by a graylag goose. If the egg is removed from the goose, it will continue to make the same movements. That is, the innate movements are independent of the environment.
Learning
Another form of behavior is learning. Learning refers to a change in a behavior brought about by an experience (which is what you’re doing this very moment). Animals learn in a number of ways. For the AP Biology Exam, we’ll take a look at the key types of learning.
Imprinting
Have you ever seen a group of goslings waddling along after their mother? How is it that they recognize her? Well, the mother arrives and gives out a call that the goslings recognize. The goslings, hearing the call, know that this is their mother and follow her around until they are big enough to head out on their own.
Now imagine the same goslings, newly hatched. If the mother is absent, they will accept the first moving object they see as their mother. This process is known as imprinting.
Animals undergo imprinting within a few days after birth in order to recognize members of their own species. While there are different types of imprinting, including parent, sexual, and song imprinting, they all occur during a critical period—a window of time when the animal is sensitive to certain aspects of the environment.
Remember:
Imprinting is a form of learning that occurs during a brief period of time, usually early in an organism’s life.
Classical Conditioning
If you have a dog or a cat, you know that every time you hit the electric can opener, your cat or dog comes running. This is a form of classical conditioning. To feed your pet, you need to open its can of food. For your pet, the sound of the opener has come to be associated with eating: every time it hears the opener, it thinks that it is about to be fed. We can say that your pet has been “conditioned” to link the buzz of the can opener with its evening meal.
This type of learning is now known as associative learning.
Operant Conditioning
Another type of associative learning is operant conditioning (or trial-and-error learning).
In operant conditioning, an animal learns to perform an act in order to receive a reward.
If the behavior is not reinforced, the conditioned response will be lost. This is called extinction. Here’s one thing to remember:
In operant conditioning, the animal’s behavior determines whether it gets the reward or the punishment. Classical conditioning is just the learned association of two things.
Habituation is another form of learning. It occurs when an animal learns not to respond to a stimulus. For example, if an animal encounters a stimulus over and over again without any consequences, the response to it will gradually lessen and may altogether disappear.
To recap, there are four basic types of learning:
• Imprinting occurs early in life and helps organisms recognize members of their own species.
• Classical conditioning involves learning through association.
• Operant conditioning occurs when a response is associated with new stimuli (also a form of associative learning).
Internal Clocks: The Circadian Rhythm
There are other instinctual behaviors that occur in both animals and plants. One such behavior deals with time. Have you ever wondered how roosters always know when to start crowing? The first thought that comes to mind is that they’ve caught a glimpse of the sun. Yet many crow even before the sun has risen.
Roosters do have internal alarm clocks. Plants have them as well. These internal clocks, or cycles, are known as circadian rhythms.
Watch out though: seasonal changes, like the loss of leaves by deciduous trees or the hibernation of mammals, are not examples of circadian rhythms. Circadian refers only to daily rhythms. Need a mnemonic? Just think how bad your jet lag would be after a trip around the world. In other words: Circling the globe screws up your circadian rhythm.
The Science Behind Your Jet Lag
If you’ve ever flown overseas, you know all about circadian rhythms. They’re the cause of jet lag. Our bodies tell us it’s one time, while our watches tell us it’s another. The sun may be up, but our body’s internal clock is crying, “Sleep!” This sense of time is purely instinctual: you don’t need to know how to tell time in order to feel jet lag.
HOW ANIMALS COMMUNICATE
Some animals use signals as a way of communicating with members of their species. These signals, which can be chemical, visual, electrical or tactile, are often used to influence mating and social behavior.
Chemical signals are one of the most common forms of communication among animals. Pheromones, for instance, are chemical signals between members of the same species that stimulate olfactory receptors and ultimately affect behavior. For example, when female insects give off their pheromones, they attract males from great distances.
Visual signals also play an important role in the behavior observed among members of a species. For example, fireflies produce pulsed flashes that can be seen by other fireflies far away. The flashes are sexual displays that help male and female fireflies identify and locate each other in the dark.
Other animals use electrical channels to communicate. For example, some fish generate and receive weak electrical fields. Finally, tactile signals are found in animals that have mechanoreceptors in their skin to detect prey. For instance, cave-dwelling fishes use mechanoreceptors in their skin for communicating with other members and detecting prey.
Social Behavior
Many animals are highly social species, and they interact with each other in complex ways. Social behaviors can help members of the species survive and reproduce more successfully. Several behavioral patterns for animal societies are summarized below:
• Agonistic behavior is aggressive behavior that occurs as a result of competition for food or other resources. Animals will show aggression toward other members that tend to use the same resources. A typical form of aggression is fighting between competitors.
• Dominance hierarchies (or pecking orders) occur when members in a group have established which members are the most dominant. The more dominant male will often become the leader of the group and will usually have the best pickings of the food and females in the group. Once the dominance hierarchy is established, competition and tension within the group is reduced.
• Territoriality is a common behavior when food and nesting sites are in short supply. Usually, the male of the species will establish and defend his territory (called a home range) within a group in order to protect important resources. This behavior is typically found among birds.
• Altruistic behavior is defined as unselfish behavior that benefits another organism in the group at the individual’s expense because it advances the genes of the group. For example, when ground squirrels give warning calls to alert other squirrels of the presence of a predator, the calling squirrel puts itself at risk of being found by the predator.
Symbiotic Relationships
Many organisms that coexist exhibit some type of symbiotic relationship. These include remoras, or “sucker fish,” which attach themselves to the backs of sharks, and lichen—the fuzzy, mold-like stuff that grows on rocks. Lichen appears to be one organism, when in fact it is two organisms—a fungus and an alga, or photosynthetic bacterium—living in a complex symbiotic relationship.
Overall, there are three basic types of symbiotic relationships:
• Mutualism—in which both organisms win (for example, the lichen)
• Commensalism—in which one organism lives off another with no harm to the host organism (for example, the remora)
• Parasitism—in which the organism actually harms its host
There are many special types of relationships between multicellular organisms and unicellular organisms. One important example is bacteria living inside the guts of many mammals. We have a mutualistic relationship. Humans and other mammals provide them a nice home with nutrients, and they help us in many ways by breaking things down and helping us make things. It is also good that we are filled with “good” bacteria because otherwise we might be susceptible to dangerous pathogenic bacteria.
PLANT BEHAVIOR
Plants have also evolved specific ways to respond to their environment. The plant behaviors covered on the AP Biology Exam are photoperiodism and tropisms. Plants flower in response to changes in the amount of daylight and darkness they receive. This is called photoperiodism. Although you’d think that plants bloom based on the amount of sunlight they receive, they actually flower according to the amount of uninterrupted darkness.
Tropisms
Plants need light. Notice that all the plants in your house tip toward the windows. This movement of plants toward the light is known as phototropism. As you know, plants generally grow up and down: the branches grow upward, while the roots grow downward into the soil, seeking water. This tendency to grow toward or away from the earth is called gravitropism. All of these tropisms are examples of behavior in plants.
A tropism is a turning in response to a stimulus.
There are three basic tropisms in plants. They’re easy to remember because their prefixes the stimuli to which plants react:
• Phototropism refers to how plants respond to sunlight. For example, bending toward light.
• Gravitropism refers to how plants respond to gravity. Stems exhibit negative gravitropism (they grow up, away from the pull of gravity), whereas roots exhibit positive gravitropism (they grow downward into the earth).
• Thigmotropism refers to how plants respond to touch. For example, ivy grows around a post or trellis.
ECOLOGY
The study of the interactions between living things and their environments is known as ecology. We’ve spent most of our time discussing individual organisms. However, in the real world, organisms are in constant interaction with other organisms and the environment. The best way for us to understand the various levels of ecology is to progress from the big picture, the biosphere, down to the smallest ecological unit, the population.
There is a hierarchy within the world of ecology. Each of the following terms represents a different level of ecological interaction:
• Biosphere: The entire part of the earth where living things exist. This includes soil, water, light, and air. In comparison to the overall mass of the earth, the biosphere is relatively small. If you think of the earth as a basketball, the biosphere is equivalent to a coat of paint over its surface
• Ecosystem: The interaction of living and nonliving things
• Community: A group of populations interacting in the same area
• Population: A group of individuals that belong to the same species and that are interbreeding
Biosphere
The biosphere can be divided into large regions called biomes. Biomes are massive areas that are classified mostly on the basis of their climates and plant life. Because of the different climates and terrains on the earth, the distribution of living organisms varies.
Remember that biomes tend to be arranged along particular latitudes. For instance, if you hiked from Alaska to Kansas, you would pass through the following biomes: tundra, taiga, temperate deciduous forests, and grasslands.
Ecosystem
Ecosystems are self-contained regions that include both living and nonliving factors. Living factors are called biotic factors and nonliving factors are called abiotic factors. Abiotic factors include water, humidity, temperature, soil/atmosphere composition, light, and radiation. For example, a lake, its surrounding forest, the atmosphere above it, and all the organisms that live in or feed off of the lake would be considered an ecosystem. As you probably know, there is an exchange of materials between the components of an ecosystem.
Take a look at the flow of carbon through a typical ecosystem.
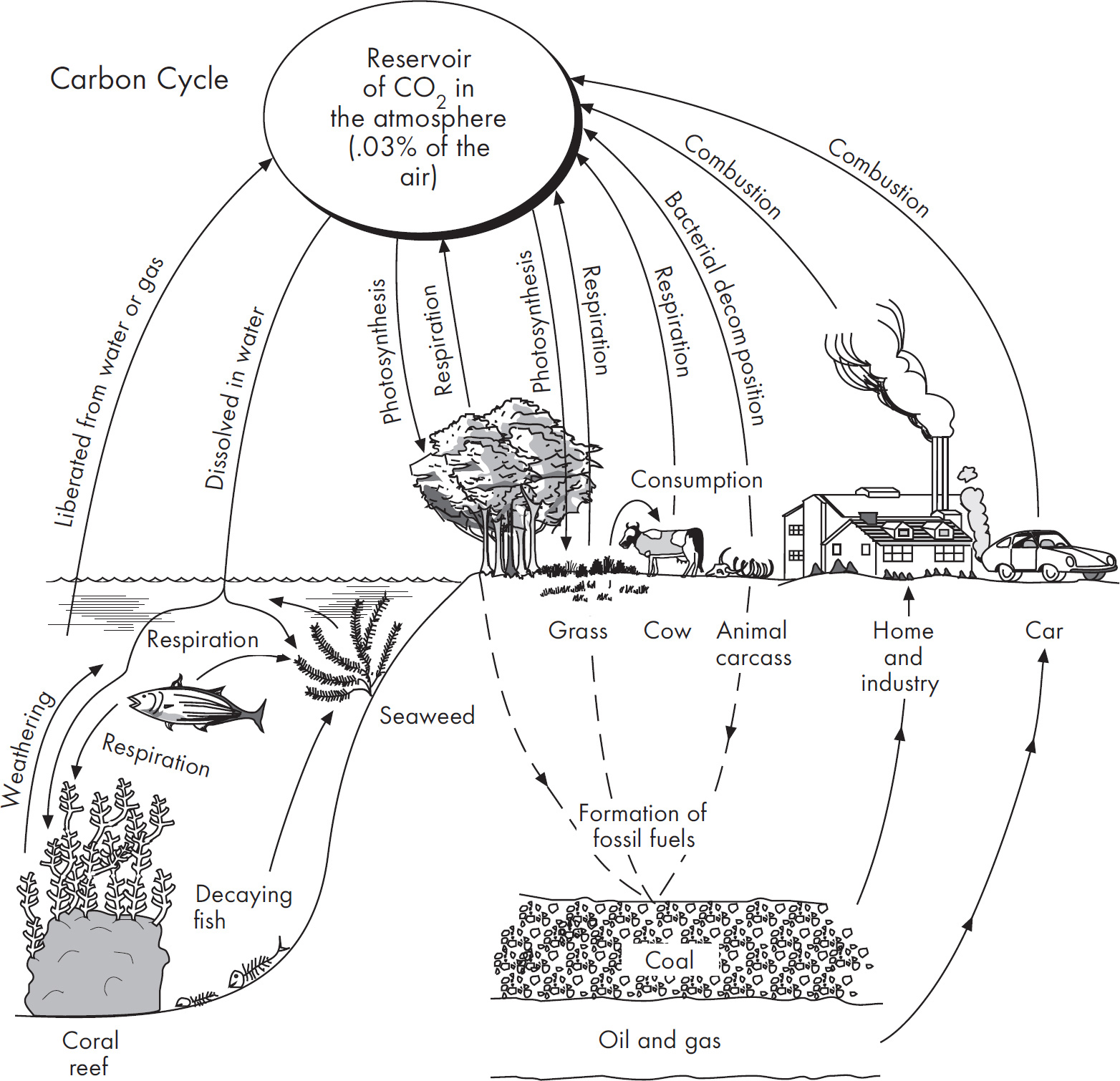
You’ll notice how carbon is recycled throughout the ecosystem—this is called the carbon cycle. In other words, carbon flows through ecosystems. Other substances have similar cycles. The interactions between the biotic and the abiotic factors are crucial for the balance of the ecosystem.
Community
The next smaller level is the community. A community refers to a group of interacting plants and animals that show some degree of interdependence. For instance, you, your dog, and the fleas on your dog are all members of the same community.
Each organism has its own niche—its position or function in a community. Because every species occupies a niche, it’s going to have an impact on all the other organisms. When two organisms occupy the same niche, they will compete for the resources within that niche. If a species can occupy an unoccupied niche, it will usually thrive without competition. These connections are shown in the food chain. A food chain describes the way different organisms depend on one another for food. There are basically four levels to the food chain: producers, primary consumers, secondary consumers, and tertiary consumers.
Producers (autotrophs)
Producers, or autotrophs, have all of the raw building blocks to make their own food. From water and the gases that abound in the atmosphere and with the aid of the sun’s energy, autotrophs convert light energy to chemical energy. They accomplish this through photosynthesis.
Primary productivity is the rate at which autotrophs convert light energy into chemical energy in an ecosystem. There are two types of primary productivity:
1. The gross productivity just from photosynthesis cannot be measured because cell respiration is occurring at the same time.
2. Net productivity only measures organic materials that are left over after photosynthetic organisms have taken care of their own cellular energy needs. This is calculated by measuring oxygen production in the light, when photosynthesis and cell respiration are both occurring. This equation is listed on the Biology Equations and Formulas sheet.
Autotrophs produce all of the available food. They make up the first trophic (feeding) level. They possess the highest biomass (the total weight of all the organisms in an area) and the greatest numbers. Did you know that plants make up about 99 percent of the earth’s total biomass?
Consumers (heterotrophs)
Consumers, or heterotrophs, are forced to find their energy sources in the outside world. Basically, heterotrophs digest the carbohydrates of their prey into carbon, hydrogen, and oxygen and use these molecules to make organic substances.
The bottom line is that heterotrophs, or consumers, get their energy from the things they consume.
Primary consumers are organisms that directly feed on producers. A good example is a cow. These organisms are also known as herbivores. They make up the second trophic level.
The next level consists of organisms that feed on primary consumers. They are the secondary consumers, and they make up the third trophic level. Above these are tertiary consumers.
Decomposers
All organisms at some point must finally yield to decomposers. Decomposers are the organisms that break down organic matter into simple products. Generally, fungi and bacteria are the decomposers. They serve as the “garbage collectors” in our environment.
• Producers make their own food.
• Primary consumers (herbivores) eat producers.
• Secondary consumers (carnivores and omnivores) eat producers and primary consumers.
• Tertiary consumers eat all of the above.
• Decomposers break things down.
Keystone Species
Sometimes one organism is particularly important to an ecosystem. Maybe they are the only producer or maybe they are the only thing that keeps a particularly deadly predator in check. Important species like this are called keystone species. If a keystone species is removed from the ecosystem, the whole balance can be undone very quickly.
The 10% Rule
In a food chain, only about 10 percent of the energy is transferred from one level to the next—this is called the 10% rule. The other 90 percent is used for things like respiration, digestion, running away from predators—in other words, it’s used to power the organism doing the eating! As a level, the producers have the most energy in an ecosystem; the primary consumers have less energy than producers; the secondary consumers have less energy than the primary consumers; and the tertiary consumers will have the least energy of all.
The energy flow, biomass, and numbers of members within an ecosystem can be represented in an ecological pyramid. Organisms that are higher up on the pyramid have less biomass and energy, and they are fewer in number.
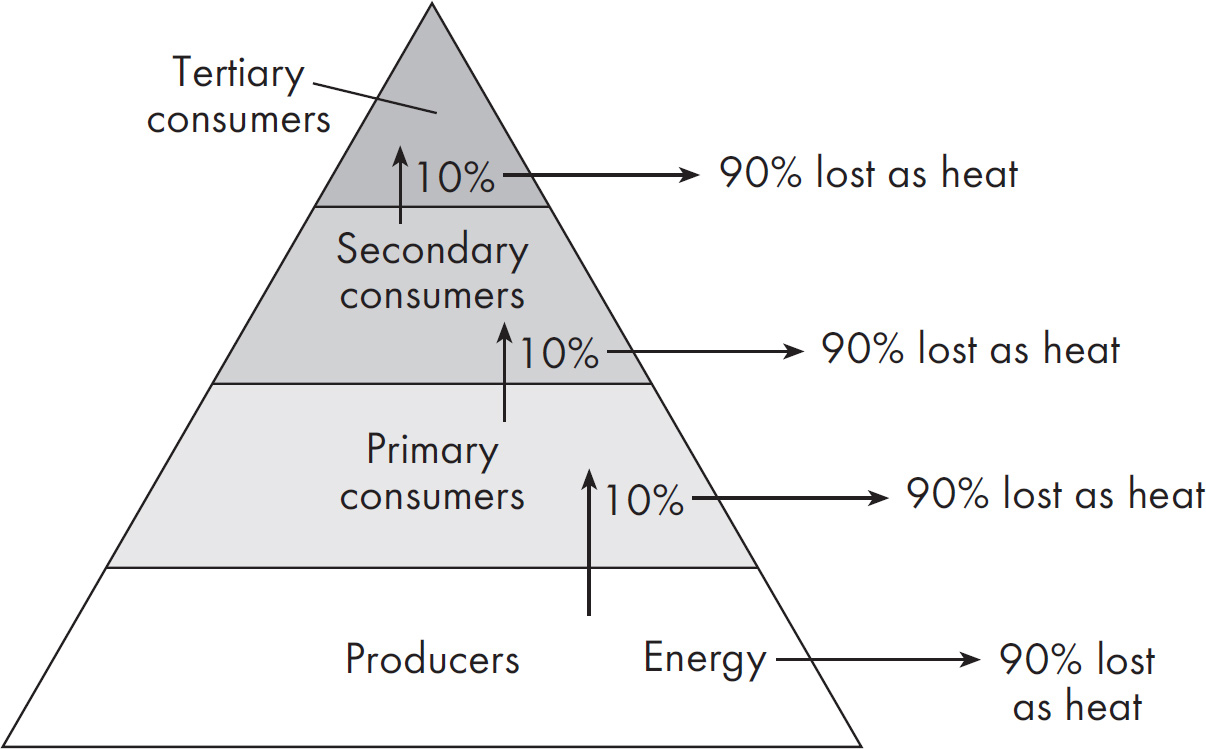
Build-up of Toxins
The downside of food pyramids is that when the consumers at the top eat something beneath them it is like they are eating that thing AND the thing that thing ate AND and the thing that thing ate, etc. So, if there is a toxin in the environment, the consumers at the top are getting the most of it because it becomes more concentrated at each level.
Toxins in an ecosystem are more concentrated and thus more dangerous for animals further up the pyramid.
This simply means that if a toxin is introduced into an ecosystem, the animals most likely to be affected are those at the top of the pyramid. This occurs because of the increasing concentration of such toxins. The classic example of this phenomenon is DDT, an insecticide initially used to kill mosquitoes. Although the large-scale spraying of DDT resulted in a decrease in the mosquito population, it also wound up killing off ospreys.
Ospreys are aquatic birds whose diet consists primarily of fish. The fish that ospreys consumed had, in turn, been feeding on contaminated insects (bioaccumulation). Because fish eat thousands of insects, and ospreys hundreds of fish, the toxins grew increasingly concentrated (biomagnification). Though the insecticide seemed harmless enough, it resulted in the near-extinction of certain osprey populations. What no one knew was that in sufficient concentrations, DDT weakened the eggshells of ospreys. Consequently, eggs broke before they could hatch, killing the unborn ospreys.
Such environmental tragedies still occur. All ecosystems, small or great, are intricately woven, and any change in one level invariably results in major changes at all the other levels.
Disrupting the Balance
Ecosystems are never exactly stable, existing in a precarious balance. Many things can disrupt the balance. If the biomass at any level changes, this will affect the surrounding levels and spread throughout the ecosystem. Geological changes, weather changes, new species, diseases, lack of resources, new resources, and many other things can disrupt the balance.
Population Ecology
Population ecology is the study of how populations change. Whether these changes are long-term or short-term, predictable or unpredictable, we’re talking about the growth and distribution patterns of a population.
When studying a population, you need to examine four things: the size (the total number of individuals), the density (the number of individuals per area), the distribution patterns (how individuals in a population are spread out), and the age structure.
For example, the graph below shows there is a high death rate among the young of oysters, but those that survive do well. On the other hand, there is a low death rate among the young of humans, but, after age sixty, the death rate is high.
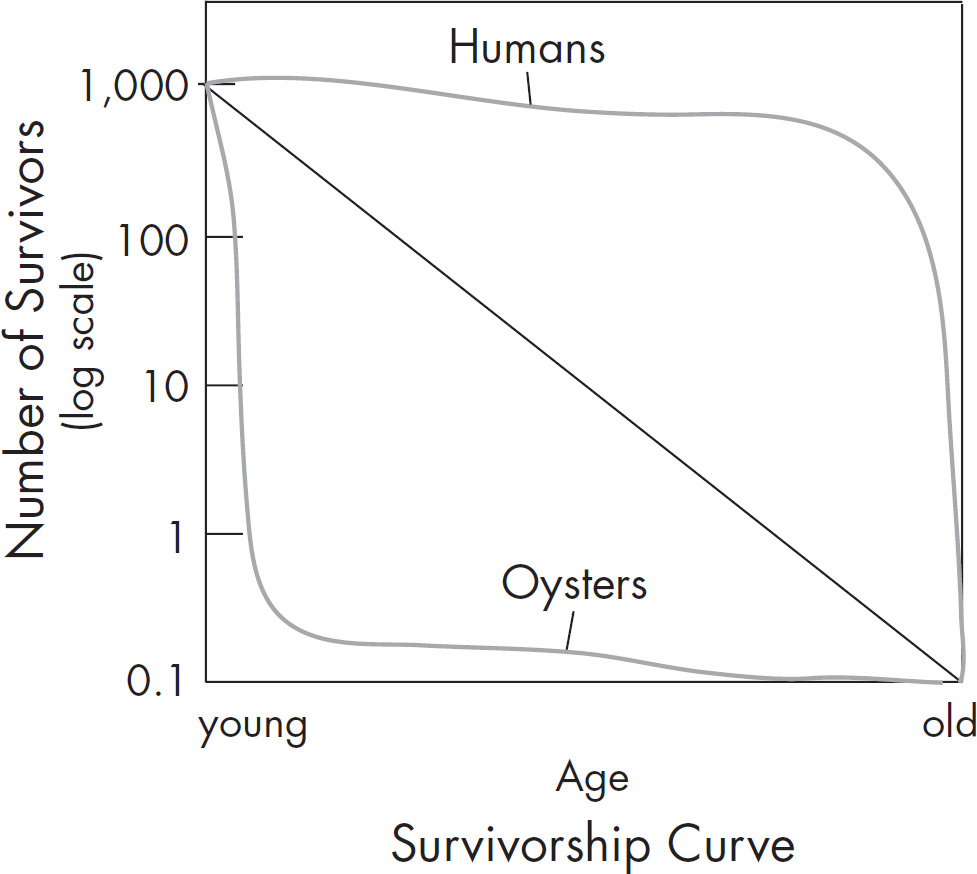
The growth of a population can be represented as the number of crude births minus the number of crude deaths divided by the size of the population.
r = (births – deaths)/N
(r is the reproductive rate, and N is the population size.)
Population growth can also be calculated in the following way:
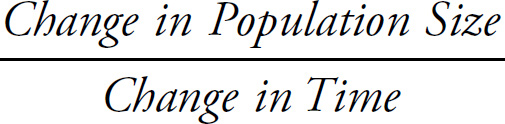 = Birth Rate – Death Rate
= Birth Rate – Death Rate
Each population has a carrying capacity—the maximum number of individuals of a species that a habitat can support. Most populations, however, don’t reach their carrying capacity because they’re exposed to limiting factors.
One important factor is population density. The factors that limit a population are either density-independent or density-dependent. Density-independent factors are factors that affect the population, regardless of the density of the population. Some examples are severe storms and extreme climates. On the other hand, density-dependent factors are those with effects that depend on population density. Resource depletion, competition, and predation are all examples of density-dependent factors. In fact, these effects become even more intense as the population density increases.
Exponential Growth
The growth rates of populations also vary greatly. There are two types of growth: exponential growth and logistic growth. Equations for both are included on the AP Biology Equations and Formulas sheet. Exponential growth occurs when a population is in an ideal environment. Growth is unrestricted because there are lots of resources, space, and no disease or predation. Here’s an example of exponential growth. Notice that the curve arches sharply upward—the exponential increase.
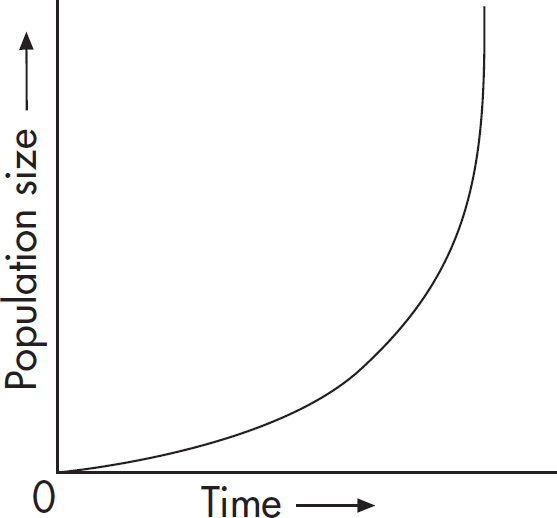
Exponential growth occurs very quickly, resulting in a J-shaped curve. A good example of exponential growth is the initial growth of bacteria in a culture. There’s plenty of room and food, so they multiply rapidly.
However, as the bacterial population increases, the individual bacteria begin to compete with each other for resources. When the resources become limited, the population reaches its carrying capacity, and the curve levels off.
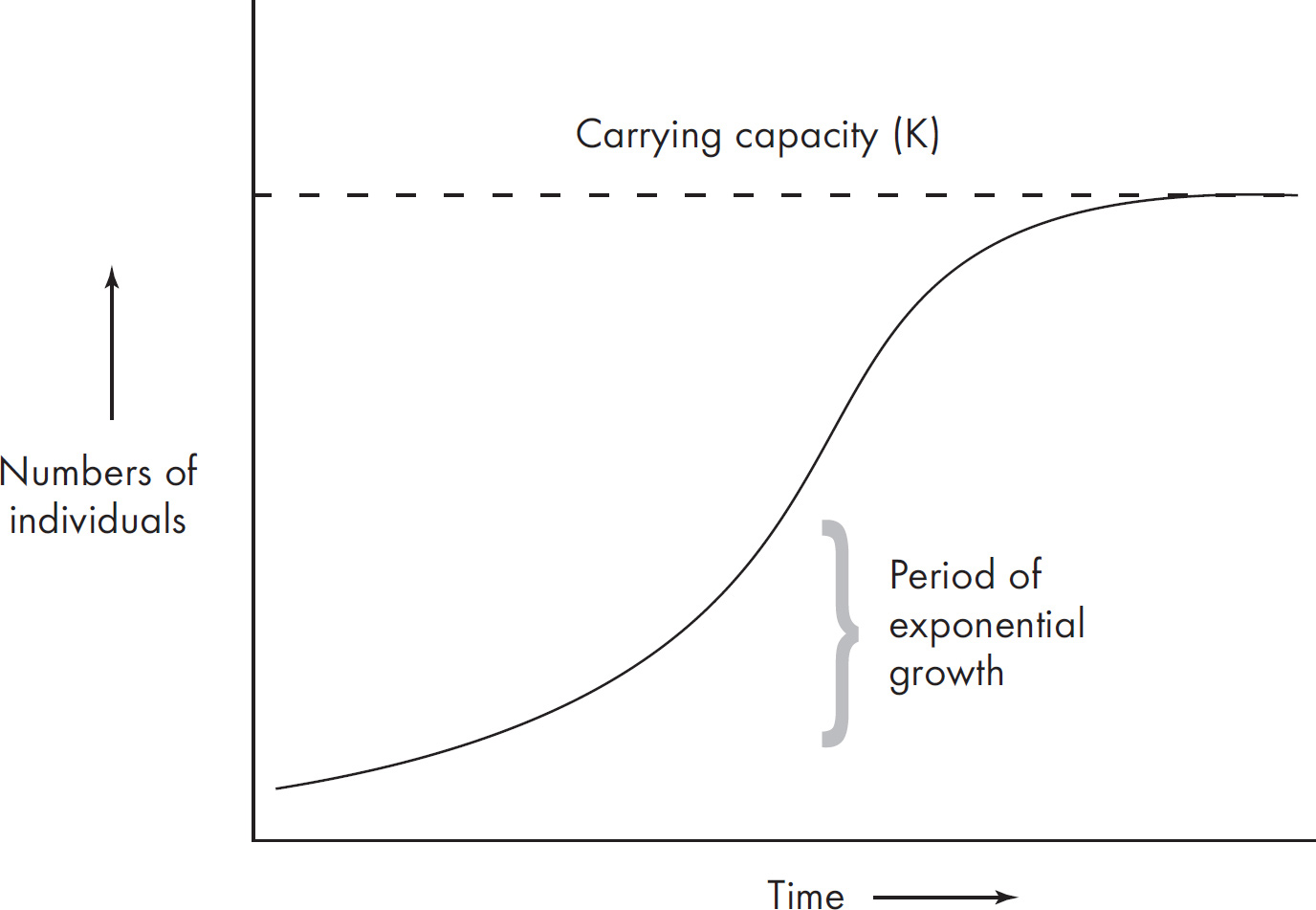
The population becomes restricted in size because of limited resources. This is referred to as logistic growth. Notice that the growth forms an S-shaped curve.
These growth patterns are associated with two kinds of life-history strategies: r-selected species and k-selected species.
R-strategists tend to thrive in areas that are barren or uninhabited. Once they colonize an area, they reproduce as quickly as possible. Why? They know they’ve got to multiply before competitors arrive on the scene! The best way to ensure their survival is to produce lots of offspring. Typical examples are common weeds, dandelions, and bacteria.
At the other end of the spectrum are the k-strategists. These organisms are best suited for survival in stable environments. They tend to be large animals, such as elephants, with long lifespans. Unlike r-strategists, they produce few offspring. Given their size, k-strategists usually don’t have to contend with competition from other organisms.
ECOLOGICAL SUCCESSION
Communities of organisms don’t just spring up on their own; they develop gradually over time. Ecological succession refers to the predictable procession of plant communities over a relatively short period of time (decades or centuries).
Centuries may not seem like a short time to us, but if you consider the enormous stretches of time over which evolution occurs, hundreds of thousands or even millions of years, you’ll see that it is pretty short.
The process of ecological succession in which no previous organisms have existed is called primary succession.
The Pioneers
How does a new habitat full of bare rocks eventually turn into a forest? The first stage of the job usually falls to a community of lichens. Lichens are hardy organisms. They can invade an area, land on bare rocks, and erode the rock surface, over time turning it into soil. Lichens are considered pioneer organisms.
Once lichens have made an area more habitable, they’ve set the stage for other organisms to settle in. Communities establish themselves in an orderly fashion. Lichens are replaced by mosses and ferns, which in turn are replaced by tough grasses, then low shrubs, then evergreen trees, and, finally, the deciduous trees. Why are lichens replaced? Because they can’t compete with the new plants for sunlight and minerals.
The entire sequence is called a sere. The final community is called the climax community. The climax community is the most stable. In our example, the deciduous trees are part of the climax community.
Now what happens when a forest is devastated by fire? The same principles apply, but the events occur much more rapidly. The only exception is that the first invaders are usually not lichens, but grasses, shrubs, saplings, and weeds. When a new community develops where another community has been destroyed or disrupted, this event is called secondary succession.
HUMAN IMPACT ON THE ENVIRONMENT
Unfortunately, humans have disturbed the existing ecological balance, and the results are far-reaching. Soils have been eroded and various forms of pollution have increased. The potential consequences on the environment are summarized below.
• Greenhouse effect: The atmospheric concentrations of carbon dioxide increased by the burning of fossil fuels and forests have contributed to the warming of the earth. Higher temperatures may cause the polar ice caps to melt and flooding to occur. Other potential effects of global warming include changes in precipitation patterns, changes in plant and animal populations, and detrimental changes in agriculture.
• Ozone depletion: Pollution has also led to the depletion of the atmospheric ozone layer by such chemicals as chlorofluorocarbons (CFCs), which are used in aerosol cans. Ozone (O3) forms when UV radiation reacts with O2. Ozone protects the earth’s surface from excessive ultraviolet radiation. Its loss could have major genetic effects and could increase the incidence of cancer.
• Acid rain: Burning fossil fuels produces pollutants such as sulfur dioxide and nitrogen dioxide. When these compounds react with droplets of atmospheric water in clouds, they form sulfuric and nitric acids, respectively. The rain that falls from these clouds is weakly acidic and is called acid rain. Acid rain lowers the pH of aquatic ecosystems and soil, which damages water systems, plants, and soil. For example, the change in soil pH causes calcium and other nutrients to leach out, which damages plant roots and stunts their growth. Furthermore, useful microorganisms that release nutrients from decaying organic matter into the soil are also killed, resulting in less nutrients being available for the plants. Low pH also kills fish, especially those that have just hatched.
• Desertification: When land is overgrazed by animals, it turns grasslands into deserts and reduces the available habitats for organisms.
• Deforestation: When forests are cleared (especially by the slash and burn method), erosion, floods, and changes in weather patterns can occur.
• Pollution: Another environmental concern is the toxic chemicals in our environment. One example is DDT, a pesticide used to control insects. DDT was overused at one time and later found to damage plants and animals worldwide. DDT is particularly harmful because it resists chemical breakdown, and it can still be found in the tissues of nearly every living organism. The danger with toxins such as DDT is that as each trophic level consumes DDT, the substance becomes more concentrated by a process called biomagnification.
• Reduction in biodiversity: As different habitats have been destroyed, many plants and animals have become extinct. Some of these plants could have provided us with medicines and products that may have been beneficial.
KEY TERMS
behavior
ecology
instinct
fixed action pattern
learning
imprinting
critical period
classical conditioning
associative learning
operant conditioning (or trial-and-error learning)
habituation
circadian rhythms
pheromones
agonistic behavior
dominance hierarchy
territoriality
altruistic behavior
symbiotic relationship
mutualism
commensalism
parasitism
photoperiodism
tropism
phototropism
gravitropism
thigmotropism
biosphere
ecosystem
community
population
biomes
abiotic factors
biotic factors
carbon cycle
niche
food chain
producers
primary productivity
primary consumers
herbivores
secondary consumers
tertiary consumers
decomposer
keystone species
10% rule
ecological pyramid
population growth
carrying capacity
population density
density-independent factors
density-dependent factors
exponential growth
logistic growth
r-strategists
k-strategists
ecological succession
primary succession
pioneer organisms
sere
climax community
secondary succession
greenhouse effect
ozone depletion
acid rain
desertification
deforestation
pollution
biomagnification
reduction in biodiversity
Summary
Behavior
 Behavior is an organism’s response to the environment. Behavior can be instinctual (inborn), learned, or social:
Behavior is an organism’s response to the environment. Behavior can be instinctual (inborn), learned, or social:
• Types of learned behaviors include imprinting, classical conditioning and operant conditioning, and insight.
• Types of social behaviors include agonistic behavior, dominance hierarchies, territoriality, and altruistic behavior.
 There are also plant-specific behaviors known as tropisms. The three basic tropisms are phototropism, gravitropism, and thigmotropism.
There are also plant-specific behaviors known as tropisms. The three basic tropisms are phototropism, gravitropism, and thigmotropism.
Ecology
 There are several major biomes (tundra, taiga, temperate deciduous forest, grasslands, deserts, tropical rainforests) that make up the biosphere. Each biosphere contains ecosystems.
There are several major biomes (tundra, taiga, temperate deciduous forest, grasslands, deserts, tropical rainforests) that make up the biosphere. Each biosphere contains ecosystems.
 Within an ecosystem are communities, which consist of organisms fulfilling one of three main roles:
Within an ecosystem are communities, which consist of organisms fulfilling one of three main roles:
• Producers, or autotrophs, convert light energy to chemical energy via photosynthesis.
• Consumers, or heterotrophs, acquire energy from the things they consume. Their digestion of carbohydrates produces carbon, hydrogen, and oxygen, which are then used to make organic substances.
• Decomposers form fossil fuels from the detritus of other organisms in the ecosystem.
• The 10% rule says that only 10% of the energy consumed from one level will be retained by the higher level that consumed it. The other energy will be spent to perform normal daily activities.
 The smallest unit of ecology is the population. The growth of a population can be found with the equation (r) = (births – deaths) / N.
The smallest unit of ecology is the population. The growth of a population can be found with the equation (r) = (births – deaths) / N.
 The carrying capacity is the maximum number of individuals that can be supported by a habitat. Most populations do not reach carrying capacity due to factors such as population density (density-independent factors and density-dependent factors).
The carrying capacity is the maximum number of individuals that can be supported by a habitat. Most populations do not reach carrying capacity due to factors such as population density (density-independent factors and density-dependent factors).
 Exponential growth (J-shaped curve) occurs when a population is an ideal environment. Logistic growth (S-shaped curve) of a population occurs when there are limited resources in an environment.
Exponential growth (J-shaped curve) occurs when a population is an ideal environment. Logistic growth (S-shaped curve) of a population occurs when there are limited resources in an environment.
 Organisms are generally either r-strategists or k-strategists. R-strategists ensure their survival by producing lots of offspring. K-strategists, which are usually large animals, produce few offspring but have longer life spans and less competition from other organisms for resources.
Organisms are generally either r-strategists or k-strategists. R-strategists ensure their survival by producing lots of offspring. K-strategists, which are usually large animals, produce few offspring but have longer life spans and less competition from other organisms for resources.
 Succession describes how ecosystems recover after a disturbance in terms of pioneers, sere, climax community, and secondary succession.
Succession describes how ecosystems recover after a disturbance in terms of pioneers, sere, climax community, and secondary succession.
 Human impact on the planet includes the following issues:
Human impact on the planet includes the following issues:
• greenhouse effect
• ozone depletion
• acid rain
• desertification
• deforestation
• pollution
• reduction in biodiversity
Chapter 12 Drill
Answers and explanations can be found in Chapter 16.
1. A chimpanzee stacks a series of boxes on top of one another to reach a bunch of bananas suspended from the ceiling. This is an example of which of the following behaviors?
(A) Operant learning
(B) Imprinting
(C) Instinct
(D) Insight
2. Viruses are obligate intracellular parasites, requiring their host cells for replication. Consequently, viruses generally attempt to reproduce as efficiently and quickly as possible in a host. Below is a graph depicting the initial growth pattern of a bacteriophage within a population of E. coli. This reproductive strategy is most similar to which of the following?
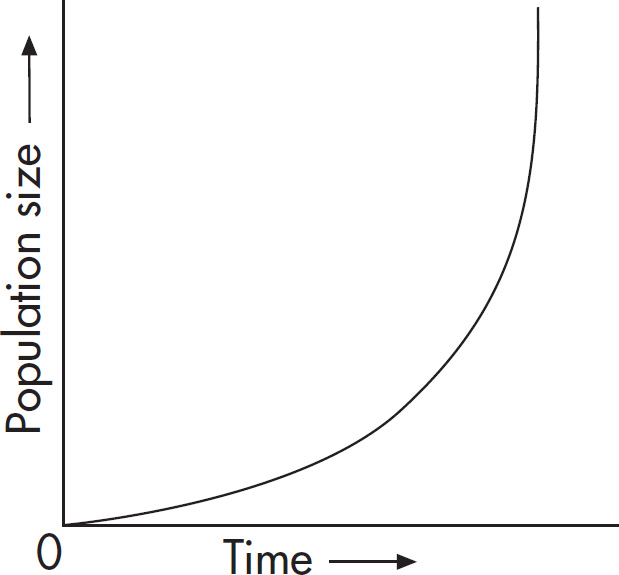
(A) An r-strategist, because it aims to produce a large abundance of offspring to ensure survival
(B) A k-strategist, because it aims to produce a large abundance of offspring to ensure survival
(C) An r-strategist, because it is best suited to thrive in stable environments and over a long life-span
(D) A k-strategist, because it is best suited to thrive in stable environments and over a long life-span
3. In a pond ecosystem, spring rains trigger an expansion of species at levels of the food chain. Runoff from nearby hills brings nutrients which, when combined with warming temperatures, trigger an algae bloom. The populations of small protozoans such as plankton expand by ingesting the algae. These plankton, subsequently, are consumed by small crustaceans such as crayfish, which ultimately become prey for fish such as catfish or carp. In this ecosystem, which of the following accurately describes the crayfish?
(A) They are producers.
(B) They are primary consumers.
(C) They are secondary consumers.
(D) They are tertiary consumers.
Questions 4-6 refer to the following table, chart and paragraph.
Table 1: Minimum pH Tolerance of Common Aquatic Organisms
| Animal | pH Minimum |
| Brook Trout | 4.9 |
| American Bullfrog | 3.8 |
| Yellow Perch | 4.6 |
| Tiger Salamander | 4.9 |
| Crayfish | 5.4 |
| Snails | 6.1 |
| Clams | 6.0 |
Over the period of 1980 to 2000, the average pH in Richard Creek changed drastically. An ecological survey was performed to evaluate the effect of detectable decreases in pH on the aquatic life of the creek. Four times a year, ecological surveys were performed to identify the amount of snails (a primary consumer), crayfish (a secondary consumer), and brook trout (a tertiary consumer) present at five different locations. The percent change relative to 1980 is shown in Chart 1 above. Many aquatic organisms cannot live in low pH conditions. The minimum pH necessary for common aquatic organisms to sustain life is shown in Table 1.
4. According to the data in Chart 1 and Table 1, the average pH in the creek is most nearly which of the following?
(A) 5.9
(B) 5.5
(C) 5.1
(D) 4.7
5. Which of the following is the most likely explanation for the abrupt decrease in pH in Richard Creek?
(A) Greenhouse effect
(B) Deforestation
(C) Acid rain
(D) Pollution
6. In order for snails to return to Richard Creek, the pH of the creek must exceed
(A) 5.4
(B) 5.7
(C) 6.0
(D) 6.1
7. Mimosa pudica is a plant often called the “sensitive plant” because when you touch the leaves, they immediately close up. One theory about the purpose of this type of movement is that herbivores avoid the plant due to this movement. The movement of Mimosa pudica is an example of
(A) phototropism
(B) gravitropism
(C) aquatropism
(D) thigmotropism
Question 8 refers to the following graph.
8. A proponent of the vegan diet writes a blog post that claims that the vegan diet is the only way to “reduce your carbon footprint, with respect to food, because it cuts your carbon footprint in half” and shows the graph above. Which of the following directly supports this claim?
(A) According to the 10% rule, only 10% of energy is passed from producers to consumers, so when humans consume animals, the energy harvested is greatly reduced.
(B) Livestock produce methane emissions and consume large amounts of plants every year.
(C) Deforestation for grazing livestock contributes to the loss of plants for carbon sequestration.
(D) Beef and lamb livestock—not chicken, veal, or pork—has the most dramatic impact on carbon footprint.
9. Costa Rica is plentiful with many beautiful tropical rainforests. For a long time, cattle farmers cleared much of the forests for their livestock. However, in recent decades, the government has placed more emphasis on reforestation. Several fields are now being reclaimed by the rainforest. One such field has many long grasses, a few tall palms, and some dense shrubbery along the periphery of the field. This field can be said to be
(A) invaded by pioneer species
(B) a product of secondary succession
(C) a climax community
(D) filled with invasive species
10. Probiotics are beneficial bacteria or yeast applied to maintain intestinal health in humans or livestock. In theory, these organisms inhabit the intestines and aid in digestion of food. Also, these organisms have bioprotective effects. How can addition of bacteria to the intestine protect the host from disease?
(A) Probiotic bacteria crowd the intestine, preventing pathogenic bacteria from growing due to density-dependent limitation on their population.
(B) Probiotic bacteria crowd the intestine, preparing a niche for pathogenic bacteria to grow.
(C) Bacteria populations grow exponentially until the carrying capacity is reached.
(D) Pathogenic bacteria do not recognize the intestinal cells because they are coated with probiotic bacteria.
REFLECT
Respond to the following questions:
• Which topics from this chapter do you feel you have mastered?
• Which content topics from this chapter do you feel you need to study more before you can answer multiple-choice questions correctly?
• Which content topics from this chapter do you feel you need to study more before you can effectively compose a free response?
• Was there any content that you need to ask your teacher or another person about?
Chapter 13
Quantitative Skills and Biostatistics
There will be six designated quantitative questions on the exam, but this does not mean that the multiple-choice and the free-response questions will not be quantitative. These types of questions pop up all over the place. Even if think you are a whiz, check out the formula page at the back of this book. Expect at least a couple questions using formulas from this sheet.
It is often easy to see patterns in data, but it is not always easy to determine if a pattern is valid or significant. Quantitative data analysis is the first step in figuring this out. In this chapter, we will review how data can be summarized, presented, and tested for validity. This depends on what kind of question was being asked at the beginning of the experiment. When you’re reading this chapter, pay attention to which techniques are used when. On the AP Biology Exam, you should be able to justify which techniques are best.
This list is a good reference for the types of quantitative questions you should be able to tackle. We will not walk though all of these in detail, but you will find some examples in the end-of-chapter drill and in the practice tests. Use the formula sheet to familiarize yourself with these concepts.
Must-Know Types of Quantitative Problems
□ Hardy-Weinberg Equilibrium
□ Water Potential/Osmosis
□ Energy Pyramids/Biomass
□ Chi-squared Analysis
□ Gene Linkage Analysis
□ Inheritance I Probability
□ Reading from Graphs
□ Predicting from Graphs
□ Gibbs Free Energy
□ Dilutions of Solutions
□ Population Growth
□ Primary Productivity
□ Q10
SUMMARIZING AND PRESENTING DATA
Instead of presenting raw data, scientists use descriptive statistics and graphs to summarize large datasets, present patterns in data, and communicate results. Descriptive statistics summarize and show variation in the data. There are many types of descriptive statistics, including measures of central tendency (such as mean, median, and mode) and measures of variability (such as standard deviation, standard error, and range). The AP Biology Equations and Formulas sheet contains definitions for mean, median, mode and range, and equations for mean, standard deviation, and standard error. That said, you will not need to calculate standard deviation or standard error. Instead, focus on understanding how these values are used.
Descriptive statistics are used to summarize the data collected from samples, but may also describe the entire population you’re trying to study. Experiments are designed to include a sample, which is a subset of the population being studied. The best experiments use random sampling, which makes sure there is no bias when picking which individuals from the population will be included in the sample. It is important that the sample is big enough that the data from the experiment are a good representation of what would happen if the whole population were measured.
Graphs are visual representations of data and are often used to reveal trends that might not be obvious by looking at a table of numbers. There are six types of graphs you should be familiar with:
1. Bar graph
2. Pie graph
3. Histogram
4. Line graph
5. Box-and-whisker plot
6. Scatterplot
Each is described below. Pay attention to which graph is used when. On the AP Biology Exam, you may need to decide which graph is best and then create it. You may also have to analyze graphs given to you on the exam.
A good graph must include the following things:
• a good title that describes what the experiment was and what was measured
• axes labeled with numbers, labels and units, and index marks
• a frame or perimeter
• data points that are clearly marked (e.g., easily identifiable lines or bars)
TYPES OF DATA
Data can be quantitative (based on numbers or amounts that can be measured or counted) or qualitative (data that is descriptive, subjective, or difficult to measure). For example, behavioral observations are qualitative. Most data on the AP Biology Exam will be quantitative, with either counts or measurements.
Count data
Count data are generated by counting how many of an item fit into a category. For example, you could count the number of organisms with a particular phenotype or the number of animals in one habitat versus another. Count data also include data that is collected as percentages or the results of a genetic cross. This type of data is usually summarized using a bar graph or a pie graph. For example, suppose a mixed population of E. coli were plated on growth plates. Some cells contained a β-galactosidase marker and grew in blue colonies, while others did not contain the marker and grew in white colonies. The colonies were counted after twenty-four hours of growth.
In another example, suppose fly populations were monitored in northern Maine to determine if population size varies over the warm months of the year.
Count data can be analyzed using hypothesis testing, which will be described later in this chapter.
Measurements
Measurements are continuous, meaning there is an infinite number of potential measurements over a given range. Size, height, temperature, weight, and response rate are all measurements. There are two types of measurement data: parametric and nonparametric.
Normal or Parametric Data
Normal or parametric data are measurement data that fit a normal curve or distribution, usually when a large sample size is used. For example, if you took a large sample of seventeen-year-olds in America and graphed the frequency of heights, the results will be normally distributed.
Several descriptive statistics can be used to summarize normal data.
The sample size (n) is how many members of the population are included in the study. Sample size is an important consideration when you’re trying to determine how well the data in the study represents a population. Large sample sizes are always better, but for technical reasons, experiments can’t be infinitely large.
The mean (x) is the average of the sample, calculated by adding all of the individual values and dividing by the number of values you have. The mean is not necessarily a number provided in the sample. You should be able to recognize what the mean of a given dataset is and be able to calculate the mean using the equation on the AP Biology Equations and Formulas sheet. One limitation of mean is that it is influenced by outliers, or numerical observations that are far removed from the rest of the observations.
The standard deviation (s) can determine if numbers are packed together or dispersed because it is a measure of how much each individual number differs from the mean. A low standard deviation means the data points are all similar and close to the mean, while a high standard deviation means the data are more spread out. Here is the relationship between a normal distribution and standard deviation (or SD).
This means that about 70 percent of the data is within one standard deviation of the mean in a normally distributed dataset. Standard error (SE) can also be used to report how much a given dataset varies and is calculated by dividing the standard deviation by the square root of the sample size.
Nonparametric Data
Nonparametric data often include large outliers and do not fit a normal distribution. In order to determine if data is parametric or not, you can construct a histogram or frequency diagram. These graphs give information on the spread of the data and the central tendencies. Making a histogram is like setting up bins, or intervals with the same range, that cover the entire dataset (the x-axis). The number of measurements in each bin are graphed on the y-axis. For example, suppose you counted the number of pine needles on each branch of a Pinus strobus tree. A histogram of this data may look like this.
This dataset does not match a normal distribution, meaning it is a nonparametric dataset. Keep in mind that even if a certain dataset is not normal, the population itself could be. There could have been sampling bias or errors in the data collection, which can lead to sample data that doesn’t match population data. The best way to fix this is usually to increase the sample size. Histograms differ from bar graphs in that they show ranges instead of just categories.
Nonparametric data requires a different set of descriptive statistics than parametric data.
The median is the middle number in a dataset and is determined by putting the numbers in consecutive order and finding the middle number. If there is an odd number of numbers, there will be a single number that is the median. If there is an even number of numbers, the median is determined by averaging the two middle numbers. Therefore, the median is not necessarily one of the numbers in the dataset. The median is useful in gauging the midpoint of the data, but will not necessarily tell you much about outliers.
The mode is the most frequently recurring number in the dataset. If there are no numbers tha-t occur more than once, there is no mode. If there are multiple numbers that occur most frequently, each of those numbers is a mode. The mode must be one of the numbers in the sample, and modes are never averaged.
The range is the difference between the smallest and largest number in a sample. This value is less useful than standard deviation, because it does not give any information on individual values or the majority of values.
Example 1: Eleven male laboratory mice were weighed at thirty-two weeks of age. The data collected was: 32 g, 28 g, 29 g, 34 g, 30 g, 28 g, 32 g, 31 g, 30 g, 32 g, 33 g. What is the mean, median and mode of this dataset?
Solution: Let’s start by putting the 11 numbers in the dataset in order, from smallest value to largest value: 28 g, 28 g, 29 g, 30 g, 30 g, 31 g, 32 g, 32 g, 32 g, 33 g, 34 g. The mean is
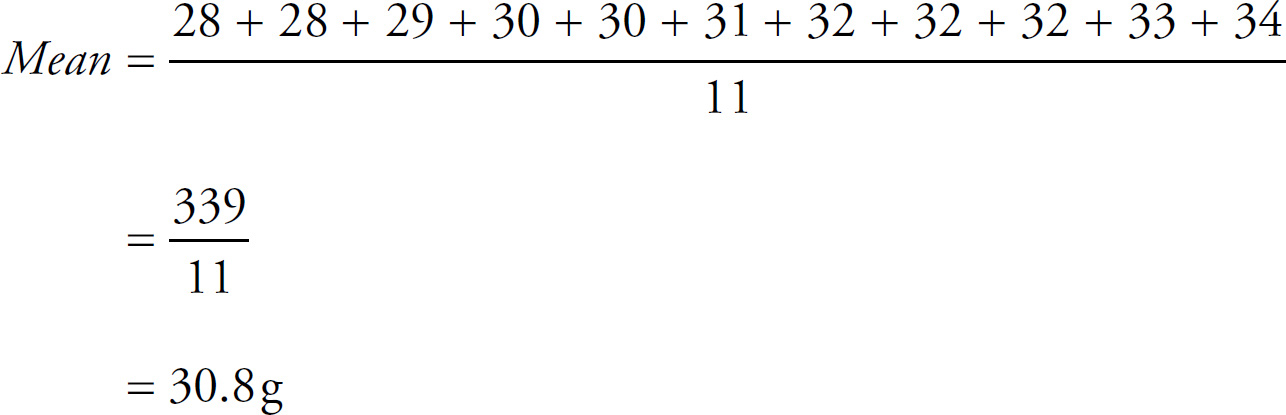
The median is the middle value, or 31 g. The mode is the most frequent number, or 32 g.
Example 2: In the following set of values, what is the range?
Values: −5, 8, 11, −1, 0, 4, 14
Solution: The smallest value in the set above is −5, and the largest is 14. The difference between these two is the range, which is 19.
TYPES OF EXPERIMENTS OR QUESTIONS
You will be tested on scientific experiments and the scientific reasoning behind different aspects of the experiments a LOT. You should be clear with the steps of the scientific method and know how to properly design an experiment and recognize a poorly designed experiment. The following list should remind you:
• Hypothesis: A well-thought-out prediction of what the outcome of the experiment will be.
• Independent Variable: The factor that you, as the experimenter, will change between the different groups in the experiment. If one plant is grown in pH 5 and the other in pH 7, then pH is the independent variable.
• Dependent Variable: The data. The thing that you measure during the experiment. The height of the plant you are growing might be the dependent variable.
• Constants (Controlled Variables): The things that are the same between all your groups. Hint: Everything should be constant except the independent variable.
• Control Groups: Any group that is needed simply so you can compare your interesting experimental groups to it. These are often the “no treatment” group.
• Statistical Significance: This is how trustworthy the results are and how sure you can be in your conclusions about them. This can always be increased by including more individuals in the groups or including more trials.
Most biological experiments do one of three things:
1. look at how something changes over time
2. compare groups of some sort
3. test for an association
How you present and summarize data depends on what kind of experiment you perform.
Time-Course Experiments
These experiments look at how something changes over time. A line graph is usually used to present this type of data. Several lines can be plotted on the same graph, but these must be clearly labeled. Also, each dot on a line graph could represent one data point or a mean of values. If mean values are plotted, standard deviation or standard error can also be shown. More generally, line graphs can also be used to compare how a dependent variable (y-axis) changes with an independent variable (x-axis).
When making line graphs, the intervals on the axes must be consistent. Also make sure it is clear whether the data start at the origin (0, 0) or not. The same guidelines apply to scatterplots, which will be discussed below.
Data points should be connected with a solid line. You can extrapolate (or continue) the line past the data points, but then must use a broken line. Finally, the slope on a line graph tells you the rate of change.
Suppose we want to determine how stress hormone levels change after the end of an important exam. If we measure cortisol and epinephrine in the serum of several people every hour for five hours after the end of an exam, we would have several values for each time point. We could plot the mean of each measurement and also show data variance using standard error bars. In this example, the distance above the data point is the standard error, and the distance below the data point is the standard error. We call this “±SE.”
Comparative Experiments
These experiments compare populations, groups, or events. There are several options for graphing data from these experiments:
• Bar graphs are helpful to compare categories of data if the data is parametric. It is also a good idea to show variance of the data by including the standard error. This gives information on how different the two means are from each other. Suppose you compare two strains of fission yeast before and after treatment with a drug that inhibits cell wall synthesis. As a measure of viability, you measure how much Myc mRNA was made in each strain with and without treatment. In this case, each bar is a mean of values and the error bars are ±SE, as described above.
• Box-and-whisker plots should be used for nonparametric data. These graphs show median (horizontal line), quartiles (top and bottom of the box), and largest and smallest values (ticks at the tops and bottoms of the lines). In other words, the middle 50% of plant weights is represented by the box, with the median shown with the line, and the highest and lowest values are marked with the top and bottom extenders (whiskers). Remember, the median is not the average, so the line doesn’t have to be in the middle. Suppose the weights of two types of Arizona plants are measured, to determine which is more plentiful in a given niche:
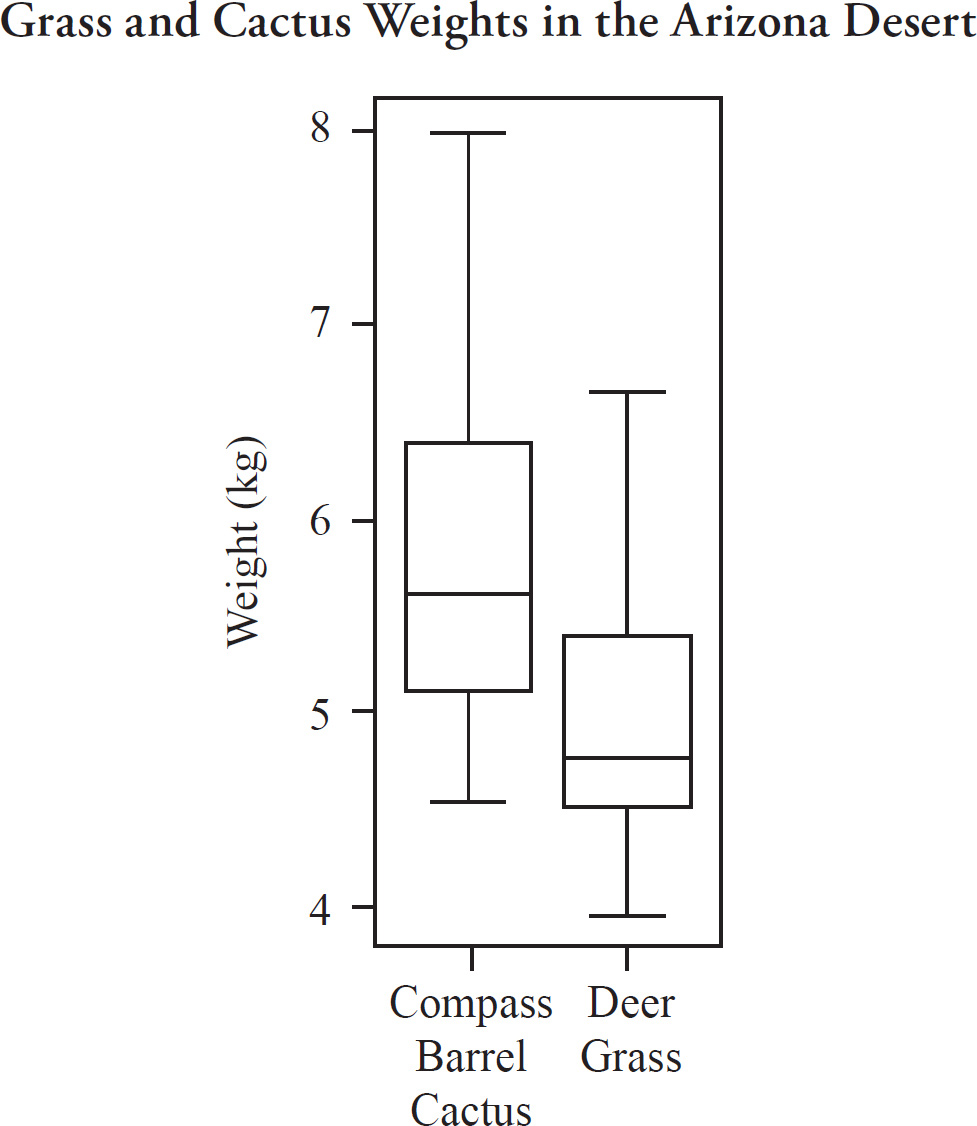
In order to conclude that the samples or groups being compared are different or not, hypothesis testing should be performed. This will be discussed later in this chapter.
Association Experiments
These experiments look for associations between variables. They attempt to determine if two variables are correlated, and additional tests can demonstrate causation. Scatterplots are used to present data from association experiments. Each data point is plotted as a dot. Suppose different substrate concentrations are used in an enzymatic reaction (using the enzyme Kozmase III), and the enzyme efficiency is measured (in percent of maximum). The data could be presented like this.
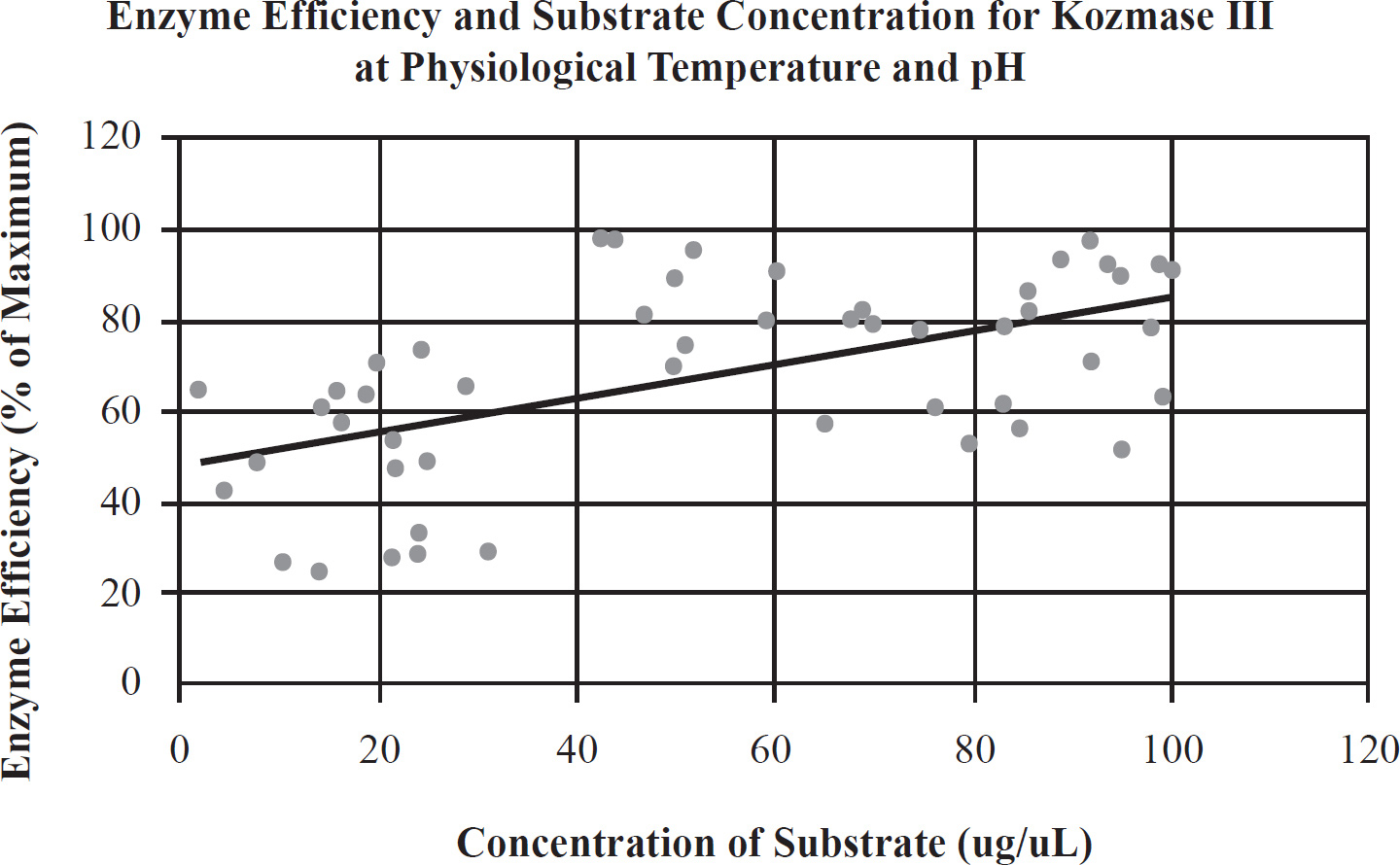
If the relationship looks linear, a linear regression line can be added. There are a few common shapes of scatterplots you should be familiar with:
• A bell-shaped curve is associated with random samples and normal distributions.
• An upward curve (concave in shape) is associated with exponentially increasing functions. A common example of this is the lag phase of bacterial growth in a liquid culture.
• A curve that looks like the sine wave is common when biological rhythms are being studied.
PROBABILITY
You already saw an introduction to probability calculations in Chapter 8 of this book. The probability (P) that an event will occur is the number of favorable cases (a) divided by the total number of possible cases (n).
P = a / n
This can be determined experimentally by observation (such as when population data is being collected) or by the nature of the event. For example, the probability of getting a two when rolling a die is 1/6, since there are six sides on a die.
Example 3: What is the probability of drawing the nine of hearts from a deck of cards?
Solution: Since there are 52 cards in a deck and this question is asking about one particular card, P = 1/52.
Combining Probabilities
Many genetics problems involve several probabilities, all being considered together to answer a larger biological question. There are three rules that are often used.
Product Rule
This rule is used for independent events and is also called the “AND rule.” The probability of independent events occurring together is the product of their separate probabilities. This rule is used when the order of the events matters.
Example 4: What is the probability of drawing a heart and a spade consecutively from a deck of cards if the first draw was replaced before the second draw?
Solution:
P(heart) = 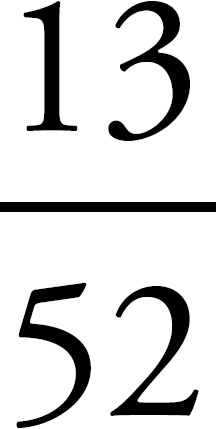 =
= 
P(spade) = 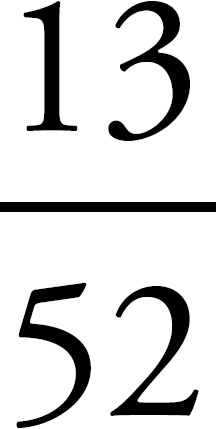 =
= 
P(heart and spade) = P(heart) × P(spade)
P(heart and spade) =  ×
× =
= 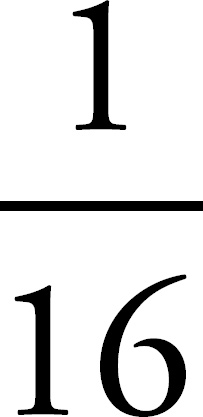
Sum Rule
This rule is used when studying two mutually exclusive events, and can be thought of as the “EITHER/OR″ rule.” The probability of either of two events occurring is the sum of their individual probabilities. This rule is used when the order of the events matters.
Example 5: What is the probability of drawing a diamond or a heart from a deck of cards?
Solution:
P(diamond) = 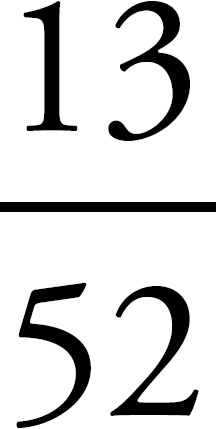 =
= 
P(heart) = 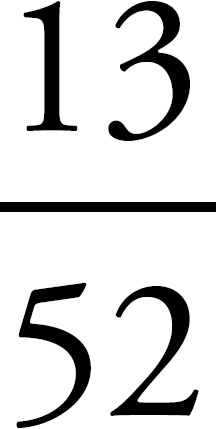 =
= 
P(diamond or heart) = P(diamond) + P(heart)
P(diamond or heart) =  +
+  =
= 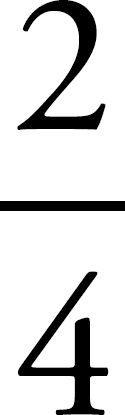 =
= 
HYPOTHESIS TESTING
t-test and p-values
Many experiments involve comparing two datasets or two groups, and a t-test can be used to calculate whether the means of two groups are significantly different from each other. This test is most often applied to datasets that are normally distributed. A p-value equal to or below 0.05 is considered significant in most biology-related fields. T-test calculations involve comparing each data point to the group’s mean and also take standard deviation and sample size into consideration.
Understanding mathematically how p-values are calculated and t-tests are done is not important here. Instead, it’s important that you understand how these values are used to interpret data.
Let’s work through an example to demonstrate how common statistics are used in biological labs: A researcher has sections of two different types of skin cancers from human patients. She stains them for the protein CD31, which is a marker of endothelial cells. She then takes digital pictures of the immunofluorescent sections and counts how many blood vessels are present in each picture. She takes five pictures of each slide.
| Slide A | Slide B |
| 10 | 3 |
| 8 | 2 |
| 7 | 4 |
| 8 | 4 |
| 11 | 4 |
(a) What is the mean, median, and mode for each dataset?
(b) The standard error of group A is 0.735, and the standard error for group B is 0.400. What does this tell you about the data?
(c) What assumptions are made about the data in performing these tests?
(d) What could the researcher do to increase her confidence in the results?
Solutions:
(a) The means are
MeanGroup A = 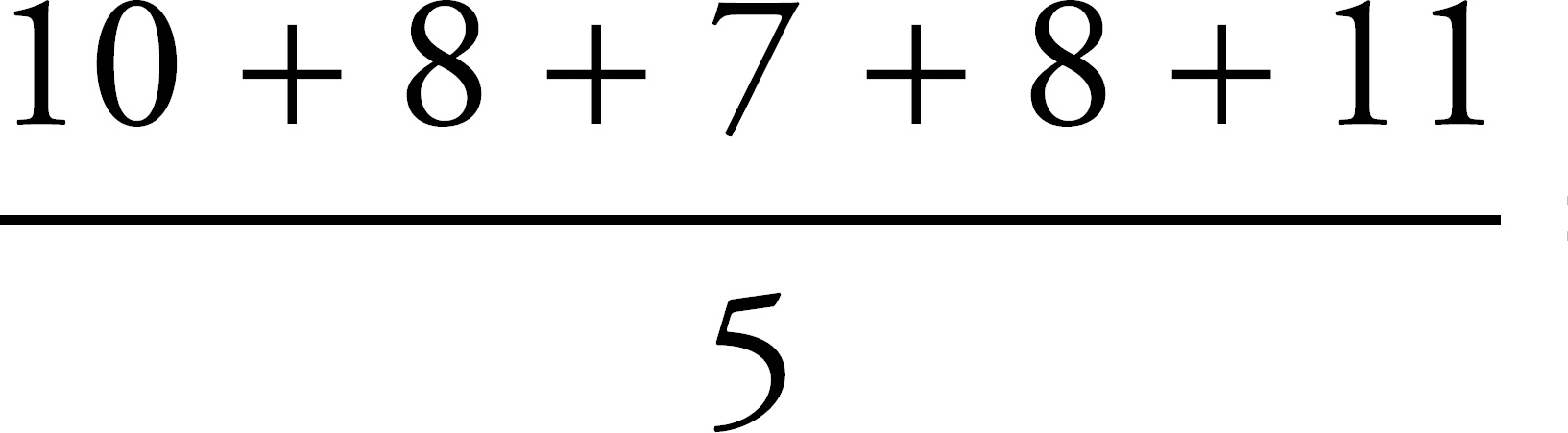 = 8.8
= 8.8
MeanGroup B = 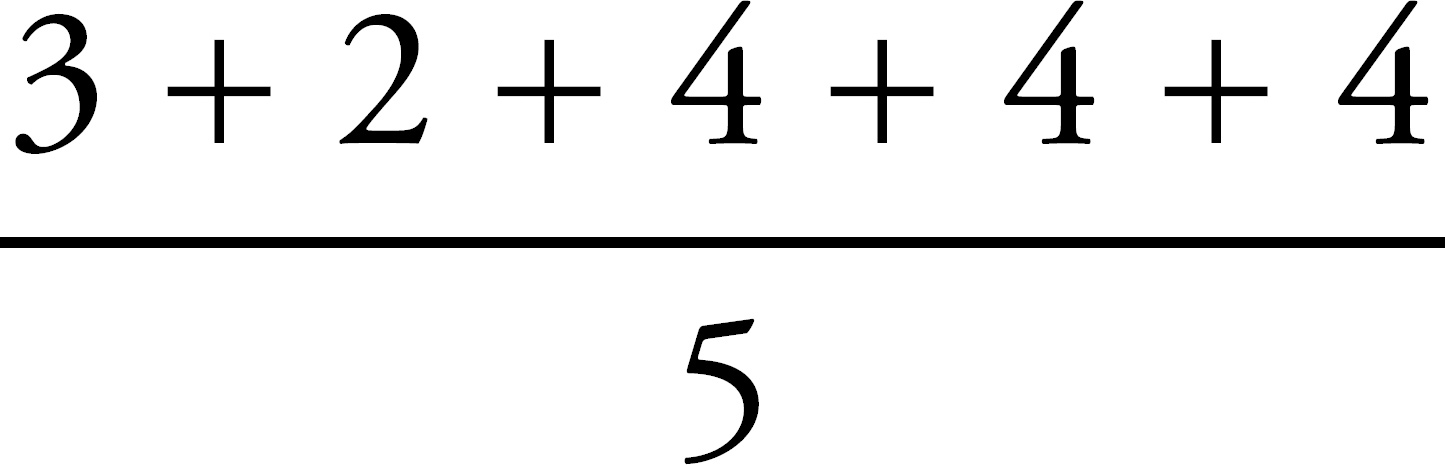 = 3.4
= 3.4
Remember, the median is the middle number, so it’s best to put the data in order first.
| Slide A | Slide B |
| 7 | 2 |
| 8 | 3 |
| 8 | 4 |
| 10 | 4 |
| 11 | 4 |
The median of group A is 8, and the median of group B is 4.
The mode is the most frequent value. For group A, this is 8. For group B, the mode is 4.
(b) Standard error is the standard deviation divided by the square root of the sample size. The sample size is five for both group A and group B because five pictures were taken for each slide. Because the standard error of group A is larger than that of group B and because the two groups have the same sample size, the standard deviation of group A must also be larger than that of group B. Note that this may or may not be the case if the sample sizes were different. A larger standard deviation means the data is more variable, so you can conclude that the data from group A has a larger spread than the data from group B.
(c) As with most datasets, the researcher is assuming the data fits a normal distribution and that her sample size is large enough to be meaningful.
(d) More data allows for more confident conclusions. The researcher could therefore take more pictures from each slide to increase the sample size.
Chi-Square (x2) Tests
A chi square test is a statistical tool used to measure the difference between observed and expected data. For example, suppose you roll a die 120 times. You would expect a 1:1:1:1:1:1 ratio between all the numbers that come up. In other words, you would expect to see each side 20 times. However, because this is an experiment and the sample size is only 120, you will probably see some variation in the data. If you rolled 24 sixes for example, the disagreement between observed (24) and expected (20) is small, and could have occurred by chance. If you rolled 40 sixes however, there is a large disagreement from the expected. Maybe it is a weighted die. The region in between these two values is trickier. Where is the line drawn between expected/normal variations and results that are so different that they must tell a different story altogether.
Chi-square tests are one way to make this decision. They start with a null hypothesis (Ho), which represents a set numerical outcome that you want to compare your data with. The chi-square test will determine if your actual data is close to the null hypothesis outcome and could have occurred due to a fluke (i.e., rolling a six 24 times) or if it is so different that it is probably not a fluke (i.e., 40 rolled sixes) and the null hypothesis outcome must not correctly describe what is occurring. In the dice case, rolling a six 40 times would cause the null hypothesis of expecting a 1:1:1:1:1:1 to be rejected and the die can be considered to be weighted. In the calculations, expected results if the null hypothesis were true are compared to the actual values, and an x2 value is calculated. This value is compared to a critical value, and a decision is made to reject or accept the null hypothesis.
For example, suppose data was collected from 100 families with three children each. Fourteen families had three female children, 36 families had two females and a male, 30 families had one female and two males, and 20 families had three male children. We would expect offspring sex to segregate independently. In other words, the sex of a given child shouldn’t be affected by the sex of previous children. Let’s test whether this is true for this dataset.
Since we expect sex to segregate independently, this is the null hypothesis (Ho). Each family has three children, so there are four possible outcomes for each family:
1. three daughters
2. two daughters and a son
3. one daughter and two sons
4. three sons
Using some advanced math (that you don’t need to worry about), the probabilities for each of these is:
1. P(three females) = 12.5%
2. P(two females, one male) = 37.5%
3. P(one female, two males) = 37.5%
4. P(three males) = (1)(0.50)3 = 12.5%
Because there are 100 families, we expect 12.5 to have three females, 37.5 to have two females and a male, 37.5 to have one female and two males, and 12.5 to have three males. This can be calculated by multiplying the probability of each event by the total (100 in this case).
Next, we compare this expected data (E) to what actually happened (the observed data, O). Note that the total numbers for observed and expected columns must be the same. To calculate a x2 value, the formula (O – E)2/E is calculated for each row of the table, and summed.
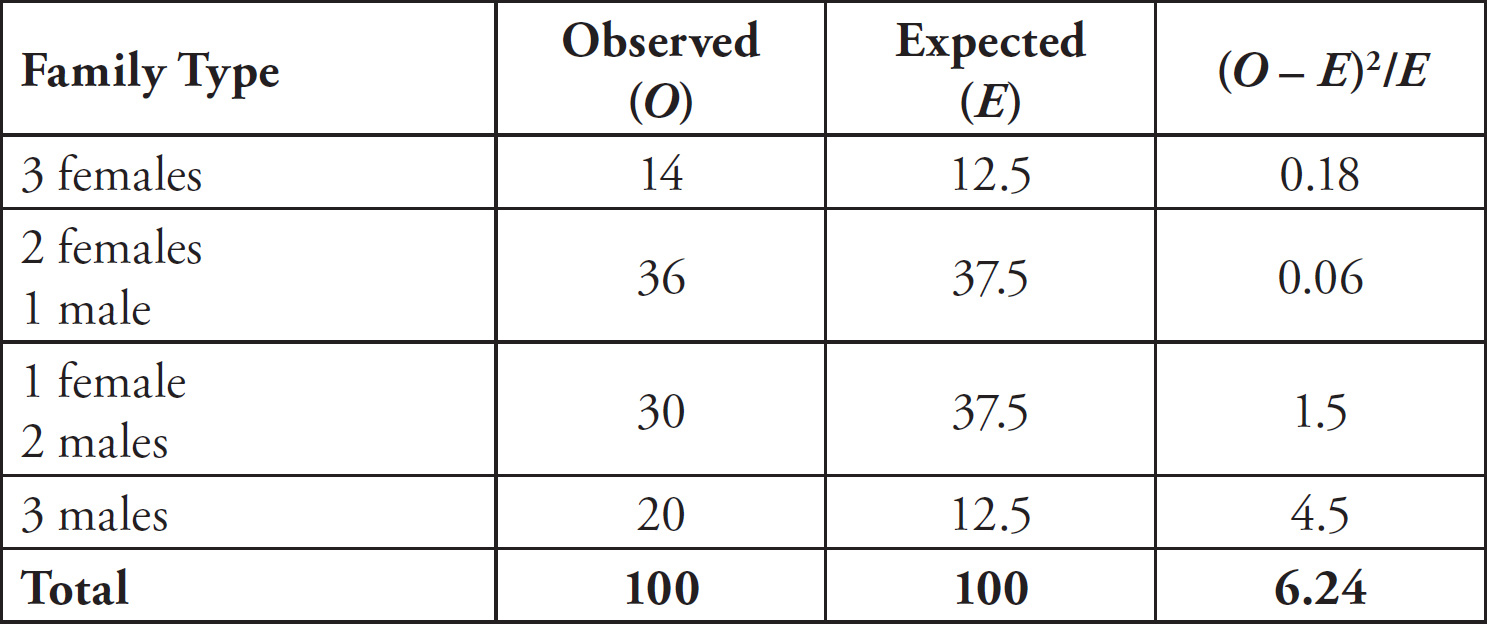
The calculated x2 value is 6.24. Next, this value needs to be compared to a critical value (CV). This is obtained from a table like this one.
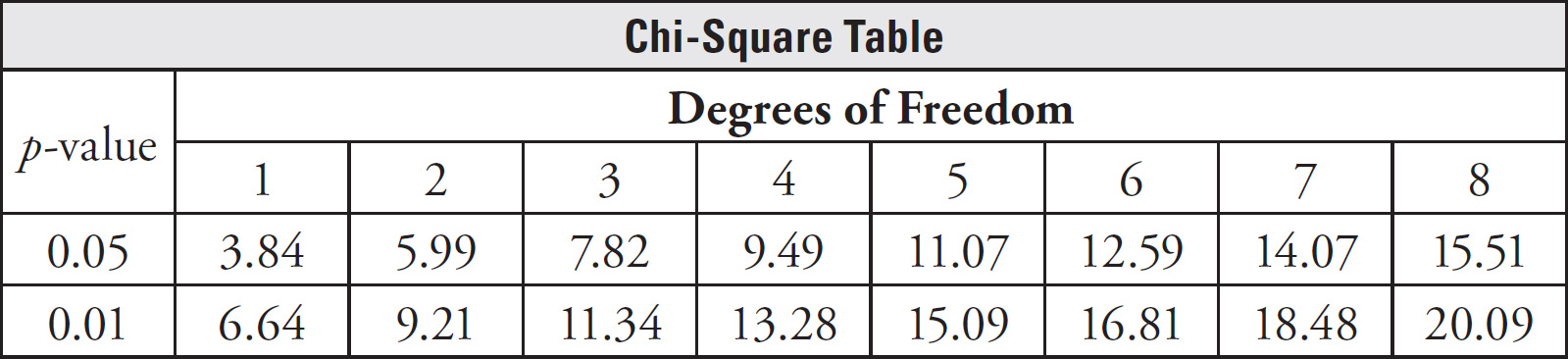
This chart is included on the AP Biology Equations and Formulas sheet. In order to use this table, you must know the degrees of freedom (DF), which represent the number of independent variables in the data. In most cases, this is the number of possibilities being compared minus one. In this example, DF = 4 – 1, because we are comparing four different family types.
You also need a reference p-value. This is the probability of observing a deviation from the expected results due to chance. For most biological tests, a p-value of 0.05 is used.
Since we are using DF = 3 and p = 0.05, the critical value is 7.82 (using the chart above). Next, the critical value is compared to the x2 value; if x2 < CV, you accept Ho. If x2 > CV, you reject Ho. We calculated a x2 value of 6.24, which is smaller than 7.82. Since x2 < CV, we accept Ho and conclude that offspring sex could, indeed, be segregating independently in this dataset.
KEY TERMS
descriptive statistics
sample
population
quantitative data
qualitative data
bar graph
pie graph
normal or parametric data
normal distribution
sample size
mean
outliers
standard deviation
standard error
nonparametric data
histogram or frequency diagram
median
mode
range
hypothesis
dependent variable
independent variable
contants (contolled variables)
controls
statistical significance
line graph
bra graph
box and whisker plot
scatterplot
probability
product rule
sum rule
t-test
p-value
chi-square test
null hypothesis
critical value
degrees of freedom
Summary
 Descriptive statistics (such as mean, standard deviation, standard error, median, mode, range) and graphs are used to summarize data and show patterns and conclusions:
Descriptive statistics (such as mean, standard deviation, standard error, median, mode, range) and graphs are used to summarize data and show patterns and conclusions:
• Normal (parametric) measurement data are summarized using mean and standard deviation or standard error.
• Parametric data are summarized using median, mode, and range.
• Histograms can be used to see if a dataset has a standard deviation or not.
 Count data are summarized using a bar graph or a pie graph.
Count data are summarized using a bar graph or a pie graph.
 Time course experiments look at how something changes over time; summarize these using a line graph.
Time course experiments look at how something changes over time; summarize these using a line graph.
 Experiments that compare groups are summarized using a bar graph (if the dataset has a standard deviation) or a box-and-whisker plot (if it does not).
Experiments that compare groups are summarized using a bar graph (if the dataset has a standard deviation) or a box-and-whisker plot (if it does not).
 Scatterplots summarize association experiments, and regression lines can be used to determine if the relationship is linear.
Scatterplots summarize association experiments, and regression lines can be used to determine if the relationship is linear.
 Probability can be calculated using the sum rule or the product rule.
Probability can be calculated using the sum rule or the product rule.
 Hypothesis testing is used to determine if two groups are significantly different from each other. They start with a null hypothesis, which is rejected or accepted, depending on how a calculated p-value or chi-square value compares to a standard value.
Hypothesis testing is used to determine if two groups are significantly different from each other. They start with a null hypothesis, which is rejected or accepted, depending on how a calculated p-value or chi-square value compares to a standard value.
Chapter 13 Drill
Answers and explanations can be found in Chapter 16.
1. Five subjects were weighed before and after an 8-week exercise program. What is the average amount of weight lost in pounds for all five subjects, rounded to the nearest pound?
| Subject | Starting Weight (pounds) | Final Weight (pounds) |
| 1 | 184 | 176 |
| 2 | 200 | 190 |
| 3 | 221 | 225 |
| 4 | 235 | 208 |
| 5 | 244 | 225 |
(A) 12 pounds
(B) 13 pounds
(C) 14 pounds
(D) 15 pounds
2. The height of six trees is measured. Is plant 6 taller than the median for all six trees?
| Plant | Height (inches) |
| 1 | 67 |
| 2 | 61 |
| 3 | 72 |
| 4 | 71 |
| 5 | 66 |
| 6 | 68 |
(A) Yes, the median is 67.3.
(B) No, the median is 67.3.
(C) Yes, the median is 67.5.
(D) No, the median is 67.5.
3. In the following set of test scores, what is the mode and what is the range?
Test Scores: 71, 67, 75, 65, 66, 32, 69, 70, 72, 82, 73, 68, 75, 68, 75, 78
(A) Mode: 68; Range: 75
(B) Mode: 69; Range: 50
(C) Mode: 75; Range: 70.5
(D) Mode: 75; Range: 50
4. Given the cross AaBbCc × AaBbCc, what is the probability of having an AABbCC offspring?
(A) 
(B) 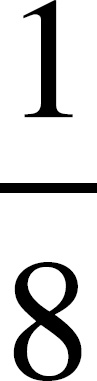
(C) 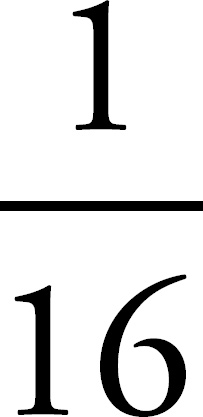
(D) 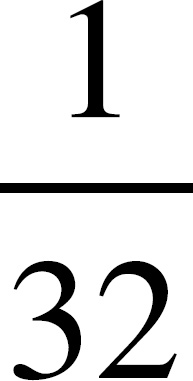
5. Given the cross AaBb x aabb, what is the probability of having an Aabb or aaBb offspring?
(A) 
(B) 
(C) 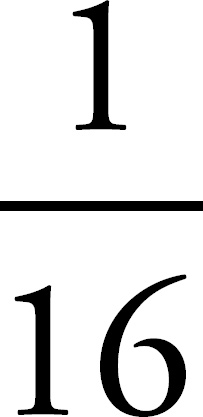
(D) 0
6. Two pea plants are crossed, and a ratio of 3 yellow plants to 1 green plant is expected in the offspring. It is found that out of 100 plants phenotyped, 84 are yellow and 16 are green. Do the experimental data match the expected data?
(A) Yes, the x2 value is greater than 3.84.
(B) Yes, the x2 value is smaller than 3.84.
(C) No, the x2 value is greater than 3.84.
(D) No, the x2 value is smaller than 3.84.
7. A mating is set up between two pure breeding strains of plants. One parent has long leaves and long shoots. The other parent has short leaves and stubby shoots. F1 plants are collected, and all have long leaves and long shoots. F1 plants are self-crossed, and 1000 F2 plants are phenotyped. The data is as follows:
| Phenotype | # of F2 |
| Long leaves, long shoots | 382 |
| Long leaves, stubby shoots | 109 |
| Short leaves, long shoots | 112 |
| Short leaves, stubby shoots | 397 |
| Total | 1000 |
Are the genes for leaf and shoot length segregating independently?
(A) Yes; the degrees of freedom are 3, and the calculated χ2 value is small.
(B) No; the degrees of freedom are 3, and the calculated χ2 value is large.
(C) Yes; the degree of freedom is 1, and the calculated χ2 value is small.
(D) No; the degree of freedom is 1, and the calculated χ2 value is large.
REFLECT
Respond to the following questions:
• Which topics from this chapter do you feel you have mastered?
• Which content topics from this chapter do you feel you need to study more before you can answer multiple-choice questions correctly?
• Which content topics from this chapter do you feel you need to study more before you can effectively compose a free response?
• Was there any content that you need to ask your teacher or another person about?
Chapter 14
Sample Free-Response Questions
SECTION II: FREE RESPONSE
Section II of the AP Biology Exam contains eight questions, consisting of two long free-response questions and six short free-response questions. You will have a total of 90 minutes—including 10 minutes for reading—to complete all questions. The directions will look something like this (read carefully!).
Directions: Questions 1 and 2 are long free-response questions that should require about 22 minutes each to answer and are worth 10 points each. Questions 3 through 8 are short free-response questions that should require about 6 minutes each to answer. Questions 3 through 5 are worth 4 points each, and questions 6 through 8 are worth 3 points each.
Read each question carefully and completely. You are advised to spend the 10-minute reading period planning your answers. You may begin writing your responses before the reading period is over. Write your response in the space provided following each question. Only material written in the space provided will be scored. Answers must be written out in paragraph form. Outlines, bulleted lists, or diagrams alone are not acceptable.
The following sections discuss scoring guidelines in more detail and provide sample questions. But first take some time to review some key tips for writing your free responses.
Free-Response Tips and Strategies
AP Biology Readers (the people who grade your free-response essays) expect students to interpret data and apply knowledge to new situations in the free-response questions. Some important things to keep in mind when writing your answers to the free-response questions are provided in the following list:
• Answer the question. Do not become distracted or waste time writing about tangential topics that you know more about than what is being asked. Focus your response.
• Think quantitatively. Use data whenever possible to explain your answer. This is especially necessary when the question features a graph, table, or other type of diagram (which many free-response questions now do).
• Be specific. Offer specific detailed examples where appropriate.
• Use vocabulary correctly and often. Hone your understanding of the words provided in the Key Terms lists in this book and the various biological ideas and situations to which they can be applied. This will help you use the correct terminology when writing your responses.
• Remember the Big Ideas. Know the four big ideas covered in the AP Biology course (this page), as they are reflected in many free-response questions. Understanding these ideas and being able to articulate them will help you craft your responses.
Okay, now it’s time for some practice. Take about 22 minutes and try to write a long free-response to the following question. Keep these tips in mind as you write.
The regulation of the expression of gene products is crucial to the development of an embryo and the maintenance of homeostasis. The table below illustrates the mRNA levels, total protein levels, and total enzyme activity levels for three different enzymes. These enzymes are encoded by genes A, B, and C after being expressed in both the absence and presence of “Protein X” or “Protein Y.”
Describe the likely role of Protein X and/or Protein Y in the expression of each of these genes. Include when during gene expression the regulation occurs and explain a possible mechanism including a potential binding partner for Protein X and/or Protein Y.
In a follow-up experiment, Protein X and Protein Y are added together. On the axes below, create a graph (or several) predicting the outcome on the mRNA, protein, and enzyme activity of Genes A, B, and C. Be sure to show the untreated levels also.
SCORING GUIDELINES
To help you grade this sample free response, we’ve put together a checklist that you can use to calculate the number of points that should be assigned to each part of this question. We’ll first explain the important points on the checklist and then give you sample student responses to show you how test reviewers would evaluate them.
Free-Response Checklist
1. Describe the roles/point of regulation/possible mechanism and binding partner (2 points for each correct combination for 6 points total)
a. gene A regulation (transcription enhanced by X)
b. gene B regulation (transcription inhibited by X and enzyme activity enhanced by Y
c. gene C regulation (enzyme activity inhibited by Y)
2. Create graph(s)
a. The graph(s) should show the correct trend for each gene in comparison to untreated. (1 point each for 3 points total)
i. gene A (mRNA, protein, and enzyme activity increased)
ii. gene B (mRNA, protein, and enzyme activity reduced, though maybe not as much of a reduction in enzyme as was seen with protein X alone)
iii. gene C (mRNA and protein unchanged, but enzyme activity reduced)
b. The graph should have properly labelled axes (most likely with the levels of mRNA, protein, and enzyme activity on the Y-axis and the different genes on the X-axis). (1 point)
SAMPLE ESSAYS
The following response would earn full points.
Gene A expression has no change with the addition of Protein Y, but it has increased transcription, protein, and enzyme activity with the addition of Protein X. It is likely that Protein X is an enhancer of transcription of gene A. Perhaps it binds directly to the promoter region of gene A and assists RNA polymerase to bind. It might also remove a repressor of transcription or work together with other transcription factors. The increase in mRNA would lead to increased protein and increased enzyme activity down the road.
Gene B has a decrease in mRNA, protein, and enzyme activity with Protein X. It has increased total activity with Protein Y. It is likely that Protein X is a repressor of transcription for gene B. It might bind to the promoter of gene B or to another transcription factor to prevent RNA polymerase binding or prevent transcription from occurring in another way. Since the enzyme activity increases with Protein Y, but the mRNA and the protein levels did not change, this means that the enzyme activity was somehow enhanced on the same amount of enzyme that was present without Protein Y. Protein Y might be an enzyme activator of Enzyme B. It might bind to Enzyme B directly or maybe the protein phosphorylates it or cleaves it to make the enzyme active.
Gene C is unaffected by Protein X, but Protein Y decreases the enzyme activity without affecting the mRNA or protein levels. This means it is acting upon the enzyme and turning it off. Protein Y might be an allosteric or a competitive inhibitor of Enzyme C. If it is an allosteric inhibitor, then it binds to a site other than the active site. If it is a competitive inhibitor, then it binds to the active site on Enzyme C.
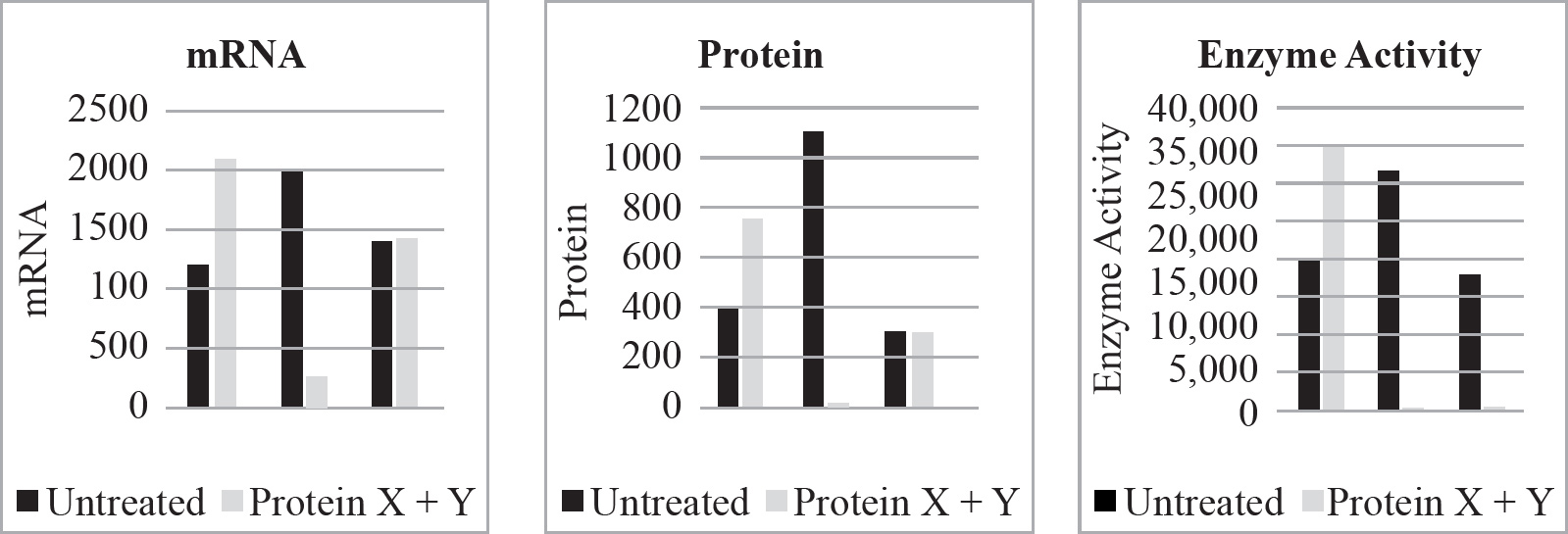
HOW TO USE THE SAMPLE FREE-RESPONSE QUESTIONS
The most important advice for this section of the test is practice, practice, practice! No matter how well you think you know the material, it’s important to practice formulating your thoughts on paper. Before you take our practice tests in the back of the book, make sure you review our sample free-response questions. Then use our checklist—which is put together just like the testing board’s—to assign points.
To give you as much practice as possible, we’ve compiled a few sample long and short free-response questions. Most of these questions refer to themes in biology that crop up time and time again on the exam, so chances are you’ll see something similar on your test. You can also access free-response questions from recent exams on the College Board website at https://apstudent.collegeboard.org/apcourse/ap-biology/exam-practice.
SAMPLE LONG FREE-RESPONSE QUESTIONS
1. Describe how the modern theory of evolution is supported by evidence from the following areas:
(a) Comparative anatomy
(b) Biogeography
(c) Embryology
2. Describe the chemical nature of genes. Discuss the replicative process of DNA in eukaryotic organisms. What are the various types of gene mutations that can occur during replication?
3. Describe the process of photosynthesis. What are the major plant pigments involved in photosynthesis? Design an experiment to measure the rate of photosynthesis.
SAMPLE SHORT FREE-RESPONSE QUESTIONS
1. Describe the structure of a generalized eukaryotic cell.
2. Discuss the following responses in plants and give one example of each.
(a) Photoperiodism
(b) Phototropism
3. Select one of the following pairs of hormones and discuss the concept of negative feedback.
(a) Thyroid-stimulating hormone (TSH) and thyroxine
(b) Parathyroid hormone and calcitonin
(c) Cortisol and ACTH
The biology knowledge you need to answer all of these free-response questions is in this book. After you have completed your essays, go back to the chapters that cover these topics and check to see if you have presented all the relevant points. Use our Free-Response Checklist as a guideline to give yourself (or have someone give you) an approximate score. Good luck!
Summary
 The free-response section features eight questions, two of which require long responses. You will have 90 minutes to complete the entire section.
The free-response section features eight questions, two of which require long responses. You will have 90 minutes to complete the entire section.
 Each question is worth a certain number of points. Generally, the long-response questions are worth 10 points, and the short-response questions are worth either 3 or 4 points each.
Each question is worth a certain number of points. Generally, the long-response questions are worth 10 points, and the short-response questions are worth either 3 or 4 points each.
 Your responses must be written in paragraph form; outlines are not acceptable. Diagrams can be used to supplement your writing but will not receive credit on their own (unless otherwise specified).
Your responses must be written in paragraph form; outlines are not acceptable. Diagrams can be used to supplement your writing but will not receive credit on their own (unless otherwise specified).
 Remember to keep your response focused and use specific data and examples (where applicable) to justify your answers.
Remember to keep your response focused and use specific data and examples (where applicable) to justify your answers.
 Familiarize yourself with the key terms in this book as well as the Big Ideas for the AP Biology course. Understanding these will help you write clear, focused, and accurate responses.
Familiarize yourself with the key terms in this book as well as the Big Ideas for the AP Biology course. Understanding these will help you write clear, focused, and accurate responses.
REFLECT
Respond to the following questions:
• Which topics from this chapter do you feel you have mastered?
• Which content topics from this chapter do you feel you need to study more before you can answer multiple-choice questions correctly?
• Which content topics from this chapter do you feel you need to study more before you can effectively compose a free response?
• Was there any content that you need to ask your teacher or another person about?
Chapter 15
Laboratory
All AP Biology courses have a laboratory component that gives students hands-on experience regarding some of the biology topics covered in class. Through these laboratory exercises you can learn the scientific method, lab techniques, and problem-solving skills. There is not an official set of labs that must be completed, and AP teachers get to choose the ones for their classroom. This means that you will not be tested on memorizing specific protocols or data from the labs. However, all labs teach the same basic skills of critical thinking, developing hypotheses, judging results, making conclusions, adapting experiments, and understanding variables. So, those are the skills that you are expected to fully understand. The exam will likely present you with example experiments and you will need to make decisions and answer questions about them. There are thirteen labs that are very popular to do in AP Biology classes, and these are the same labs that often come up on the AP test. We have given you short summaries of those 13 labs. This way, if you encounter a question about one of these labs, you will be one step ahead of students that have never even heard of it before.
LAB 1: ARTIFICIAL SELECTION
This lab explores artificial selection—the process by which humans decide traits to enhance or diminish in other species by crossing individuals with the desired phenotype. You need to understand the basic principles of natural selection and how natural selection drives evolution, listed below:
• Variation is present in any population.
• Natural selection, or differential reproduction in a population, is a major mechanism in evolution.
• Natural selection acts on phenotypic variations in populations. Some organisms will have traits that are more favorable to the environment and will survive to reproduce more than other individuals, causing the genetic makeup of the population to change over time.
This lab specifically deals with Wisconsin Fast Plants. In order to artificially select for certain traits, plants with the desired traits can be crossed, changing the genetic makeup of the population. For example, if you wanted to select for height, you could cross only the tallest plants with one another. The new population will have a higher mean height than the previous generation.
LAB 2: MATHEMATICAL MODELING: HARDY-WEINBERG
In order to understand everything you need for the AP Biology Exam, you need to be able to use the Hardy-Weinberg principles and equations to determine allele frequencies in a population. One way to study evolution is to study how the frequencies of alleles change from generation to generation:
• Know how to calculate the allele and genotype frequencies using the two Hardy-Weinberg equations: p + q = 1 and p2 + 2pq + q2 = 1. Don’t forget: if the population obeys Hardy-Weinberg’s rules, these frequencies remain constant over time.
• Review the discussion of the Hardy-Weinberg principle in this book. Know the five conditions of the Hardy-Weinberg equilibrium: (1) large population, (2) no mutations, (3) no immigration or emigration, (4) random mating, and (5) no natural selection.
• Review natural selection and how it can lead to changes in the genetic makeup of a population.
LAB 3: COMPARING DNA SEQUENCES TO UNDERSTAND EVOLUTIONARY RELATIONSHIPS WITH BLAST
This laboratory uses BLAST (Basic Local Alignment Search Tool), a database that allows you to input a DNA sequence for a gene to look for similar or identical sequences present in other species. In order to understand everything you need for the AP Exam you need to be able to do the following:
• Look at a phylogenic tree, which is a way to visually represent evolutionary relatedness. Endpoints of each branch correspond to a specific species, and each junction on the tree represents a common ancestor. Species that are closer on a phylogenic tree are more closely related.
• Understand BLAST scoring. Most phylogenic trees are constructed by examining nucleotide sequences; the more identical two species sequences are for a specific gene, the more closely related they are. BLAST is able to analyze different sequences to tell you how similar they are to one another based on a score. The higher the score, the closer the two sequences align.
LAB 4: DIFFUSION AND OSMOSIS
This lab investigates the process of diffusion and osmosis in a semipermeable membrane as well as the effects of solute concentration and water potential on these processes.
What are the general concepts you really need to know?
Fortunately, this lab covers the same concepts about diffusion and osmosis that are discussed in this book. Just remember that osmosis is the movement of water across a semipermeable membrane, from a region of high water concentration to one of low water concentration, or from a hypotonic region (low solute concentration) to a hypertonic region (high solute concentration).
• Be familiar with the concept of water potential, which is simply the free energy of water. It is a measure of the tendency of water to diffuse across a membrane. Water moves across a selectively permeable membrane from an area of higher water potential to an area of lower water potential.
• Be familiar with the effects of water gain in animal and plant cells. In animals, the direction of osmosis depends on the concentration of solutes both inside and outside of the cell. In plants, osmosis is also influenced by turgor pressure—the pressure that develops as water presses against a cell wall. If a plant cell loses water, the cell will shrink away from the cell wall and plasmolyze.
Another important concept to understand is the importance of surface area and volume in cells. There are several questions about each listed on the AP Biology Equations and Formulas sheet. Cells maintain homeostasis by regulating the movement of solutes across the cell membrane. Small cells have a large surface area-to-volume ratio, however as cells become larger, this ratio becomes smaller, giving the cell relatively less surface area to exchange solutes. A cell is limited in size by the surface area-to-volume ratio. There are many organisms that have evolved strategies for increasing surface area, like root hairs on plants and villi in the small intestines of animals.
LAB 5: PHOTOSYNTHESIS
The chemical equation for photosynthesis is
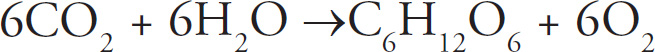
Because plants consume some of this energy during photosynthesis, measuring the oxygen produced by a plant can tell us about the net photosynthesis that is occurring. In this laboratory, photosynthesis rates are measured using leaf discs that begin to float as photosynthesis is carried out, allowing you to see that photosynthesis is occurring.
There are several properties that affect the rates of photosynthesis, including:
• light intensity, color, and direction
• temperature
• leaf color, size, type
Be able to hypothesize about the effects of these variables: for example, as light intensity increases, so does the rate of photosynthesis. Remember, plants and animals both contain mitochondria and carry out cell respiration!
LAB 6: CELLULAR RESPIRATION
In this lab the respiratory rate of germinating and nongerminating seeds and small insects is investigated. The equation for cellular respiration is
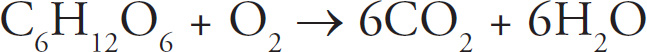
Germinating seeds respire and need to consume oxygen in order to continue to grow. Non-germinating seeds do not respire actively. In this lab, the amount of oxygen consumed by these types of seeds is measured using a respirometer. The experiment is also conducted at two temperatures, 25°C and 10°C, because seeds consume more oxygen at higher temperatures. You should know the following:
• Oxygen is consumed in cellular respiration.
• Germinating seeds have a higher respiratory rate than non-germinating seeds, which have a very low respiratory rate.
• Know how to design a study to determine the effect of temperature on cell respiration.
• Know the significance of a control using glass beads. A control is a condition held constant during an experiment. In this case, glass beads are used as a control because they will not consume any oxygen.
LAB 7: MITOSIS AND MEIOSIS
This lab highlights the differences between mitosis and meiosis. In this lab, slides of onion root tips are prepared to study plant mitosis. The important information and skills to review in this lab include the following:
• Mitosis produces two genetically identical cells, while meiosis produces haploid gametes.
• Cell division is highly regulated by checkpoints that depend, in part, on complexes of proteins called cyclins with other proteins called cyclin-dependent kinases. One such example is mitosis promoting factor (or MPF) that has its highest concentration during cell division and is thought to usher a cell into mitosis.
• Nondisjunction, or the failure of chromosomes to separate correctly, can lead to an incorrect number of chromosomes (too many or too few) in daughter cells.
• Know what each phase of the cell cycle looks like under a microscope. Chapter 8 contains diagrams of each phase.
In one section of this lab, the sexual life cycle of the fungus Sordaria fimicola is examined. Sexual reproduction in this fungus involves the fusion of two nuclei: a (+) strain and a (–) strain—to form a diploid zygote. This zygote immediately undergoes meiosis to produce asci, which contain eight haploid spores each.
• During meiosis, crossing-over can occur to increase genetic variation. If crossing-over, or recombination, has occurred, different genetic combinations will be observed in the offspring when compared with the parent strain.
• These offspring with new genetic combinations are called recombinants. By examining the numbers of recombinants with the total number of offspring, an estimate of the linkage map distance between two genes can be calculated using the following equation.
Map distance (in map units) = [(# recombinants) / (# total offspring)] × 100.
LAB 8: BIOTECHNOLOGY: BACTERIAL TRANSFORMATION
In this lab, the principles of genetic engineering are studied. Biotechnologists are able to insert genes into an organism’s DNA in order to introduce new traits or phenotypes, like inserting genes into a corn genome that help the crops ward off pests. This process is very complicated in higher plants and animals, but relatively simple in bacteria. You are responsible for knowing the ways in which bacteria can accept fragments of foreign DNA:
• Conjugation: the transfer of genetic material between bacteria via a pilus, a bridge between the two cells
• Transformation: when bacteria take up foreign genetic material from the environment
• Transduction: when a bacteriophage (a virus that infects bacteria) transfers genetic material from one bacteria to another
In addition, DNA can also be inserted into bacteria using plasmids, which are small, circular DNA fragments that can serve as a vector to incorporate genes into the host’s chromosome. Plasmids are key elements in genetic engineering. The concepts you need to know about plasmids include the following:
• One way to incorporate specific genes into a plasmid is to use restriction enzymes, which cut foreign DNA at specific sites, producing DNA fragments. A specific fragment can be mixed together with a plasmid, and this recombinant plasmid can then be taken up by E. coli.
• Plasmids can give a transformed cell selective advantage. For example, if a plasmid carries genes that confer resistance to an antibiotic like ampicillin, it can transfer these genes to the bacteria. These bacteria are then said to be transformed. That means if ampicillin is in the culture, only transformed cells will grow. This is a clever way scientists can find out which bacteria have taken up a plasmid.
• In order to make a bacteria cell take up a plasmid, you must (1) add CaCl2, (2) heat shock the cells, and (3) incubate them in order to allow the plasmid to cross the plasma membrane.
LAB 9: BIOTECHNOLOGY: RESTRICTION ENZYME ANALYSIS OF DNA
This laboratory introduces you to the technique of electrophoresis. This technique is used in genetic engineering to separate and identify DNA fragments. You need to know the steps of this lab technique for the AP exam:
1. DNA is cut with various restriction enzymes.
2. The DNA fragments are loaded into wells on an agarose gel.
3. As electricity runs through the gel, the fragments move according to their molecular weights. DNA is a negatively charged molecule; therefore, it will migrate toward the positive electrode. The longer the DNA fragment, the slower it moves through the gel.
4. The distance that each fragment has traveled is recorded.
Restriction mapping allows scientists to distinguish between the DNA of different individuals. Since a restriction enzyme will only cut a specific DNA sequence, it will cause a unique set of fragments for each individual called restriction fragments length polymorphisms, or RFLPs for short. This technology is used at crime scenes to help match DNA samples to suspects.
LAB 10: ENERGY DYNAMICS
This lab examines energy storage and transfer in ecosystems. Almost all organisms receive energy from the sun either directly or indirectly:
• Producers, or autotrophs, are organisms that can make their own food using energy captured from the sun. They convert this into chemical energy that is stored in high-energy molecules like glucose.
• Consumers, or heterotrophs, must obtain their energy from organic molecules in their environment. They can then take this energy to make the organic molecules they need to survive.
• Biomass is measure of the mass of living matter in an environment. It can be used to estimate the energy present in an environment.
LAB 11: TRANSPIRATION
This lab investigates the mechanisms of transpiration, the movement of water from a plant to the atmosphere through evaporation. What do you need to take away from this lab?
• The special properties of water that allow it to move through a plant from the roots to the leaves include polarity, hydrogen bonding, cohesion, and adhesion
• Know the vascular tissues that are involved in transport in plants. Xylem transports water from roots to leaves, while phloem transports sugars made by photosynthesis in the leaves down to the stem and roots.
• Stomata are small pores present in leaves that allow CO2 to enter for photosynthesis and are also a major place where water can exit a plant during transpiration.
LAB 12: FRUIT FLY BEHAVIOR
In this lab, fruit flies are given the choice between two environments using a choice chamber, which allows fruit flies to move freely between the two environments. Typically, fruit flies prefer an environment that provides either food or a place to reproduce. They also respond to light and gravity:
• Taxis is the innate movement of an organism based on some sort of stimulus. Movement toward a stimulus is called positive taxis, while movement away from a stimulus is negative taxis. In this lab, fruit flies exhibited a negative gravitaxis (or a movement opposite to the force of gravity) and positive phototaxis (a movement toward a light source).
LAB 13: ENZYME ACTIVITY
This lab demonstrates how an enzyme catalyzes a reaction and what can influence rates of catalysis. In this lab the enzyme peroxidase is used to catalyze the conversion of hydrogen peroxide to water and oxygen.
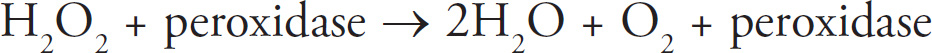
The following are the major concepts you need to understand for the AP exam:
• Enzymes are proteins that increase the rate of biological reactions. They accomplish this by lowering the activation energy of the reaction.
• Enzymes have active sites, which are pockets that the substrates (reactants) can enter that are specific to one substrate or set of substrates.
• Enzymes have optimal temperature and pH ranges at which they catalyze reactions. Enzyme concentration and substrate concentration can also influence the rates of reaction.
• If you’re asked to design an experiment to measure the effect of these four variables on enzyme activity, keep all the conditions constant except for the variable of interest. For example, to measure the effects of pH in an experiment, maintain the temperature, enzyme concentration, and substrate concentration constant as you change pH.
REFLECT
Respond to the following questions:
• Which topics from this chapter do you feel you have mastered?
• Which content topics from this chapter do you feel you need to study more before you can answer multiple-choice questions correctly?
• Which content topics from this chapter do you feel you need to study more before you can effectively compose a free response?
• Was there any content that you need to ask your teacher or another person about?
Chapter 16
Chapter Drill Answers and Explanations
CHAPTER 4 DRILL: CHEMISTRY OF LIFE
1. C Water has many unique properties that favor life, including (A), a high specific heat, (B), high surface tension and cohesive properties, and (D), high intermolecular forces due to hydrogen bonding. However, water is a very polar molecule and is an excellent solvent, making (C) inaccurate and the correct answer.
2. B The pH scale is logarithmically based, meaning that each difference of 1 on the log scale is indicative of a ten-fold difference in the hydrogen (H+) ion concentration. According to the question, the pH values of the cytoplasm of cells and gastric juices are approximately 7.4 and 2.5, respectively. Therefore, the pH values vary by 5, and the hydrogen (H+) ion concentrations much vary by 5 orders of ten (105), or 100,000-fold. Since lower pH values have more hydrogen ions, they are also more acidic.
3. C All amino acids share a carboxylic acid group, COOH, labeled (B), and an amino-group, NH2, labeled (D). They also share a hydrogen atom bound to the central carbon (A). Differences in amino acids are defined by variations in the fourth position called the R group, labeled (C).
4. B The hypothesis that life may have arisen from formation of complex molecules from the primordial “soup” of Earth is not supported by the absence of nucleic acids. All life is DNA-based, yet no nucleic acid molecules were detected. The presence of carbon molecules, amino acids, and sugars, which are common compounds and compose life, supports the hypothesis, so eliminate (A), (C), and (D). Choice (B) is correct.
5. C The amino acids cysteine and methionine contain the element sulfur (as indicated by the S in the amino acid structure shown). However, no sulfur-based compounds were included in the Miller-Urey experiment, so it was impossible to form these two amino acids under the conditions of the experiment.
6. B Silica is a mineral form of glass, is not a common component of life-forms, and is largely chemically inert. Since oxygen is already present in several compounds included in the experiment, the addition of this compound does not provide any additional elements or chemical substrates, which would permit generation of additional amino acids or synthesis of nucleic acids. The addition of sulfur compounds and phosphorus are necessary to generate some amino acids and all nucleic acids, which eliminates (A), (C), and (D).
7. D Hydrolysis adds water to a polymer to break the linkages between monomers. Therefore, (A), free water, will not result because water is being broken down in the reaction. Free water would result from the opposite reaction, for condensation reactions. Choice (B), adenine, results from the hydrolysis of DNA, as it is a monomer of DNA. Choice (C), cholesterol, is a steroid and therefore does not undergo hydrolysis. Dipeptides could result from the hydrolysis of proteins, which are composed of polypeptides.
8. B Phospholipids have a long fatty acid tail and a polar head group. The fatty acid tails associate in the membrane, and the phospholipid head group associates with water at the boundary of the membrane. Nucleotides, (A), are mostly polar and found in DNA and RNA. Water, (C), is polar and not found in plasma membranes. Amino acids, (D), are zwitterionic, that is, they have both a positive and a negative charge. Free amino acids are not found in the membrane.
9. C (CH2O)3 = C18H36O18, as in (A), but the dehydration reactions to produce two glycosidic bonds between the three monosaccharides would remove two H2O molecules, or four hydrogens and two oxygens. Therefore, the correct answer is (C).
10. A All isotopes of hydrogen will contain one proton, by definition. All atoms that only have one proton are identified as hydrogen. Tritium is an isotope that has one proton and two neutrons, for a total of three particles in its nucleus. Choice (A) is correct. Choice (B) is not correct because two protons would mean that the element is helium, not hydrogen. Choice (C) is not correct because the atomic number, one, will never change without changing the identity of the atom. Finally, (D) is not correct because radioactive atoms do not give off electrons.
CHAPTER 5 DRILL: CELLS
1. C Active transport is the movement of substances across a membrane against their concentration gradient through the use of energy (ATP). The sodium-potassium pump is a critical structure which uses ATP hydrolysis to move sodium and potassium ions against their respective concentration gradients. Diffusion of oxygen is an example of simple diffusion and can occur without the need of channels or pores. Water uses aquaporins to travel across the membrane, down its concentration gradient; therefore, it also does not require energy. Movement of sodium ions by a voltage-gated ion channel, (B), is an example of facilitated diffusion. Because the sodium ions are still undergoing diffusion from high to low concentrations without the need of ATP, this does not represent a form of active transport.
2. C Bacterial cells can be visualized using light microscopy. In fact, back in the seventeenth century, some of the earliest studies using primitive microscopes recorded the shape and organization of bacteria. Choices (A), (B), and (D) are incorrect because virus and cell organelle structures are too small to be observed using light microscopy and require electron microscopy.
3. D The cell wall is a structure that is present in bacteria but absent in animal cells. Consequently, this structure is targeted by several leading classes of antibiotics and would be an effective target of therapeutics against V. cholerae. Cytoplasm, plasma membrane, and ribosomes—(A), (B), and (C), respectively—are all structures that are present in animal cells.
4. C Based on the microscopy data, the organism has a cell wall and lacks mitochondria. The presence of a cell wall suggests that the organism is not a protozoan, so eliminate (B). The absence of mitochondria is indicative of prokaryotic structure because they lack organelles, eliminating (A) and (D). The most likely conclusion is that this organism is a bacterial species.
5. 700 The graphed line of chlorophyll a has its highest peak at 700 nm.
6. B Although many of the mentioned organelles work closely together, the only choice that can be correct is (B). Ribosomes translate and manufacture proteins, often on the rough endoplasmic reticulum. Those proteins are then packaged in the Golgi bodies into vesicles, which are then fused with the plasma membrane. Choice (A) is incorrect because the nuclear envelope and nucleolus do not interact with vacuoles; they interact with the centrioles only during mitosis. Choice (C) is incorrect because ribosomes don’t often interact with mitochondria, and lysosomes only interact with mitochondria and chloroplasts when they degrade those organelles. Choice (D) is incorrect because the nucleolus doesn’t interact with the smooth endoplasmic reticulum, lysosomes, or centrioles.
7. C Estrogen is said to be lipid-soluble. This means it can slip through the membrane and bind to an intracellular receptor. A noncompetitive inhibitor would bind to an estrogen receptor, reducing the effectiveness of this binding. Choice (A) is incorrect because estrogen binds to intracellular receptors, not receptors at the plasma membrane. Choice (B) is incorrect because testosterone and estrogen have different effects, but testosterone would not change estrogen effectiveness. Choice (D) is also incorrect because wiping out the ovaries would eliminate, rather than reduce, estrogen levels.
8. B You would need a cell that makes and secretes a lot of protein or hormones since the ER moves things out of the cell. Bacteria, (C), do not have organelles. Choice (D), blood cells, is not the best answer because leukocytes do not secrete large amounts of hormones or proteins. Choice (A), neurons, is a potential answer, but (B) is the best answer because the pancreas is a gland that secretes copious amounts of protein and hormones, such as insulin.
9. A Choice (A) is correct because contractile vacuoles expel water that accumulates in cells in a hypotonic environment. The strategy described in (B) is wrong because it would increase water intake in Paramecium. Choice (C) is true for Paramecium but does not help with water gain. Choice (D) would help it find even more dangerously hypotonic environments.
10. B ATP is consumed by the Na+K+ pump, so (B) is correct. The sodium potassium pump actively pumps both ions. A cotransporter requires one to be passive and one to be active. Choice (A) is not true. ATP is hydrolyzed and not produced. Choice (C) is not true. Choice (D) is incorrect since ATP is hydrolyzed.
CHAPTER 6 DRILL: CELLULAR ENERGETICS
1. B The Krebs cycle primarily occurs in the matrix of the mitochondria. The inner membrane and the intermembrane space—(A) and (C), respectively—are used in oxidative phosphorylation.
2. D Inhibitor Y is binding at a site outside of the active site and is inducing a conformational change in the enzyme structure. By binding outside of the active site, it must be an allosteric inhibitor, eliminating (A) and (C). Because the inhibitor is binding outside of the active site, it is not competing with the substrate for binding, so it is considered a non-competitive inhibitor.
3. B Enzymes are biological catalysts, which lower the activation energy (the energy threshold that must be met to proceed from reactant to product). The reaction coordinate diagram must reflect a decrease in the activation energy, eliminating (C). Furthermore, the enzyme does not alter the energy of the reactants or products, eliminating (A) and (D).
4. B Based on the pathway provided, consumption of one glucose and two ATP results in production of four ATP. In other words, each glucose results in a net gain of two ATP. Therefore, two glucose molecules would result in a net gain of four ATP.
5. B In fermentation, pyruvic acid is converted into either ethanol or lactic acid. During this process, NADH is recycled into NAD+.
6. B Glycolysis results in the production of ATP (energy), so it is considered an exergonic process.
7. B Thermophilic bacteria live in hot environments. Also, a DNA polymerase replicates DNA. Therefore, in the PCR technique, the stage in which DNA is elongated by a DNA polymerase is part three of the cycle, at 72°C. Therefore, bacteria growing in hydrothermal vents between 70–75°C is the answer. The conditions described in (A) and (D) do not reflect the data, which focuses on temperature. Choice (C) describes hot springs, but the temperature does not reflect the enzyme activity most likely for Taq polymerase.
8. D The free energy does not change between catalyzed and uncatalyzed reactions; therefore, (A) is incorrect. Choice (B) is a correct statement, but it does not answer the question. Choice (C) is incorrect because enzymes catalyze reactions.
9. B ATP production will increase in the treated mitochondria because the low pH provides more H+ ions in the solution. Also, oxygen provides a terminal electron acceptor for oxidative phosphorylation.
10. D Choices (A) and (B) are clearly wrong, as they defy physics. Choice (C) is not correct because the anabolic and catabolic reactions are not necessarily equal nor are they direct opposites. Choice (D) is the correct answer because it explains the increase in entropy. Even though organisms build and develop as ordered systems, heat is lost continuously. Additionally, organisms exhale gases and produce waste products that balance the effect of order.
CHAPTER 7 DRILL: MOLECULAR BIOLOGY
1. B Okazaki fragments are generated during DNA replication when the DNA polymerase must create short DNA segments due to its requirement for 5’ to 3’ polymerization. Since the newly discovered yeast cell has 3’ to 5’ activity, there would be no lagging strand and likely no Okazaki fragments.
2. C Since the gene is much shorter than expected, a stop codon must have been introduced by mutagenesis. This is an example of a nonsense mutation.
3. D The order for DNA replication is helicase, RNA primase, DNA polymerase, and ligase.
4. C If an mRNA codon is UAC, the complementary segment on a tRNA anticodon is AUG.
5. D During post-translational modification, the polypeptide undergoes a conformational change. Choices (A), intron excision, and (B), poly(A) addition, are examples of post-transcriptional modifications. Formation of peptide bonds occurs during translation, not afterwards.
6. B If twenty-one nucleotides compose a sequence and three nucleotides compose each codon, there would be seven codons and thus a maximum of seven amino acids.
7. A Choice (B) is incorrect because bacteria make membrane proteins that reside on the plasma membrane. Choice (C) is also incorrect because bacteria use transcription factors. Choice (D) is possible, but (A) is the best answer.
8. C Choice (A) is likely incorrect. Choice (B) would be deleterious for the cell. Choice (D) is also incorrect, as DNA replication occurs only during or preceding binary fission.
9. B Viruses do not have their own ribosomes, flagellum, or independent metabolism, so (A), (C), and (D) are incorrect. The only possible answer is (B).
10. C Transformation occurs when bacteria take up DNA from their surroundings. The pathogenic bacteria did not come alive again, as suggested by (A). Protein was not the transforming agent, so (B) is incorrect. Finally, (D) does not make sense, as genes cannot turn into other genes simply by being in a different cell context.
11. D Choice (D) describes the result of semiconservative replication, which is correct. After two rounds of semi-conservative replication, half the double helices will be half 15N and half 14N. The other helices will be all 14N.
12. C If there are DNA/RNA fragments, then helicase, (A), is present and has opened the strands. RNA primase, (D), is also present because RNA primers have been added. DNA polymerase, (B), must be present because there are small chunks and long chunks. If there was not polymerase, then there wouldn’t be small chunks because nothing new would be made. It is likely that DNA ligase (C) is missing and couldn’t attach the short Okazaki fragments and the longer leading strand fragments.
CHAPTER 8 DRILL: CELL REPRODUCTION
1. C During anaphase, the chromatids are separated by shortening of the spindle fibers. Chemically blocking the shortening of these fibers would arrest the cell in metaphase. The cells are arrested in metaphase as indicated by the alignment of the chromosomes in the center of the cell and their attachment to spindle fibers, eliminating (A) and (D). The chromosomes still seem to be attached to the fibers, so there doesn’t appear to be dissociation of the fibers, eliminating (B).
2. C The synthesis, or S phase, of the cell cycle represents the step in which the genetic material is duplicated. The only phase labeled in the experiment that represents an increase is phase B. Based on the time scale on the x-axis, this phase lasts approximately 30 minutes.
3. D Anaphase represents the cell division stage of the cell cycle and would be the phase that occurs right before the amount of genetic material should decrease. Phase D is the phase right before the genetic material would drop, so (D) is the correct answer.
4. 13 The organism has 13 chromosomes in gametic cells, which are haploid. Therefore, a diploid cell would normally have two copies of each chromosome or 26 making 13 homologous pairs. S phase replicates chromatids, but that won’t change the number of pairs.
5. D Choices (A) and (B) do not occur, but if they did, they would not impact the gametes. Choice (C) is not likely to occur because the mitotic spindles attach only to kinetochore protein complexes at the centromere. Therefore, (D) is the answer.
6. B Both (A) and (B) have sperm or ovum as the first type of cell. Because these are both gametes, they will both have half of the DNA of other types of cells. Choices (C) and (D) can thus be eliminated. Muscles and neurons are both terminally differentiated in G0 arrest. However, liver and taste buds potentially differ. Although liver cells can divide, taste buds divide at a higher rate, so they are most likely to be in G2 phase. Therefore, the answer is (B).
7. A Nondisjunction results in major changes to the genome, so (B) and (C) can be eliminated. Deletion of an enhancer, (D), could affect the gene expression and then phenotype. However, in (A), translocation may not produce any effect, as the same information exists in the genome.
CHAPTER 9 DRILL: HEREDITY
1. D The father and mother are both AB blood type. Since neither parent has a recessive allele, it is impossible for their child to be O blood type.
2. B When the phenotype associated with two traits is mixed, this is considered an example of incomplete dominance. In this case, neither red nor blue is dominant, and the resulting progeny exhibited a mixture of the traits (purple).
3. B Crossing the pea plant that is heterozygous for both traits (TtGg) with a plant that is recessive for both traits (ttgg) results in the following possible combinations, each of which should occur 25 percent of the time: TtGg (tall and green), Ttgg (tall and yellow), ttGg (short and green), and ttgg (short and yellow). Using the rules of probability, there is a 1/2 likelihood of it being tall and also a 1/2 likelihood of it being yellow. Multiply them together to get 1/4.
4. C Because the woman is a carrier, she must have one normal copy of the X chromosome and one diseased copy. Since the boy will receive an X chromosome from his mother, there is a 50 percent chance that he will receive a diseased copy. Because he doesn’t have a second X chromosome, he must have the disease if he receives the diseased X chromosome.
5. D They both must have a normal copy of the X chromosome. It is possible that the woman may be affected with hemophilia; however, that scenario is extremely unlikely because such a case would require two diseased copies of the X chromosome.
6. D Essentially, transmission of hemophilia to a girl born with Turner syndrome would be very similar to the conditions by which a boy would receive the disease. Both boys and girls with Turner syndrome only have one copy of the X chromosome. Therefore, they both would have a 50 percent chance of receiving the diseased copy of the X chromosome.
7. B Choice (A) is true, but it doesn’t explain why the parents are normal. Choice (B) is a better explanation. Carriers of recessive genetic diseases frequently do not have a recognizable phenotype because one normal allele provides enough functioning protein to avoid the ill effects of the disease allele. Choice (C) is also possible, but (B) is still the better answer. Finally, (D) is not correct because CLN3 is an autosomal gene.
8. C Choice (C) describes a testcross that will actually let the breeder know if the male is heterozygous. If any white spotted pups result from that cross, then the male would have contributed a spotted allele. Choice (B) describes a male that is homozygous for the spot allele, so it’s incorrect. Choice (A), stop breeding Speckle, is a way to remove spots from the line, but it will not help you identify males with the spot allele. Choice (D) also will not help you determine which black Labrador males have a spot allele.
9. A Choice (B) does not apply to this question. Choice (C) refers to a gene not following Mendel’s Law of Dominance. Choice (D) is a reason why traits would not follow independent assortment, but nondisjunction would cause many traits—whole chromosomes—to differ from the law. Choice (A)—genes are linked on the same chromosome—describes a normal reason for why certain traits would not follow Mendel’s law of independent assortment.
10. A A Choices (B) and (C) are unlikely to be true and can be eliminated. Choice (D) is true but does not answer the question. Choice (A) is the easiest and most probable explanation.
CHAPTER 10 DRILL: EVOLUTIONARY BIOLOGY
1. D Mammals and cephalopods developed similar eye structures independently due to similar selective pressures. This is an example of convergent evolution.
2. B The data provided show a transition toward one extreme (black) and away from another (white). This is an example of directional selection.
3. B Based on the data, the amount of white-bodied pepper moths decreased between 1802 and 1902, and the amount of black-bodied pepper moths increased during the same period.
4. B Longtail moths were included in the experiment as a control to compare the effects that are not associated with color.
5. B If the color of ash or soot produced by the Industrial Revolution were white or light gray, this would likely reverse the trend observed, applying additional selection against the black moths.
6. 0.42 The frequency can be calculated as shown below.
p + q = 1
p + (0.3) = 1
p = 0.7
2pq = heterozygotes = 2(0.7)(0.3) = 0.42
7. B The mole rats live in the same habitat, but over time there has been speciation as the two groups have chosen different plants as their diet. Therefore, (B) is correct. Choice (A) is incorrect because there is not a habitat barrier; the mole rats have access to each other. Choice (C) is incorrect because the mole rats are capable of breeding, they just do not. Choice (D) is incorrect because the mole rats do not interbreed.
8. D Choice (A) is incorrect because this is a small population, so genetic drift is likely. Choice (B) is incorrect because this situation does not describe convergent evolution, and the second part of the answer choice does not accurately describe the change of the population. Choice (D) is correct because the number of people that are homozygous recessives will likely become fewer as the red-head allele is mixed with the more dominant hair colors.
9. D Choice (D) is the answer because it is the only choice that does not predict evolution occurring on the island, thus supporting a claim for HW equilibrium. Choices (A), (B), and (C) all prevent a Hardy-Weinberg equilibrium and predict evolution. The population is small, mutations are inevitable, and humans usually do not mate randomly. This population will definitely evolve or undergo genetic drift.
10. A Choice (A) is correct because the trait is being selected based on female mating preferences. Longer, bigger tails indicate reproductive fitness. Peahens choose males who are healthy and strong enough to grow these big tails and will hopefully produce the best offspring. Choices (B), (C), and (D) are incorrect because they do not describe the female mating preferences.
CHAPTER 11: ANIMAL STRUCTURE AND FUNCTION
1. B Lymph nodes harbor large quantities of lymphocytes. During active infections or cancer, mass expansion of lymphocytes causes enlargement and swelling of lymph nodes. This is why doctors often feel the areas around lymph nodes nearest to where a suspected infection is.
2. C The placebo strain was included as a control for the experiment. Because no active antibodies should be produced to the placebo strain, the scientists would be able to compare antibodies elicited specifically to the vaccine strain.
3. B B-cells (or plasma cells) are responsible for creating antibodies. These are the primary cells associated with lasting memory to vaccinations.
4. A The most practical way to increase statistical significance is to increase the sample size, which is (A). More control variables will only ensure that they haven’t introduced bias. Repeating a third challenge would evaluate the length of potential protection, not the extent. Using a different statistical test would simply manipulate the data and not provide true significance.
5. C Choice (A) could work, but it would cause the entire immune system to be reduced, which could be harmful to the patient and is not a targeted response. Choice (B), removal of the thymus, is a treatment for this disease (The thymus is where the maturation of T-cells take place.), but it is incorrect because macrophages do not mature in the thymus. Choice (D) is incorrect because additional acetylcholine will not make a difference because the receptors are affected. Choice (C) is the best answer because removing acetylcholine autoantibodies could reduce the damage to the receptors.
6. B Thorns and spines are examples of innate defenses that plants have to deter herbivores. Choice (C), attractive scents, would have the opposite effect. Stomata allow gas exchange and are unrelated to the question. Long stems may be considered a defense only against certain types of threats. Therefore, (B) is the best answer.
7. A Choice (B) is incorrect because helper T-cells do not destroy pathogens. Choice (D) is incorrect, as the goal of the immune system is to prevent further infection. Choice (C) is incorrect because only memory B-cells remain, not plasma B-cells. Choice (A) is the correct answer because B-cells secrete antibodies specific to the pathogen in the complement system. This flags the pathogen for destruction.
8. B Choice (B) is the only logical answer, as the other three choices refer to different types of immune cells.
9. B The membrane potential depolarizes at –50 mV so that the voltage-gated sodium channels can open. Note that the sodium channels open and close more quickly than the potassium channels.
10. CThe blocking of voltage-gated sodium (Na+) channels by tetrodotoxin would prevent an action potential in cells and therefore the transmission of a neural response to the subsequent neurons.
11. D Choices (A) and (C) describe situations that are not directly related to ouabain. Choice (B) is incorrect, as the cell would slowly become more negative without the sodium potassium pump because potassium would leak out of the cell. Choice (D) is correct.
12. B Proteins are made primarily in the cell body of neurons. Dendrites receive postsynaptic signals. Axons and axon terminals transmit action potentials in the presynaptic neuron. Choice (B) is correct.
13. C In a neuromuscular junction, the neuron synapses with a muscle. The neuron releases acetylcholine as a neurotransmitter to tell the muscle to contract. Thus, acetylcholinesterase removes extra acetylcholine from the synapse after the transmission. Without this enzyme, the neurotransmitter will build up in the synapse, causing overstimulation of the acetylcholine receptors on the muscle and hyper-contraction of the muscles. Choices (A), (B), and (D) are incorrect because they describe the opposite of what would occur.
14. A By definition, the threshold is the point at which enough voltage-gated channels have opened to permit a switch in the polarity of the membrane and an action potential to perpetuate through the axon of the neuron. Choices (B), the release of neurotransmitters, and (C), peak depolarization, both occur after the threshold is met. Choice (D), sodium ions enter the cell during depolarization, is vague and not as strong as (A).
15. B Both Schwann cells and oligodendrocytes act to wrap around the axons of neurons to insulate the membrane depolarization and expedite the action potential. Choice (B) is correct.
16. B Choices (A), (C), and (D) are incorrect because testosterone and estradiol will easily diffuse across the membrane, as they are hydrophobic, steroid-type molecules. Receptors are not needed on the outside of the plasma membrane. Instead, receptors inside the cell will receive and transduce the signal. Choice (B) is the correct answer.
17. B Choice (B) is the correct answer because the receptor for vasopressin is in the G-protein–coupled receptor family. These receptors sit on the plasma membrane. Only cells with the vasopressin receptor will respond to the presence of vasopressin, even if other cells are exposed to the hormone. Therefore, (A) and (C) are incorrect. Choice (D) is also incorrect because vasopressin does not enter cells.
CHAPTER 12 DRILL: BEHAVIOR AND ECOLOGY
1. D The behavior displayed by the chimpanzee represents insight because the chimpanzee has figured out how to solve the problem without external influence or learning.
2. A Viruses would display reproductive strategies most similar to r-strategists because they aim to reproduce as fast as possible and create as many progeny as possible in order to increase their odds of transmission to other hosts.
3. C They would be considered secondary consumers because they consume the primary consumers (plankton) which consume algae (the producers).
4. C Since brook trout can tolerate pH values as low as 4.9 and do not appear to diminish, the pH of the creek must exceed 4.9, so we can eliminate (D). Since crayfish cannot tolerate pH levels lower than 5.4, the pH of the stream must have dropped lower than this value. Only (C) falls in this range.
5. C Only acid rain would directly explain why the pH would drop in the creek over the time period.
6. D Based on Table 1, the pH must exceed 6.1 for snails to be able to return to the creek ecosystem.
7. D Choices (A) and (B) refer to light and gravity responses of plants. Choice (C) is not an actual tropism—it is a made-up word. Thigmotropism is the term for how plants respond to touch. The correct answer is (D).
8. D If (D) is a correct statement, then simply cutting out beef and lamb from your diet can dramatically decrease your carbon footprint—even if you do not decide to eat vegan. Choices (A), (B), and (C) are all true statements with respect to reducing the carbon footprint of food.
9. B A community that has been destroyed and then rebuilds is a result of secondary succession. Choice (A) would be the answer only if there were no life present before the new growth. Choice (C) is true only of a mature forest. Choice (D) is incorrect, as the species taking over the field is native to the rainforests on the periphery of the field. Nothing in the question suggests invasive species. Therefore, (B) is correct.
10. A If the intestines are occupied with beneficial bacteria, there will not be any room available for pathogenic bacteria to grow by density-dependent limitations on the population. Choice (B) is not accurate because pathogenic bacteria do not require a niche prepared for them by probiotic bacteria. Choice (C) is true, but it does not answer the question. Choice (D) is also incorrect, leaving (A) as your answer.
CHAPTER 13 DRILL: QUANTITATIVE SKILLS AND BIOSTATISTICS
1. A In order to answer this question, you must first calculate how much weight each subject lost and then divide by the number of subjects (in this case, five).
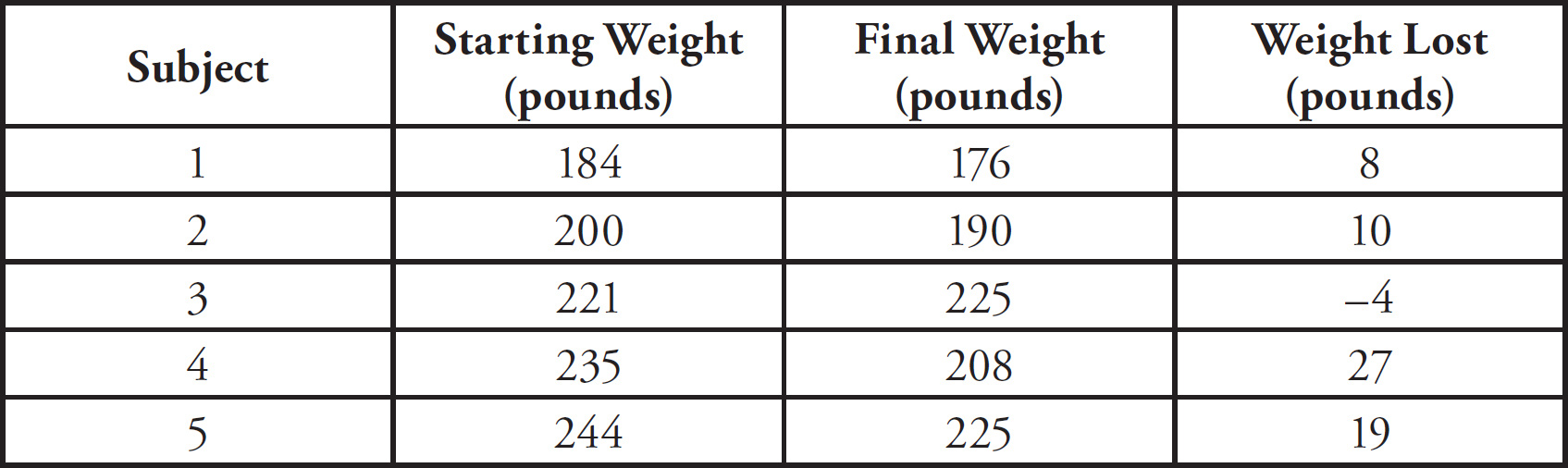
Note that subject 3 gained four pounds. Total weight lost is 60 pounds (remember to subtract 4 pounds for subject 3, not add), divided by 5 subjects is 12 pounds. The average weight lost is 12 pounds ((A) is correct).
2. C Plant 6 is 68 inches. All the answer choices list median values smaller than this, so the answer must start with “Yes” (eliminate (B) and (D)). In order to determine the median height for all six plants, their heights must first be organized in ascending order: 61, 66, 67, 68, 70, 72. The middle two numbers are 67 and 68; when averaged, this produces a median of 67.5 inches ((C) is correct). Notice that if all the data points are whole numbers, the median value must either end in .0 or .5, so you can quickly eliminate (A) and (B) in this question. In fact, (A) and (B) are giving the value for mean, not median.
3. D The test scores ordered from smallest to largest are:
32, 65, 66, 67, 68, 68, 69, 70, 71, 72, 73, 75, 75, 75, 78, 82
The most frequently recurring number in the set above is 75, so this is the mode (eliminate (A) and (B)). The smallest number is 32 and the largest is 82. The range is the difference between the two, or 50 ((D) is correct).
4. D This question is testing the product rule.
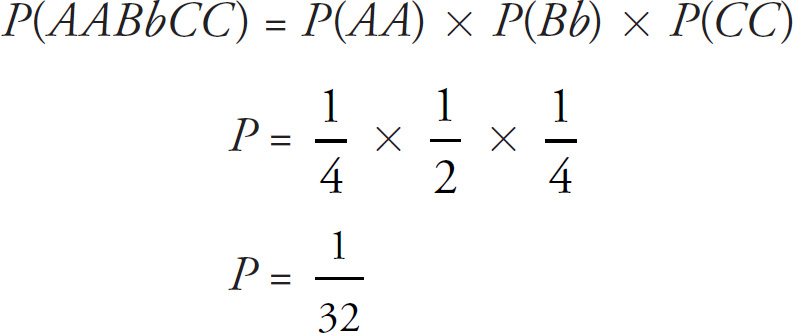
5. A This question is testing the sum rule, but you also need to use the product rule.
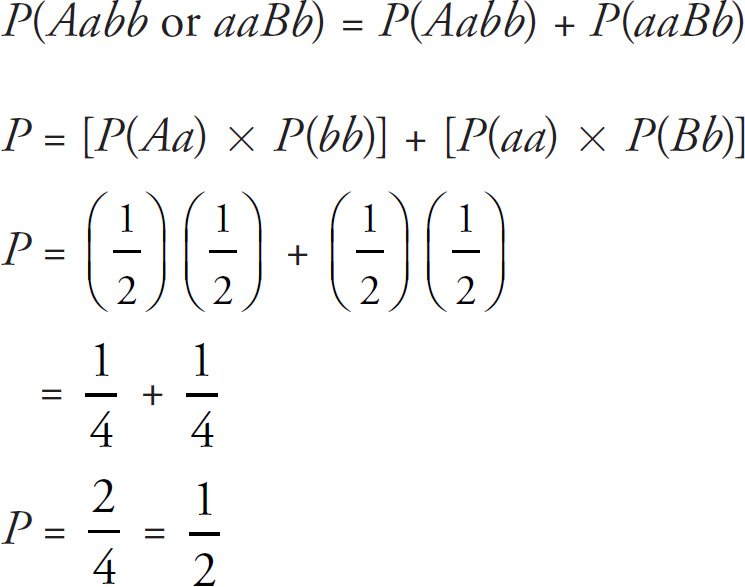
6. C In chi-square tests, a calculated x2 value is compared to a critical value (CV) from a chi-square table, like the one on the AP Biology Equations and Formulas sheet. If x2 < CV, you accept the null hypothesis (Ho). If x2 > CV, you reject Ho. Our Ho is that the observed and expected data match and that the experimental plants have a 3:1 ratio of yellow to green plants. Based on this, the only possible answer choices are (B) and (C); the other choices mix up how x2 values and critical values are compared (eliminate (A) and (D)).
One hundred plants were studied, and we expect three-fourths of them to be yellow (75 plants) and one-fourth of them to be green (25 plants). Next, we compare expected (E) and observed (O) data and calculate a x2 value.
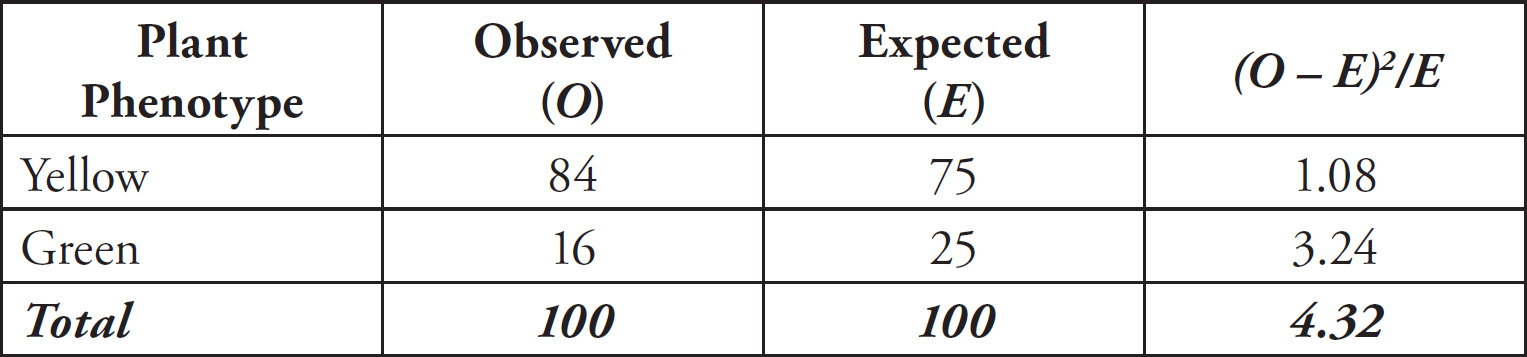
Because two possibilities are being compared (green and yellow), the degrees of freedom in this test = 2 – 1 = 1. Using p = 0.05, the critical value is 3.84 (which you don’t need to look up because it is listed in all answer choices). Since x2 > CV, you reject Ho. The observed data do not match the expected data, and the correct answer is (C).
7. B The first thing you need to do is sort out the alleles for each gene. Since the F1 generation had long phenotypes, you know these must be dominant to the short phenotypes. Let’s use:
L = long leaves
l = short leaves
S = long shoots
s = stubby shoots
The parental cross must have been LLSS × llss, and all F1 plants were LlSs. Our null hypothesis (Ho) is that the two genes are segregating independently. This means the expected ratio of F2 phenotypes will be 9LS:3Ls:3lS:1ls. Since 1000 F2 plants were generated, we expect:
P(long leaves, long shoots) = P(LS phenotype) = 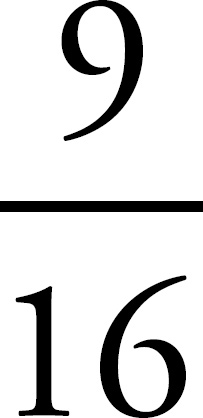 × 1000 = 562.5
× 1000 = 562.5
P(long leaves, stubby shoots) = P(Ls phenotype) = 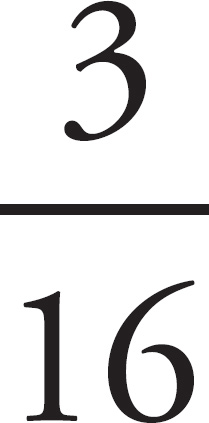 × 1000 = 187.5
× 1000 = 187.5
P(short leaves, long shoots) = P(lS phenotype) = 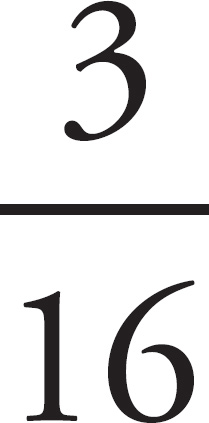 × 1000 = 187.5
× 1000 = 187.5
P(short leaves, stubby shoots) = P(ls>phenotype) = 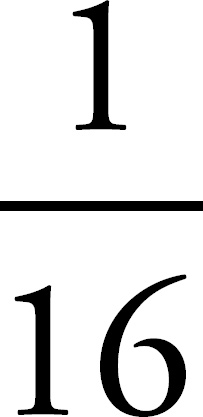 × 1000 = 62.5
× 1000 = 62.5
Next, generate a chart:
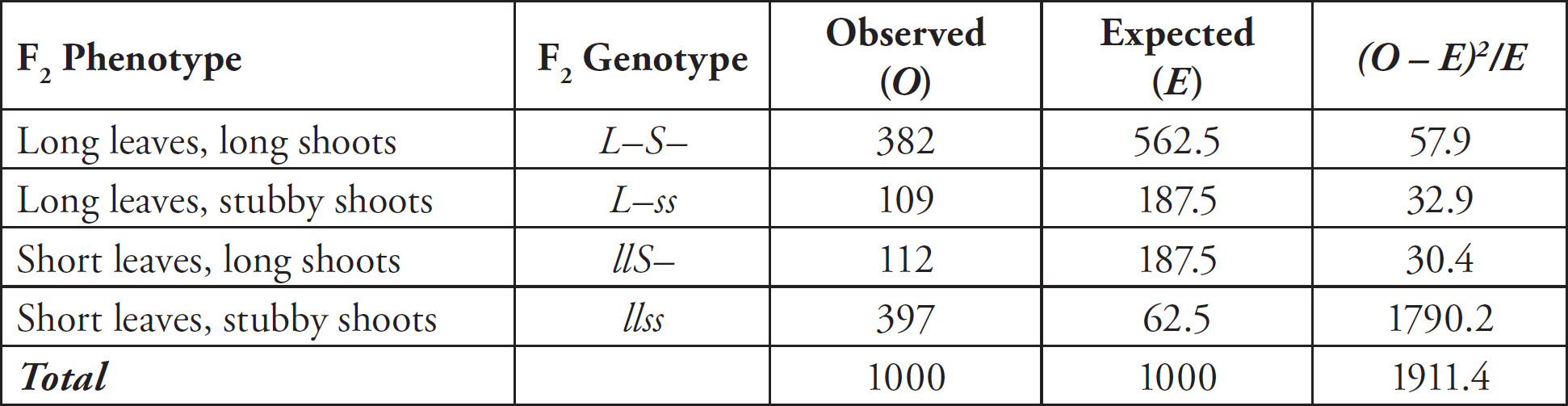
x2 = 1911.4 and degrees of freedom = # of possibilities – 1 = 4 – 1 = 3 (eliminate (C) and (D)). Using p = 0.05, we get a critical value of 7.82. Since x2 > CV, we reject Ho. These two genes are not segregating independently (eliminate (A); (B) is correct).
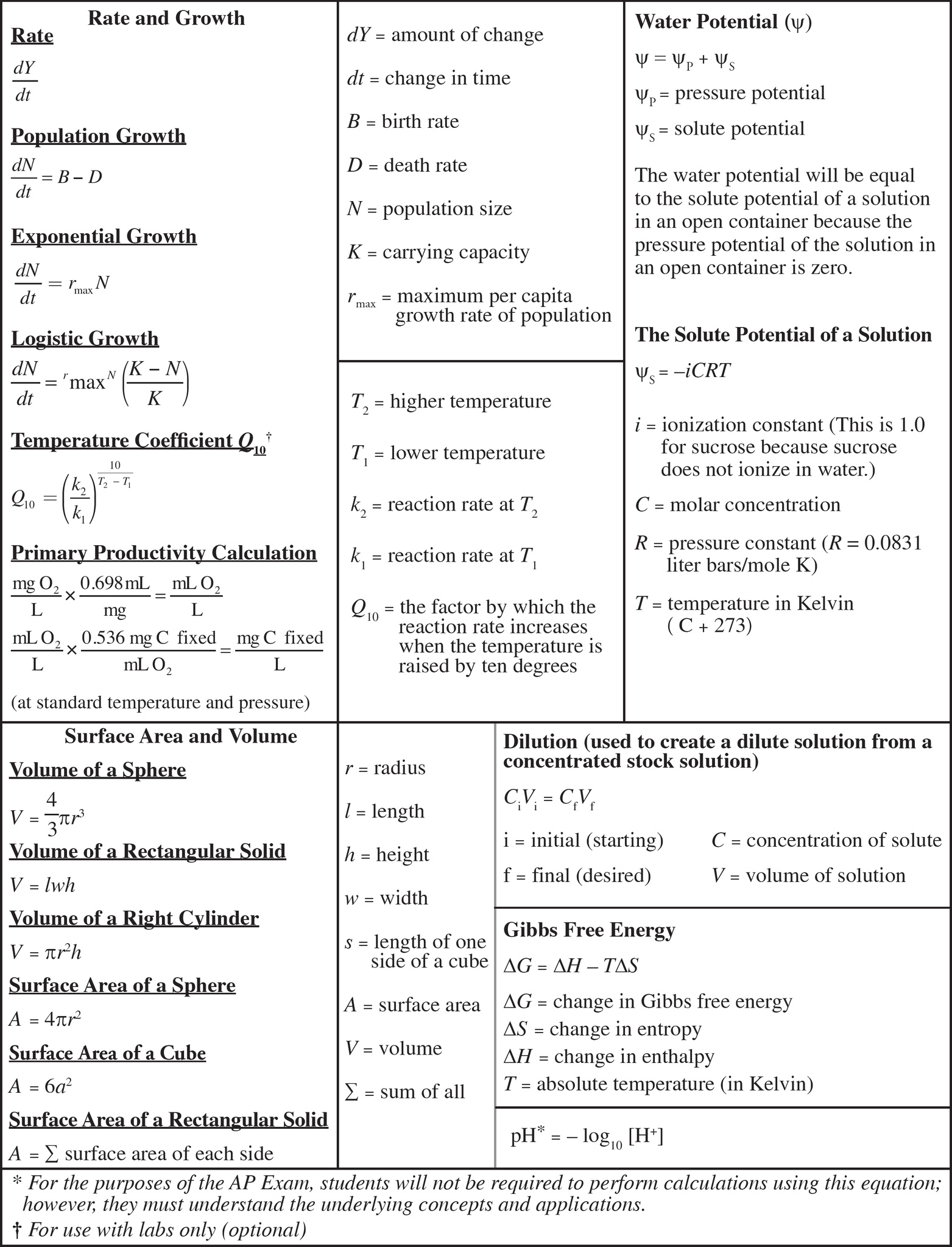
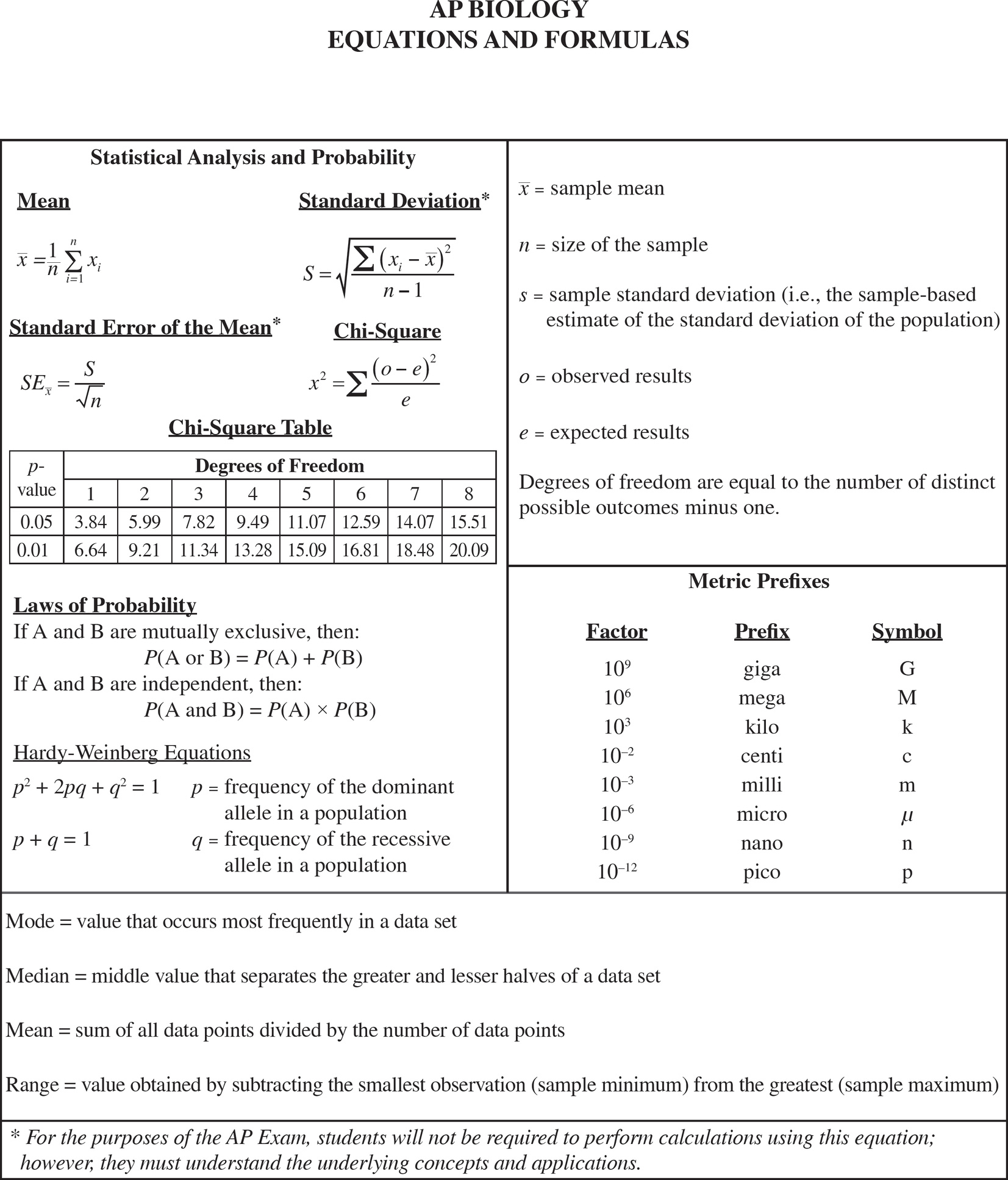

Part VI
Additional Practice Tests
Practice Test 2
Practice Test 2: Answers and Explanations
Practice Test 3
Practice Test 3: Answers and Explanations
Practice Test 4
Practice Test 4: Answers and Explanations
Practice Test 2
Click here to download a PDF of Practice Test 2
The Exam
AP® Biology Exam
SECTION I: Multiple-Choice Questions
DO NOT OPEN THIS BOOKLET UNTIL YOU ARE TOLD TO DO SO.
At a Glance
Total Time
1 hour and 30 minutes
Number of Questions
69
Percent of Total Score
50%
Writing Instrument
Pencil required
Instructions
Section I of this examination contains 69 multiple-choice questions. These are broken down into Part A (63 multiple-choice questions) and Part B (6 grid-in questions).
Indicate all of your answers to the multiple-choice questions on the answer sheet. No credit will be given for anything written in this exam booklet, but you may use the booklet for notes or scratch work. After you have decided which of the suggested answers is best, completely fill in the corresponding oval on the answer sheet. Give only one answer to each question. If you change an answer, be sure that the previous mark is erased completely. Here is a sample question and answer.
Sample Question
Chicago is a
(A) state
(B) city
(C) country
(D) continent
Sample Answer

Use your time effectively, working as quickly as you can without losing accuracy. Do not spend too much time on any one question. Go on to other questions and come back to the ones you have not answered if you have time. It is not expected that everyone will know the answers to all the multiple-choice questions.
About Guessing
Many candidates wonder whether or not to guess the answers to questions about which they are not certain. Multiple choice scores are based on the number of questions answered correctly. Points are not deducted for incorrect answers, and no points are awarded for unanswered questions. Because points are not deducted for incorrect answers, you are encouraged to answer all multiple-choice questions. On any questions you do not know the answer to, you should eliminate as many choices as you can, and then select the best answer among the remaining choices.
BIOLOGY
SECTION I
69 Questions
Time—90 minutes
Directions: Each of the questions or incomplete statements below is followed by four suggested answers or completions. Select the one that is best in each case and then fill in the corresponding oval on the answer sheet.
1. In general, animal cells differ from plant cells in that animal cells have
(A) a cell wall made of cellulose
(B) lysosomes
(C) large vacuoles that store water
(D) centrioles within centrosomes
2. A cell from the leaf of the aquatic plant Elodea was soaked in a 15 percent sugar solution, and its contents soon separated from the cell wall and formed a mass in the center of the cell. All of the following statements are true about this event EXCEPT
(A) the vacuole lost water and became smaller
(B) the space between the cell wall and the cell membrane expanded
(C) the large vacuole contained a solution with much lower water potential than that of the sugar solution
(D) the concentration of solutes in the extracellular environment is hypertonic with respect to the cell’s interior
3. A chemical agent is found to denature acetylcholinesterase in the synaptic cleft. What effect will this agent have on acetylcholine?
(A) Acetylcholine will not be released from the presynaptic membrane.
(B) Acetylcholine will not bind to receptor proteins on the postsynaptic membrane.
(C) Acetylcholine will not diffuse across the cleft to the postsynaptic membrane.
(D) Acetylcholine will not be degraded in the synaptic cleft.
4. The base composition of DNA varies from one species to another. Which of the following ratios would you expect to remain constant in the DNA?
(A) Cytosine : Adenine
(B) Pyrimidine : Purine
(C) Adenine : Guanine
(D) Guanine : Deoxyribose
Questions 5–6 refer to the following passage.
Consider the following pathway of reactions catalyzed by enzymes (shown in numbers):
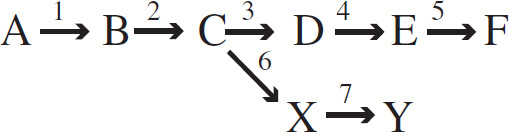
5. Which of the following situations represents feedback inhibition?
(A) Protein D activating enzyme 4
(B) Protein B stimulating enzyme 1
(C) Protein 7 inhibiting enzyme C
(D) Protein X inhibiting enzyme 2
6. An increase in substance F leads to the inhibition of enzyme 3. All of the following are results of the process EXCEPT
(A) an increase in substance X
(B) increased activity of enzyme 6
(C) decreased activity of enzyme 4
(D) increased activity of enzyme 5
7. If a competitive inhibitor bound to enzyme 1, which of the following would be decreased?
I. Protein A
II. Protein C
III. Protein X
(A) I only
(B) II only
(C) II and III
(D) I, II, and III
Questions 8–11 refer to the following passage.
The affinity of hemoglobin for oxygen is reduced by many factors, including low pH and high CO2. The graph below shows the different dissociation curves that maternal (normal) hemoglobin and fetal hemoglobin have.
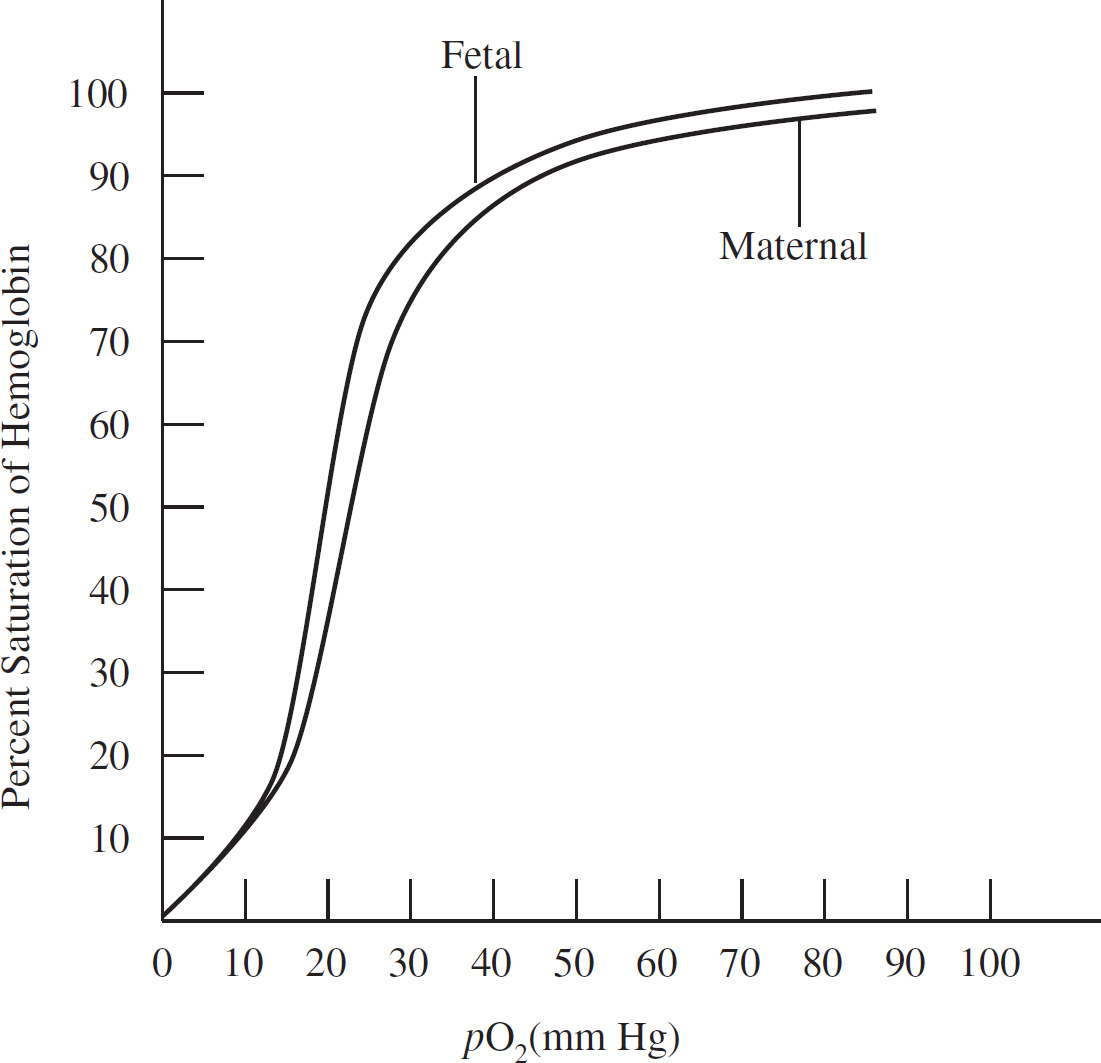
8. Based on the graph, it can be concluded that
(A) fetal hemoglobin surrenders O2 more readily than maternal hemoglobin
(B) the dissociation curve of fetal hemoglobin is to the right of maternal hemoglobin
(C) fetal hemoglobin has a higher affinity for O2 than does maternal hemoglobin
(D) fetal and maternal hemoglobin differ in structure
9. Which of the following processes would likely shift the normal dissociation curve to the right?
(A) Photosynthesis
(B) Respiration
(C) Fermentation
(D) Mitosis
10. Hemoglobin’s affinity for O2
(A) decreases as blood pH decreases
(B) increases as H+ concentration increases
(C) increases as blood pH decreases
(D) decreases as OH− concentration increases
11. How much pO2 would it take in an extremely CO2-rich environment to saturate hemoglobin 90 percent?
(A) 15
(B) 30
(C) 45
(D) 60
12. All of the following are differences between prokaryotes and eukaryotes EXCEPT
(A) eukaryotes have linear chromosomes, while prokaryotes have circular chromosomes
(B) eukaryotes possess double stranded DNA, while prokaryotes possess single stranded DNA
(C) eukaryotes process their mRNA, while in prokaryotes, transcription and translation occur simultaneously
(D) eukaryotes contain membrane-bound organelles, while prokaryotes do not
13. In minks, the gene for brown fur (B) is dominant over the gene for silver fur (b). Which set of genotypes represents a cross that could produce offspring with silver fur from parents that both have brown fur?
(A) BB × BB
(B) BB × Bb
(C) Bb × Bb
(D) Bb × bb
14. All viruses contain at least these two principal components:
(A) DNA and proteins
(B) nucleic acids and a capsid
(C) DNA and a cell membrane
(D) RNA and a cell wall
15. All of the following are examples of hydrolysis EXCEPT
(A) conversion of fats to fatty acids and glycerol
(B) conversion of proteins to amino acids
(C) conversion of starch to simple sugars
(D) conversion of pyruvic acid to acetyl-CoA
16. In cells, which of the following can catalyze reactions involving hydrogen peroxide, provide cellular energy, and make proteins, in that order?
(A) Peroxisomes, mitochondria, and ribosomes
(B) Peroxisomes, mitochondria, and lysosomes
(C) Peroxisomes, mitochondria, and Golgi apparatus
(D) Lysosomes, chloroplasts, and ribosomes
Questions 17 and 18 refer to the following passage.
The Loop of Henle is a structure within each of the million nephrons within a kidney. As shown in the figure, the two sides have different permeabilities, and there is differential movement across each membrane. The Loop acts as a counter-current multiplier that makes the medulla of the kidney very osmotic. The longer the loop, the higher and more powerful the osmolarity gradient that is created. The gradient is required for the reclamation of water from the urine collecting duct. On the right side of the figure is the urine collecting duct. If the body needs to retain water, and anti-diuretic hormone makes this region permeable to water, and the osmotic pull of the medulla reclaims the water, out of the collecting duct, which makes the urine more concentrated.
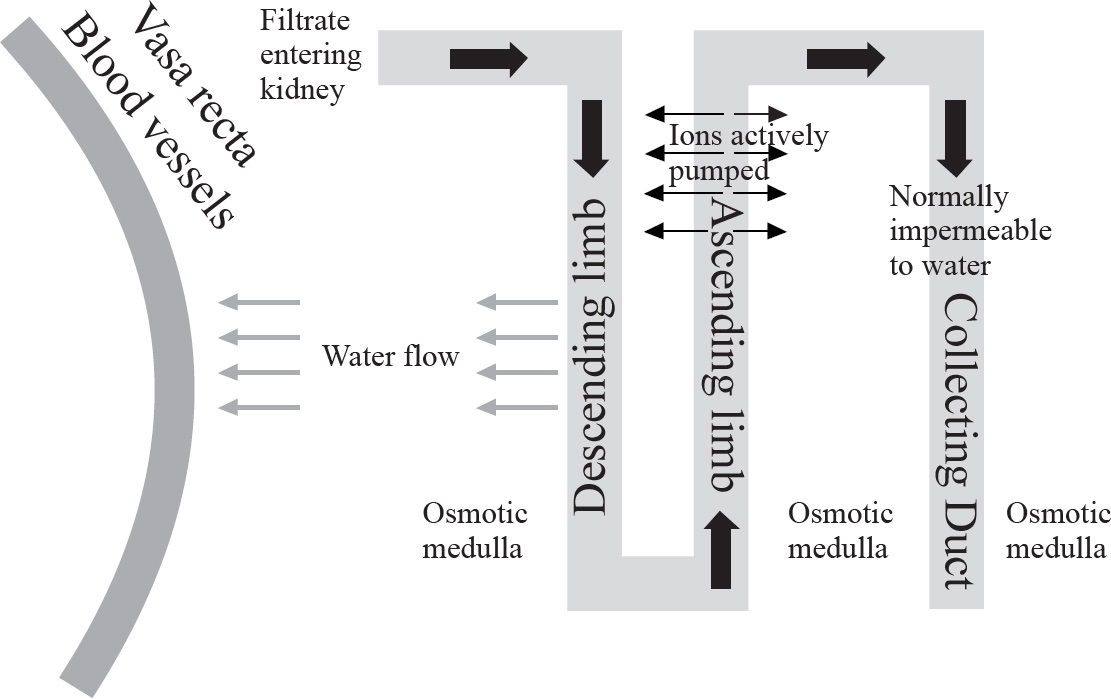
17. Which of the following statements correctly describes the state of things near the top of the descending limb?
(A) The fluid within the descending limb is hypotonic to fluid in the space surrounding the tubule.
(B) The blood within the vasa recta is hypotonic to the filtrate within the descending limb.
(C) The fluid in the area surrounding the tubule is hypertonic to the blood in the vasa recta.
(D) The water in the area surrounding the tubule has a higher water potential than the water in the descending tubule.
18. What type of transport is occurring when water flows out of the descending tubule?
(A) Simple diffusion
(B) Facilitated diffusion
(C) Active transport
(D) Secondary active transport
19. If an inhibitor of anti-diuretic hormone, such as caffeine, was ingested, what would be the result?
(A) Aquaporins would appear in the collecting duct.
(B) Aquaporins would increase in number in the collecting duct.
(C) It would block aquaporins like a competitive inhibitor.
(D) Aquaporins would not appear in the collecting duct.
20. The ability to reclaim water from the collecting duct is directly related to the osmotic pull of the medulla. Kangaroo rats are know to produce extremely concentrated urine. Compared to a human, the nephrons in kangaroo rats must have
(A) thick walls that are impermeable to water
(B) shorter Loops of Henle
(C) longer Loops of Henle
(D) shorter collecting ducts
Questions 21 and 22 refer to the following graph.
The graph below shows the growth curve of a bacterial culture.

21. Which of the following represents the carrying capacity of the environment?
(A) 2
(B) 3
(C) 4
(D) 5
22. Which of the following shows the exponential growth curve of the population?
(A) 1
(B) 2
(C) 3
(D) 4
23. In plants, the tendency of climbing vines to twine their tendrils around a trellis is called
(A) thigmotropism
(B) hydrotropism
(C) phototropism
(D) geotropism
24. Females with Turner’s syndrome have a high incidence of hemophilia, a recessive, X-linked trait. Based on this information, it can be inferred that females with this condition
(A) have an extra X chromosome
(B) have an extra Y chromosome
(C) lack an X chromosome
(D) have red blood cells that clump
25. When a retrovirus inserted its DNA into the middle of a bacterial gene, it altered the normal reading frame by one base pair. This type of mutation is called
(A) duplication
(B) translocation
(C) inversion
(D) frameshift mutation
26. The principal inorganic compound found in living things is
(A) carbon
(B) oxygen
(C) water
(D) glucose
27. Metafemale syndrome, a disorder in which a female has an extra X chromosome, is the result of nondisjunction. This failure in oogenesis would first be apparent when
(A) the homologous chromosomes are lined up in the middle
(B) the sister chromatids are lined up in the middle
(C) the nuclear envelope breaks down before meiosis
(D) the homologous chromosomes are pulling apart
28. All of the following are modes of asexual reproduction EXCEPT
(A) sporulation
(B) fission
(C) budding
(D) meiosis
29. Invertebrate immune systems possess which of the following?
(A) Killer T-cells
(B) Phagocytes
(C) B-cells
(D) Helper T-cells
Questions 30–32 refer to the following passage.
The following are important pieces of replication and transcription machinery:

30. Which of the following figures would be present without helicase?
(A) 
(B) 
(C) 
(D) 
31. What situation might have occurred to produce the following situation?
(A) Stalled DNA replication
(B) Initiation of transcription
(C) Repression of transcription
(D) Crossing-over in meiosis
32. Put these four situations in the correct order:
I. 
II. 
III. 
IV. 
(A) I, III, IV, II
(B) I, IV, II, III
(C) IV, I, III, II
(D) I, III, II, IV
33. Photosynthesis requires
(A) glucose, light, CO2
(B) light, CO2, water
(C) water, soil, O2
(D) O2, water, light
34. Which of following conclusions was made by Avery, MacLeod, and McCarty?
(A) The double helix is the structure of a DNA molecule
(B) DNA is the hereditary material
(C) DNA replication is semi-conservative
(D) DNA polymerase only adds bases to the 3′ side of DNA
35. Which of the following processes occur in the cytoplasm of an eukaryotic cell?
I. DNA replication
II. Transcription
III. Translation
(A) I only
(B) III only
(C) II and III only
(D) I, II, and III
36. Crossing-over during meiosis permits scientists to determine
(A) the chance for variation in zygotes
(B) the rate of mutations
(C) the distance between genes on a chromosome
(D) which traits are dominant or recessive
37. An animal cell that is permeable to water but not salts has an internal NaCl concentration of 10%. If placed in freshwater the cell will
(A) plasmolyze
(B) swell and eventually lyse
(C) endocytose water into a large central vacuole
(D) shrivel
38. Three distinct bird species, flicker, woodpecker, and elf owl, all inhabit a large cactus, Cereus giganteus, in the desert of Arizona. Since competition among these birds rarely occurs, the most likely explanation for this phenomenon is that these birds
(A) have a short supply of resources
(B) have different ecological niches
(C) do not live together long
(D) are unable to breed
39. Lampreys attach to the skin of lake trout and absorb nutrients from its body. This relationship is an example of
(A) commensalism
(B) parasitism
(C) mutualism
(D) gravitropism
40. The nucleotide sequence of a template DNA molecule is 5′-C-A-T-3′. A mRNA molecule with a complementary codon is transcribed from the DNA. What would the sequence of the anticodon that binds to this mRNA be?
(A) 5′-G-T-A-3′
(B) 5′-G-U-A-3′
(C) 5′-C-A-U-3′
(D) 5-′U-A-C-3′
41. Viruses are considered an exception to the cell theory because they
(A) are not independent organisms
(B) have only a few genes
(C) move about via their tails
(D) have evolved from ancestral protists
Questions 42–44 refer to the following passage.
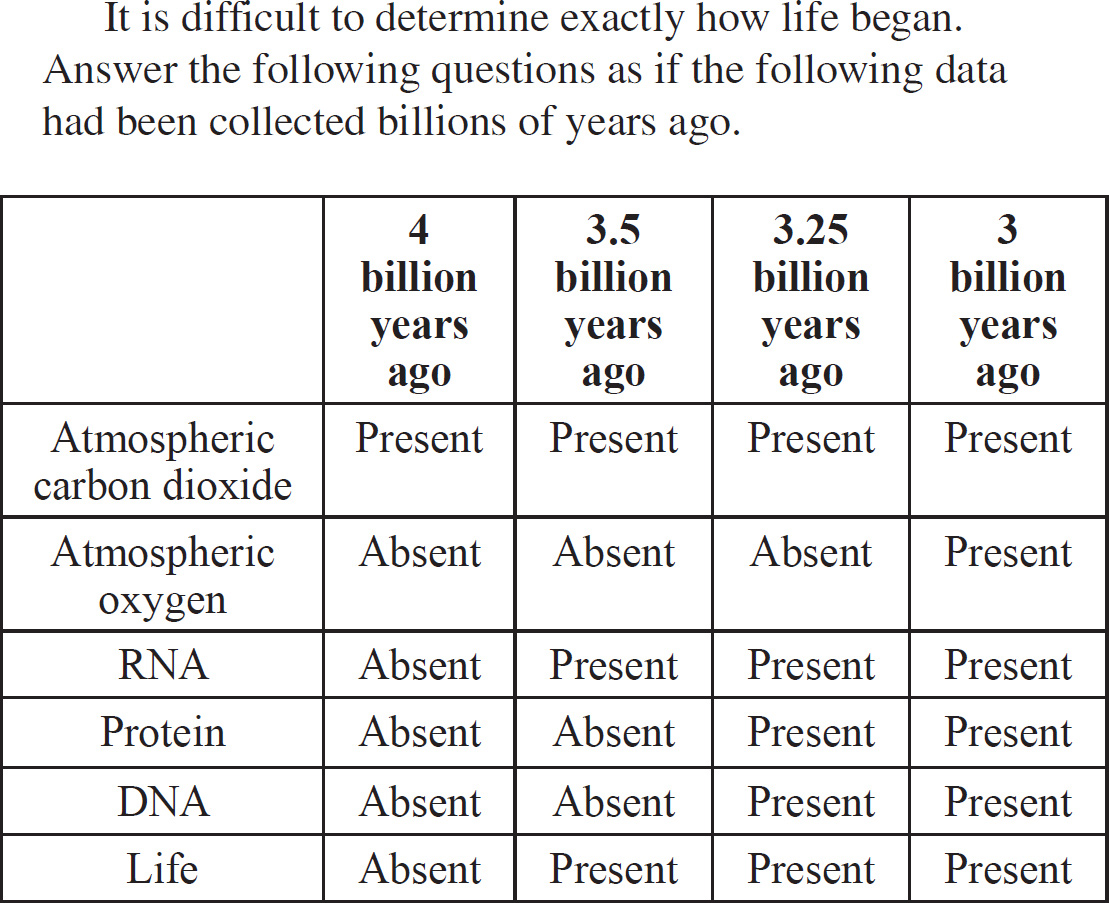
42. Which best describes the origin of life according to the data?
(A) Life required an environment with atmospheric oxygen and any type of nucleic acids
(B) Life required an environment with atmospheric carbon dioxide, but not atmospheric oxygen
(C) Life required an environment with self-replicating nucleic acids that can take on many shapes
(D) Life required an environment with nucleic acids and proteins, but not atmospheric oxygen
43. When is the earliest that functional ribosomes could have been found?
(A) Between 4 billion and 3.5 billion years ago
(B) Between 3.5 billion and 3.25 billion years ago
(C) Between 3.25 billion and 3 billion years ago
(D) Between 3 billion years ago and the present
44. Photosynthesis likely began ______ billion years ago when the first __________ appeared.
(A) 4.5; autotrophs
(B) 3.2; autotrophs
(C) 3.5; heterotrophs
(D) 3.2; heterotroph
45. The sequence of amino acids in hemoglobin molecules of humans is more similar to that of chimpanzees than it is to the hemoglobin of dogs. This similarity suggests that
(A) humans and dogs are more closely related than humans and chimpanzees
(B) humans and chimpanzees are more closely related than humans and dogs
(C) humans are related to chimpanzees but not to dogs
(D) humans and chimpanzees are closely analogous
46. Two individuals, one with type B blood and one with type AB blood, have a child. The probability that the child has type O blood is
(A) 0%
(B) 25%
(C) 50%
(D) 100%
Questions 47 and 48 refer to the following bar graph, which shows the relative biomass of four different populations of a particular food pyramid.
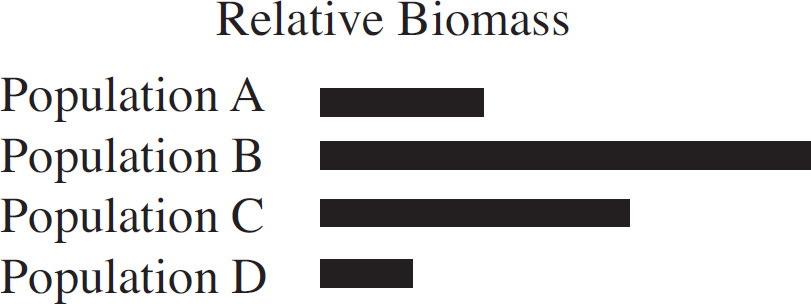
47. The largest amount of energy is available to
(A) population A
(B) population B
(C) population C
(D) population D
48. Which of the following would be the most likely result if there was an increase in the number of organisms in population C?
(A) The biomass of population D will remain the same.
(B) The biomass of population B will decrease.
(C) The biomass of population A will steadily decrease.
(D) The food source available to population C would increase.
Questions 49–52 refer to the following illustration and information.
The cell cycle is a series of events in the life of a dividing eukaryotic cell. It consists of four stages: G1, S, G2, and M. The duration of the cell cycle varies from one species to another and from one cell type to another. The G1 phase varies the most. For example, embryonic cells can pass through the G1 phase so quickly that it hardly exists, whereas neurons are arrested in the cell cycle and do not divide.
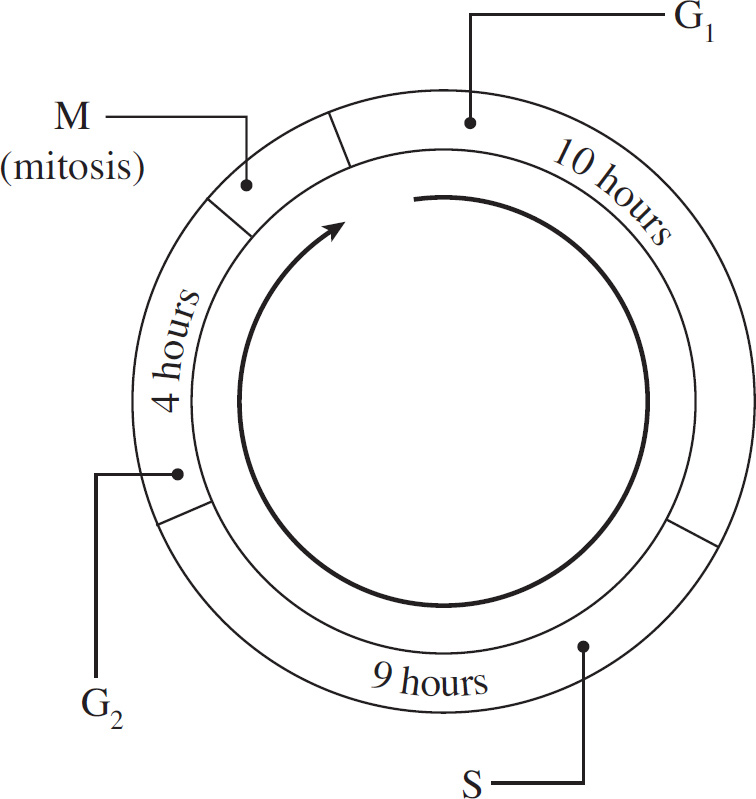
49. During which phase do chromosomes replicate?
(A) G1
(B) S
(C) G2
(D) M
50. In mammalian cells, the first sign of mitosis beginning is the
(A) appearance of chromosomes
(B) separation of chromatids
(C) disappearance of the cell membrane
(D) replication of chromosomes
51. Mitosis occurs in all of the following types of cells EXCEPT
(A) epidermal cells
(B) hair cells
(C) red blood cells
(D) pancreatic cells
52. Since neurons are destined never to divide again, what conclusion can be made?
(A) These cells will go through cell division.
(B) These cells will be permanently arrested in the G1 phase.
(C) These cells will be permanently arrested in the M phase.
(D) These cells will quickly enter the S phase.
Questions 53–56 refer to the following graphs, which show the permeability of (p) ions during an action potential in a ventricular contractile cardiac fiber. The left graph shows the membrane potential changes, and the right shows the corresponding ion permeabilities over the time frame of the action potential.
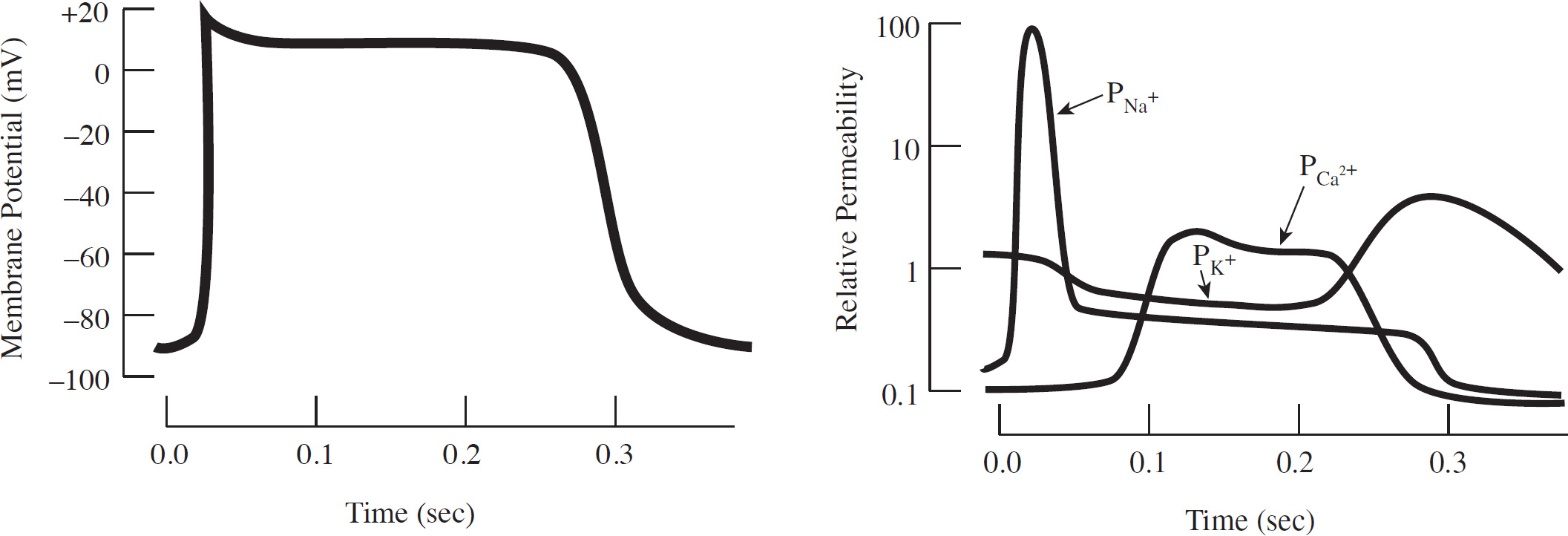
53. Based on the graph, the resting membrane potential of the muscle fibers is closest to
(A) −90 mV
(B) –70 mV
(C) 0 mV
(D) +70 mV
54. Which of the following statements is true concerning the initial phase of depolarization?
(A) Voltage-gated K+ channels open in the plasma membrane.
(B) The concentration of Ca2+ ions within the plasma membrane becomes more negative.
(C) The membrane potential stays close to −40 mV.
(D) The permeability to Na+ ions increases.
55. In cardiac fibers, the duration of an action potential is approximately
(A) 0.10 secs
(B) 0.20 secs
(C) 0.25 secs
(D) 0.30 secs
56. The action potential of skeletal muscle cells is much shorter than that of cardiac cells. Which of the following facts about cardiac muscle best explains this?
(A) The membrane is permeable to Na+, not K+.
(B) Voltage-gated K+ channels open during depolarization, not repolarization.
(C) Depolarization is prolonged compared to that in skeletal muscle fibers.
(D) The refractory period is shorter than that of skeletal muscle fibers.
Questions 57 and 58 refer to the data below concerning the general animal body plan of four organisms.

Note: + indicates a feature presence in an organism.
57. The two most closely related organisms are
(A) sea anemone and hagfish
(B) eel and salamander
(C) hagfish and eel
(D) sea anemone and salamander
58. The correct order of evolution for the traits above is
(A) jaws – vertebral column – walking legs
(B) walking legs – jaws – vertebral column
(C) jaws – walking legs – vertebral column
(D) vertebral column – jaws – walking legs
59. Pre- and post-zygotic barriers exist that prevent two different species from producing viable offspring. All of the following are pre-zygotic barriers EXCEPT
(A) anatomical differences preventing copulation
(B) different temporality of mating
(C) sterility of offspring
(D) incompatible mating songs
60. Birds and insects have both adapted wings to travel by flight. The wings of birds and insects are an example of
(A) divergent evolution
(B) convergent evolution
(C) speciation
(D) mutation
Questions 61–63 refer to the following synthetic pathway of nRNA pyrimidine, cytidine 5′ triphosphate, CTP. This pathway begins with the condensation of two small molecules by the enzyme aspartate transcarbamylase (ATCase).
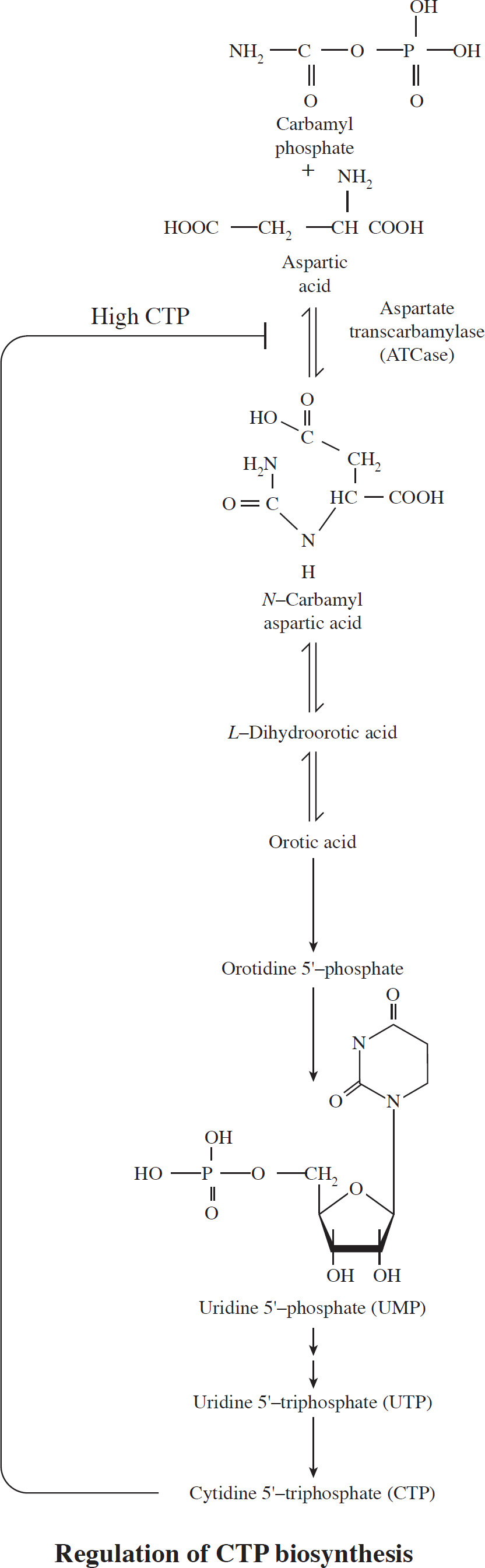
61. Which of the following is true when the level of CTP is low in a cell?
(A) CTP is converted to ATCase.
(B) The metabolic traffic down the pathway increases.
(C) ATCase is inhibited, which slows down CTP synthesis.
(D) The final product of the pathway is reduced.
62. This enzymatic phenomenon is an example of
(A) transcription
(B) feedback inhibition
(C) dehydration synthesis
(D) photosynthesis
63. The biosynthesis of cytidine 5′-triphosphate requires
(A) a ribose sugar, a phosphate group, and a nitrogen base
(B) a deoxyribose sugar, a phosphate group, and a nitrogen base
(C) a ribose sugar, phosphate groups, and a nitrogen base
(D) a deoxyribose sugar, phosphate groups, and a nitrogen base
Directions: Part B consists of questions requiring numeric answers. Calculate the correct answer for each question.
64. In a diploid organism with the genotype AaBbCCDDEE, how many genetically distinct kinds of gametes would be produced?

65. Under favorable conditions, bacteria divide every 20 minutes. If a single bacterium replicated according to this condition, how many bacterial cells would one expect to find at the end of three hours?

66. In snapdragon plants that display intermediate dominance, the allele CR produces red flowers and CW produces white flowers. If a homozygous, red-flowered snapdragon is crossed with a homozygous, white-flowered snapdragon, what will the percentage of pink offspring be?

67. Translation is an energy-intensive process. Each tRNA that brings an amino acid costs 2 ATP to make. Every codon that binds in the ribosome costs 1 ATP. There is also a cost of 1 ATP every time the mRNA must shift to the next codon. Approximately how many ATPs are required to synthesize a protein containing 115 amino acids?

Question 68 refers to the following experiment.
A group of Daphnia, small crustaceans known as water fleas, was placed in one of three culture jars of different sizes to determine their reproductive rate. There were 100 females in the jar. The graph below shows the average number of offspring produced per female each day in each jar of pond water.
68. What is the total number of offspring born in the 0.5-liter jar on the twentieth day?

69. On average, there is a 90 percent reduction of productivity for each trophic level. Based on this information, 10,000 pounds of grass should be able to support how many pounds of crickets?

STOP
END OF SECTION I
IF YOU FINISH BEFORE TIME IS CALLED, YOU MAY CHECK YOUR WORK ON THIS SECTION. DO NOT GO ON TO SECTION II UNTIL YOU ARE TOLD TO DO SO.
BIOLOGY
SECTION II
8 Questions
Planning Time—10 minutes
Writing Time—80 minutes
Directions: Questions 1 and 2 are long free-response questions that should require about 22 minutes each to answer and are worth 10 points each. Questions 3 through 8 are short free-response questions that should require about 6 minutes each to answer. Questions 3 through 5 are worth 4 points each, and questions 6 through 8 are worth 3 points each.
Read each question carefully and completely. Write your response in the space provided following each question. Only material written in the space provided will be scored. Answers must be written out in paragraph form. Outlines, bulleted lists, or diagrams alone are not acceptable.
1. Chlorophyll is one of a class of pigments that absorb light energy in photosynthesis.
(a) Describe the function of chlorophyll and what is meant by its absorption spectrum.
(b) Design an experiment to investigate the absorption spectrum of a pigment given to you in a solution and make a mock graph. Using the mock results of your experiment, what color should the pigment appear as?
2. Natural selection is said to act on individuals, but evolution occurs in populations. Explain each process and show that the statement is true using an example with individuals in a population. Identify variation within the population. Identify a selective pressure. Create a graph illustrating the changes to the population makeup over time.
3. Describe the chemical nature of genes. Name two types of gene mutations that could occur during replication.
4. In large multicellular organisms, signaling is necessary to maintain homeostasis. Compare and contrast the endocrine system and the nervous system with a focus on how they send signals.
5. Describe why fermentation is a less efficient way to produce energy than aerobic respiration.
6. Define analogous structures and describe how similar selective pressures can occur in two unrelated species.
7. Describe symbiosis and give an example involving humans.
8. Genes can be transferred from parent to offspring, but there are many other ways that they can be transferred, either naturally or using biotechnology. Describe two ways that genes can be transferred and explain why these types of transfers are helpful.
STOP
END OF EXAM
Practice Test 2: Answers and Explanations
PRACTICE TEST 2 ANSWER KEY
1. D
2. C
3. D
4. B
5. D
6. D
7. C
8. C
9. C
10. A
11. D
12. B
13. C
14. B
15. D
16. A
17. A
18. B
19. D
20. C
21. D
22. D
23. A
24. C
25. D
26. C
27. D
28. D
29. B
30. C
31. C
32. D
33. B
34. B
35. B
36. C
37. B
38. B
39. B
40. C
41. A
42. C
43. B
44. B
45. B
46. A
47. B
48. B
49. B
50. A
51. C
52. B
53. A
54. D
55. D
56. C
57. B
58. D
59. C
60. B
61. B
62. B
63. C
64. 4
65. 512
66. 100
67. 460
68. 200
69. 1,000
PRACTICE TEST 2 EXPLANATIONS
Section I: Multiple Choice
1. D Animal cells have centrioles. Both animal and plant cells have an endoplasmic reticulum, membrane-bound organelles, and lysosomes. Only plants have a cell wall made of cellulose and large vacuoles.
2. C If the contents of the cell separated from the cell wall, then water was moving out of the cell. This would cause the space between the cell wall and the cell membrane to expand, so eliminate (B). This also means that the concentration of solutes in the extracellular environment would be hypertonic with respect to the cell’s interior. You can also eliminate (A): because the fluid in the cell was hypotonic to the sugar solution, fluid was moving out of the vacuole, which caused it to become smaller. Choice (C) is the correct answer because if the water was rushing out of the cell then the water potential inside the vacuole would have been higher than outside the cell. High water potential means the water wants to move. Choice (C) says the water potential would be lower.
3. D This question tests your ability to associate what happens when enzymes are denatured and what would happen in the synaptic cleft. Acetylcholinesterase is an enzyme that degrades acetylcholine in the synaptic cleft. If acetylcholinesterase is denatured, acetylcholine will still be released from the presynaptic membrane and continue to diffuse across the synaptic cleft. Acetylcholine will still bind to the postsynaptic membrane because acetylcholine is not degraded. Therefore, you can eliminate (A), (B), and (C).
4. B The ratio of purines to pyrimidines should be constant because purines always bind with pyrimidines, no matter which ones they may be.
5. D Feedback inhibition is when something created by a process then inhibits that process. Choices (A) and (B) involve stimulations, so they can be eliminated. Choice (C) doesn’t make sense because it says that “C” is an enzyme and the passage says that the enzymes are the numbers shown between the steps.
6. D If substance F leads to the inhibition of enzyme 3, then substances D and E and enzymes 3, 4, and 5 will be affected. The activity of enzyme 5 will be decreased, not increased.
7. C If enzyme 1 were inhibited, then everything that comes after that in the pathway would be reduced. A would not be reduced, but both C and X would. Choice (C) is correct.
8. C Based on the graph, fetal hemoglobin has a higher affinity for oxygen than maternal hemoglobin. Fetal hemoglobin does not give up oxygen more readily than maternal hemoglobin, so eliminate (A). You can also get rid of (B), as the dissociation curve of fetal hemoglobin is to the left of the maternal hemoglobin. Finally, eliminate (D) because fetal hemoglobin and maternal hemoglobin are different structurally, but you can’t tell this from the graph.
9. C The passage says that high CO2 and low pH lead to reduced affinity. A right shift curve would represent a reduced affinity. Photosynthesis consumes CO2, so this would not increase the CO2. Mitosis and digestion are not related. Fermentation is done in cells that are doing anaerobic cell respiration, so CO2 should be released, possibly along with lactic acid, which would lower pH.
10. A Hemoglobin’s affinity for O2 decreases as the concentration of H+ increases (or the pH decreases) and as the concentration of OH− increases (or the pH increases). As the pH decreases, the affinity for oxygen will decrease, and as the pH increases the affinity for oxygen will increase.
11. D The CO2-rich environment would decrease the affinity of hemoglobin for oxygen, so the curve should shift towards the right. The regular curve seems to hit 90% saturation around 45–50 pO2, so the CO2-rich curve would take more oxygen to get there.
12. B Both prokaryotes and eukaryotes possess double-stranded DNA. Choices (A), (C), and (D) are correct differences between prokaryotes and eukaryotes.
13. C Bb and Bb is the correct set of genotypes that represents a cross that could produce offspring with silver fur from parents that both have brown fur. Complete a Punnett square for this question. In order for the offspring to have silver fur, both parents must have the silver allele.
14. B Viruses are made up of nucleic acid surrounded by a protein coat called a capsid. Don’t forget that some viruses are RNA viruses and don’t have a DNA genome.
15. D All of the choices are examples of hydrolysis except the conversion of pyruvic acid to glucose. Hydrolysis is the breaking of a covalent bond by adding water. In all of the correct examples, complex compounds are broken down to simpler compounds. The conversion of pyruvic acid to acetyl-CoA is an example of decarboxylation—a carboxyl group is removed as carbon dioxide and the 2-carbon fragment is oxidized.
16. A Peroxisomes catalyze reactions that produce hydrogen peroxide, ribosomes are involved in protein synthesis, and mitochondria contain enzymes involved in cellular respiration. Eliminate (B) and (D) because lysosomes are the sites of degradation; they contain hydrolytic enzymes but do not produce hydrogen peroxide. Choice (C) is incorrect, as the Golgi apparatus sorts and packages substances that are destined to be secreted out of the cell.
17. A The figure shows that water leaves the descending limb and flows by osmotic pressure into the surrounding space and then into the vasa recta. This means that each of those places is more hypertonic than the previous. The most hypotonic is the descending tubule (This is why the water leaves.).
18. B When water flows out of the tubule, it is moving by simple osmosis. However, because it is a polar molecule, it must pass through an aquaporin channel, which is facilitated diffusion.
19. D ADH makes the collecting duct permeable to water. If it is inhibited, then it will be unable to do that, meaning choice (D) is correct. The question doesn’t say that the inhibitor is inhibiting the aquaporins directly. Instead, the inhibitor says it is inhibiting the ADH.
20. C To make concentrated urine, the medulla must be very osmotic because it must reclaim a lot of water from the collecting duct. To make a very osmotic medulla, the Loop of Henle must be very long. The collecting duct should be very permeable, not impermeable, because concentrated urine means that a lot of water has been reclaimed.
21. D The carrying capacity is the maximum number of organisms of a given species that can be maintained in a given environment. Once a population reaches its carrying capacity, the number of organisms will fluctuate around it.
22. D During the exponential growth phase of a population, the size doubles during each time interval. This part of the graph looks like a parabola.
23. A The tendency of climbing vines to twine their tendrils around a trellis is called thigmotropism. Thigmotropism refers to growth stimulated by contact with an object. Choice (B), hydrotropism, refers to growth of a plant toward water. Choice (C), phototropism, refers to growth toward light. Choice (D), geotropism, refers to growth toward or against gravity.
24. C Females with Turner’s syndrome lack an X chromosome. If females with this syndrome have a high rate of hemophilia, they must not have the second X to mask the expression of the disease.
25. D This type of mutation is called frameshift mutation. The insertion of DNA leads to a change in the normal reading frame by one base pair. The other answer choices refer to chromosomal aberrations. Choice (A), duplication, is when an extra copy of a chromosome segment is introduced. Choice (B), translocation, is when a segment of a chromosome moves to another chromosome. Choice (C), inversion, is when a segment of a chromosome is inserted in the reverse orientation.
26. C The principal inorganic compound found in living things is water. Water is a necessary component for life.
27. D An extra X-chromosome could be created by a problem separating the homologous chromosomes or sister chromatids. When they are lined up it would not be apparent that they were not separating properly, but once they start to pull apart one could see if a mistake had occurred.
28. D All of the following are modes of asexual reproduction except meiosis, which is the production of gametes for sexual reproduction. Choice (A), sporulation, is a form of asexual reproduction in which spores are produced. Choice (B), fission, is the equal division of a bacterial cell. Choice (C), budding, is a form of asexual reproduction in yeasts in which small cells grow from a parent cell.
29. B Invertebrates lack specific immune responses including B-cells and T-cells. Phagocytes is the only answer choice that describes non-specific immune responses.
30. C Helicase is the enzyme that unwinds the helix. The closed helix would exist without helicase.
31. C The RNA polymerase is not bound, but there is a repressor bound. The transcription must be repressed.
32. D These four phases must be transcription. Transcription begins with a closed helix, then an enhancer binds, and then RNA polymerase is seen making RNA. Finally, the closed helix and the completed mRNA strand exist.
33. B Photosynthesis requires light, carbon dioxide, and water and produces oxygen and glucose.
34. B Avery, MacLeod, and McCarty concluded that DNA is the hereditary material because it was able to transform non-pathogenic bacteria into pathogenic bacteria. Watson, Crick, and Franklin determined the double helix, and Meselson determined that replication is semi-conservative.
35. B Translation, the synthesis of proteins from mRNA, occurs in the cytoplasm. DNA replication (I) occurs in the nucleus. Transcription (II), the synthesis of RNA from DNA, occurs in the nucleus.
36. C Crossing-over permits scientists to determine chromosome mapping. Chromosome mapping is a detailed map of all the genes on a chromosome. The frequency of crossing-over between any two alleles is proportional to the distance between them. The farther apart the two linked alleles are on a chromosome, the more often the chromosome will break between them. Crossing-over does not tell us about the chance of variation in zygotes, the rate of mutations, or whether the traits are dominant, recessive, or masked.
37. B In this scenario, the inside of the cell is hypertonic to the outside environment—this will cause water to move into the cell and can cause the cell to lyse.
38. B The most likely explanation for this phenomenon is that these birds have different ecological niches. An ecological niche is the position or function of an organism or population in its environment. Eliminate (A) because we do not know if there is a short supply of resources. Choice (C) can also be eliminated because we do not know how long the bird species live together. The breeding patterns of the bird species do not explain the lack of competition, which eliminates (D) as well.
39. B This relationship is an example of parasitism. Parasitism is a form of symbiosis in which one organism benefits and the other is harmed. Choice (A), commensalism, is a form of symbiosis in which one organism benefits and the other is unaffected. Choice (C), mutualism, is a form of symbiosis in which both organisms benefit. Choice (D), gravitropism, is the growth of a plant toward or away from gravity.
40. C If the nucleotide sequence of a DNA molecule is 5′-C-A-T-3′, then the transcribed DNA strand (mRNA) would be 3′-G-U-A-5′. The nucleotide sequence of the tRNA codon would be 5′-C-A-U-3′.
41. A Viruses are considered an exception to the cell theory because they are not independent organisms. They can only survive by invading a host. Choice (B) is incorrect, as all viruses have genomes and some have many genes. Choice (C) is also wrong because not all viruses have a tail. You can also eliminate (D), as viruses did not evolve from ancestral protists.
42. C The origin of life refers to when the first life began. Life is shown to be present 3.5 billion years ago when there was not atmospheric oxygen present yet, so (A) can be eliminated. Also, there was no life at 4 billion years despite having CO2 and no O2; therefore something else must be required, and (B) can be eliminated. There were not proteins when the first life arose so (D) is incorrect. Choice (C) states that self-replicating nucleic acids were necessary, which is why life had to occur when RNA appeared.
43. B Functional ribosomes would not likely be found before the proteins they make. Therefore, this would be between 3.5 and 3.25 billion years ago.
44. B The sign that photosynthesis began was when oxygen appeared. Oxygen was absent at 3.25 billion years and present at 3 billion years so photosynthesis must have begun between those times, eliminate (A) and (C). Autotrophs perform photosynthesis not heterotrophs (they rely on eating other things for their energy).
45. B The similarity suggests that humans and chimpanzees are more closely related than humans and dogs. Because these two organisms share similar amino acid sequences, humans must share more recent common ancestors with chimpanzees than with dogs.
46. A There is no way for these two parents to produce a type O child. The only genotype that will result in type O blood is two “O” alleles, one from each parent, since the allele for O blood is recessive to the alleles for A or B blood. The AB parent only has alleles for A and B blood, so it is impossible for these two individuals to produce a child with type O blood.
47. B The largest amount of energy is available to producers. Population B is most likely composed of producers because they have the largest biomass.
48. B An increase in the number of organisms in population C would most likely lead to a decrease in the biomass of B because population B is the food source for population C. Make a pyramid based on the biomasses given. If population C increases, population B will decrease. Eliminate (A) and (C), as we cannot necessarily predict what will happen to the biomass of populations that are above population C. Choice (D) can also be eliminated because the food source available to population C would most likely decrease, not increase.
49. B Chromosomes replicate during interphase, the S phase. Choices (A) and (C) are incorrect because during G1 and G2, the cell makes protein and performs other metabolic duties.
50. A The first sign of prophase in mammalian cells is the appearance of chromosomes, which are usually invisible. They are condensed at the start of mitosis.
51. C Mitosis occurs in all of the following type of cells except mature red blood cells. Mature red blood cells are short-lived and do not divide.
52. B Because neurons are not capable of dividing, it is reasonable to conclude that these cells will not complete the G1 phase. This is a reading comprehension question. The passage states that cells that do not divide are arrested at the G1 phase. Choice (A) is incorrect because these cells will not be committed to go through cell division. You can also eliminate (C) and (D), as the cells will not enter the M or S phase.
53. A According to the graph, the resting membrane potential of the muscle fiber is close to −90mV.
54. D Refer to the second graph about the membrane permeability of ions during a muscle contraction. During depolarization, the membrane is permeable to Na+. Choice (A) can be eliminated because the voltage-gated K+ channels do not open until after depolarization. Eliminate (B) because the concentration of Ca2+ does not become more negative. Choice (C) is also incorrect, as the membrane potential changes from −90 mV to +20 mV.
55. D Refer to both graphs in the passage. In cardiac fibers, the duration of an action potential—a neuronal impulse—is approximately 0.3 seconds.
56. C In cardiac muscle fibers, depolarization is prolonged compared to that in skeletal muscle fibers. Eliminate (A) because the membrane is permeable to both Na+ and K+. Choice (B) can also be eliminated; in cardiac muscle fibers, voltage-gated K+ channels open during repolarization. Finally, (D) is wrong because the refractory period is longer in cardiac muscle fibers compared to skeletal muscle fibers.
57. B The two most closely related organisms are the two with the most shared derived characteristics.
58. D Shared derived characteristics are newly evolved traits that are shared with every group on a phylogenic tree except for one. Vertebral columns are present in every group except for the sea anemone, so it must have evolved first. Walking legs are only found in the salamander, indicating that it most likely evolved most recently.
59. C Pre-zygotic barriers to reproduction are those that prevent fertilization, so you can eliminate (A), (B), and (D). Choice (C) is an example of a post-zygotic barrier to reproduction.
60. B Convergent evolution occurs when two organisms that are not closely related independently evolve similar traits, such as the wings of insects and birds. Divergent evolution occurs when two closely related individuals become different over time and can lead to speciation.
61. B When the level of CTP is low in a cell, the metabolic traffic down the pathway increases. This pathway is controlled by feedback inhibition. The final product of the pathway inhibits the activity of the first enzyme. When the supply of CTP is low, the pathway will continue to produce CTP.
62. B This enzymatic phenomenon is an example of feedback inhibition. Feedback inhibition is the metabolic regulation in which high levels of an enzymatic pathway’s final product inhibit the activity of its rate-limiting enzyme. Transcription, (A), is the production of RNA from DNA. Dehydration synthesis, (C), is the formation of a covalent bond by the removal of water. In photosynthesis, (D), radiant energy is converted to chemical energy.
63. C The biosynthesis of cytidine 5-triphosphate requires a nitrogenous base, three phosphates, and a sugar ribose. Pyrimidines are a class of nitrogenous bases with a single ring structure. The sugar they contain is ribose, which is shown in the pathway diagram.
64. 4 There are four genetically distinct kinds of gametes that could be produced, ABCDE, AbCDE, aBCDE, and abCDE. Notice that the only alleles that vary are A and B (22 = 4).
65. 512 If bacteria divide every twenty minutes, you would produce 512 bacterial cells. One method would be to use the equation 2x, where x equals the number of twenty minute intervals in three hours; 29 = 512.
66. 100 Because snapdragon plants display intermediate dominance, the heterozygous phenotype is affected by the alleles of both homozygotes. If a homozygous red plant is crossed with a homozygous white plant, all offspring would be heterozygous and pink.
67. 460 A 115 amino acid protein requires 115 aa × 2 ATP = 230. A total of 116 codons will bind (115 amino acids and the stop codon). A total of 114 translocations will occur (The start codon begins in the P-spot, but there will be a translocation after the remaining 114 codons.). 230 + 116 + 114 = 460.
68. 200 The graph shows the average number of offspring per female per day. Because there are 100 females in the half liter, the total number of offspring on the twentieth day would be 100 females times 2 offspring per day, which equals 200 offspring.
69. 1,000 A 90 percent reduction of productivity in the grass is a reduction of 9,000 pounds. That means the grass should be able to support 1,000 pounds of crickets.
Section II: Free Response
Sample Long-Form Free Response 1
Chlorophyll is the principal light-harnessing pigment found in the thylakoid membrane of chloroplasts. Chlorophyll comes in two main types, chlorophyll a and chlorophyll b. Each type has a different wavelength absorption spectrum. When chlorophyll absorbs light, electrons within it are energized. Photosynthesis occurs when sunlight activates chlorophyll by exciting its electrons. Chlorophyll reflects green light, which is why plants appear green.
To determine the absorption spectrum of an unknown pigment, the pigment would have to be exposed to many different wavelengths of light and the amount of light absorbed at each should be measured. This can be done with a spectrophotometer. The wavelength of the entering light can be varied to see which wavelength is most absorbed by the solution. The least absorbed wavelengths are reflected back and that determines the color we see. As the graph shows, it would be green.
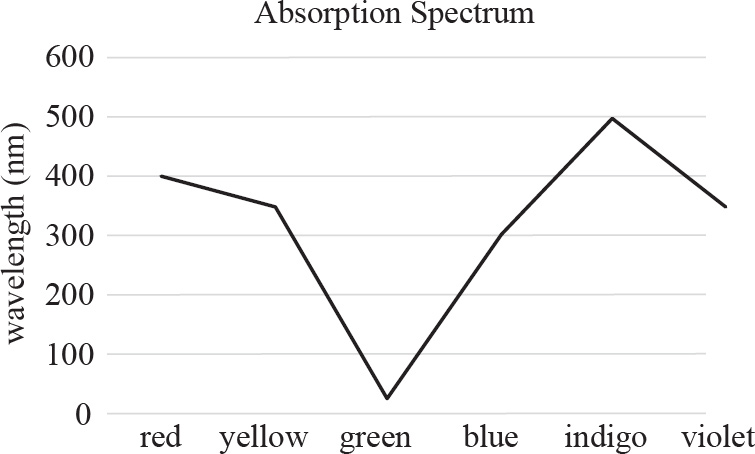
Sample Long-Form Free Response 2
Natural selection is the superior reproduction of individuals with certain inheritable traits. Those traits will contribute to the next generation, and less desirable traits will not be “selected” or passed along to the next generation. This cannot happen to a population because populations do not reproduce.
Evolution is the change of the gene pool of a population over time as certain traits disappear and others are selected. This cannot happen to an individual because an individual will have the same DNA throughout its lifetime.
As an example, let’s look at a population of squirrels. Half of them are dark brown and half of them are grey. The grey squirrels are a very similar color to the pavement of the road, cause them to blend in. Motorists often hit the grey squirrels on accident because they cannot see them until it is too late. The brown squirrels are very easy to see against the grey pavement, and the motorists slow down and drive around the squirrels. Over time, the population of squirrels changes and more and more brown squirrels are seen. This is because the brown is selected because it has an advantage over the grey squirrels (less likely to be killed by a motorist). Each squirrel is still either brown or grey, but the population overall is changing. The selection is at the squirrel level, but the evolution is at the population level.
Sample Short-Form Free Response 3
A gene is a heredity unit located at a specific locus along a chromosome. Genes are made up of DNA, and DNA is made up of repeating subunits of nucleotides. A nucleotide consists of three parts, a five-carbon sugar, a phosphate group, and a nitrogenous base.
A gene mutation is a change in the sequence of base pairs in a DNA molecule. It results from defects in the sequence of bases. This can be caused by an error of the DNA polymerase or because of some other type of DNA damage. Mutations that involve a single base change in the DNA sequence are called point mutations. This type of a mutation can cause a change in the mRNA codon, which might cause a change in the sequence of the polypeptide. These are base substitutions involving a single DNA nucleotide being replaced by another nucleotide. Another type of mutation is a frameshift mutation, caused by an insertion or deletion of bases in the DNA sequence. This causes a shift in the reading frame of DNA. This often causes major changes in the sequence of the poylpeptide because many codons can be disrupted.
Sample Short-Form Free Response 4
The endocrine system works when glands release messengers called hormones into the bloodstream. The hormones can be either peptide based or steroid based. They travel to a particular area of the body and then they bind to either an extracellular receptor (peptide hormones) or an intracellular receptor (steroid hormones). This causes changes in the cell, which causes the desired effect of the hormone.
The nervous system is a system of neuron pathways that go throughout the body. Signals are sent from out in the body and into the central nervous system (brain and spinal cord). The message format of neurons is action potentials, which are waves of positive charges that travel down neuron axons. Neuron axons synapse with other neurons, and neurotransmitters are sent across the synapse to the next neuron or effector cell. Certain types of cells, like muscle cells, will contract when they are stimulated by a neuron signal.
The similarity is that each system is responding to a stimulus and passing a message to another place in the body, The nervous system uses action potentials traveling down neuron circuitry, but the endocrine system uses hormones traveling through the bloodstream. The nervous system is also much faster acting than the endocrine system.
Sample Short-Form Free Response 5
Fermentation occurs when oxygen is not available to act as the final electron acceptor. When this does not occur, then NADH and FADH2 are not able to be regenerated into NAD+ and FAD, respectively. Instead, the only step in respiration that is occurring is glycolysis, which makes 2 ATP, 2 NADH, and pyruvate. Pyruvate can act as the final electron acceptor and allow NAD+ to be regenerated from the NADH made in glycolysis, but the only energy that has been produced is the net of 2 ATP. Since not all the energy can be mined from the carbon when processed in fermentation, the process is less efficient. Additionally, the process of reducing pyruvate makes lactic acid or ethanol, which are toxic in large amounts. Aerobic respiration is better.
Sample Short-Form Free Response 6
Analogous structures are ones which have similar functions, but whose underlying structures are very different. They are often the product of convergent evolution, which is when differing organisms end up in the same environment and thus are subjected to the same forces of natural selection. An example of analogous structures is the wing of bird and the wing of a bee. Both are used for flight, but their components are completely different. For those same adaptations to occur in both unrelated species, the same selective pressures must have occurred. For example, the ability to fly may give animals access to an unoccupied niche. It might also be a good way to avoid predators. So, in both birds and bees there was a benefit to flying.
Sample Short-Form Free Response 7
Symbiosis describes a situation in which two organisms co-exist. If the host provides a benefit, but remains unimpacted by the other organism, then the relationship is commensalistic, but if both organisms benefit, then it is mutualistic. An example involving humans is the bacterial flora of the skin or of the intestine. Many of these bacteria provide a protective or digestive benefit to humans, while some harmlessly utilize the body for food. Parasitic relationships are another type of symbiosis, but in this instance one individual is harmed in the process.
Sample Short-Form Free Response 8
In bacteria, genes are passed by conjugation between nearby bacteria. This is not a type of reproduction, but it is just gene swapping. A plasmid is passed between two bacteria, who then go their separate ways.
Viruses can also pass genetic information around, especially to bacteria. When a virus integrates inside a bacterial cell genome, it can leave a bit of genetic material and take some with it on accident. These bits join the bacterial genome. This process if called transduction.
These processes are helpful because they increase the genetic variation among the population. Bacteria replicate by fission, which is basically just making copies of themselves. Conjugation mixes up the gene pool a bit. Transduction does the same thing by picking up and dropping bits of DNA here and there. More genetic variation is better for survival of a population.
Practice Test 3
Click here to download a PDF of Practice Test 3
The Exam
AP® Biology Exam
SECTION I: Multiple-Choice Questions
DO NOT OPEN THIS BOOKLET UNTIL YOU ARE TOLD TO DO SO.
At a Glance
Total Time
1 hour and 30 minutes
Number of Questions
69
Percent of Total Score
50%
Writing Instrument
Pencil required
Instructions
Section I of this examination contains 69 multiple-choice questions. These are broken down into Part A (63 multiple-choice questions) and Part B (6 grid-in questions).
Indicate all of your answers to the multiple-choice questions on the answer sheet. No credit will be given for anything written in this exam booklet, but you may use the booklet for notes or scratch work. After you have decided which of the suggested answers is best, completely fill in the corresponding oval on the answer sheet. Give only one answer to each question. If you change an answer, be sure that the previous mark is erased completely. Here is a sample question and answer.
Sample Question
Chicago is a
(A) state
(B) city
(C) country
(D) continent
Sample Answer

Use your time effectively, working as quickly as you can without losing accuracy. Do not spend too much time on any one question. Go on to other questions and come back to the ones you have not answered if you have time. It is not expected that everyone will know the answers to all the multiple-choice questions.
About Guessing
Many candidates wonder whether or not to guess the answers to questions about which they are not certain. Multiple choice scores are based on the number of questions answered correctly. Points are not deducted for incorrect answers, and no points are awarded for unanswered questions. Because points are not deducted for incorrect answers, you are encouraged to answer all multiple-choice questions. On any questions you do not know the answer to, you should eliminate as many choices as you can, and then select the best answer among the remaining choices.
BIOLOGY
SECTION I
69 Questions
Time—90 minutes
Directions: Each of the questions or incomplete statements below is followed by four suggested answers or completions. Select the one that is best in each case and then fill in the corresponding oval on the answer sheet.
1. Klinefelter’s syndrome is a genetic condition in males that results in the creation of an extra X chromosome during the early stages of embryo development, which must be inactivated. Which of the following karyotypes represents a person with Klinefelter’s?
(A)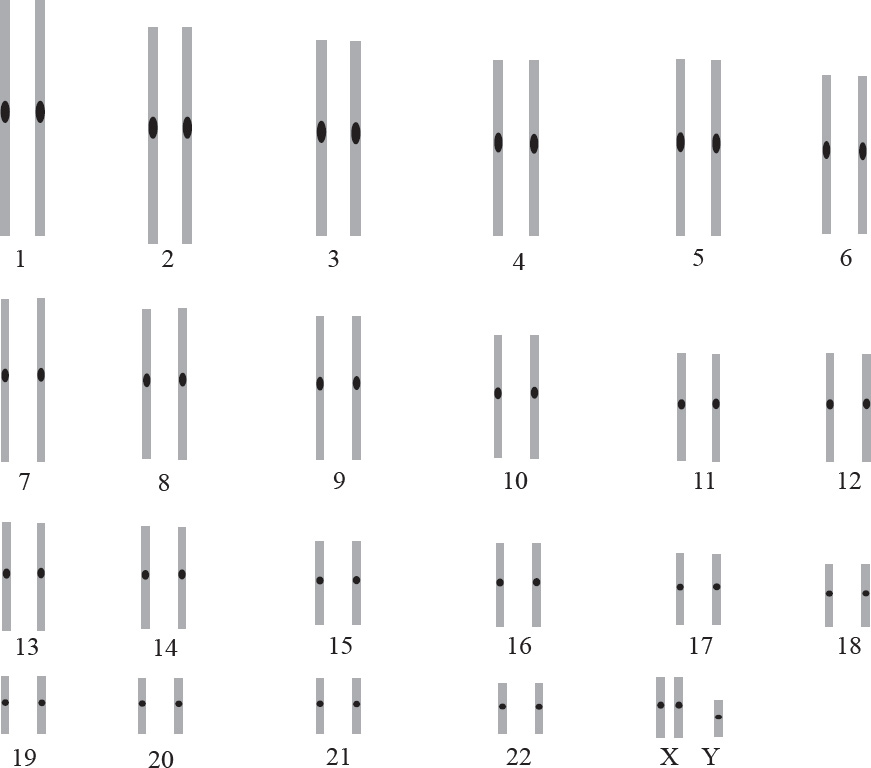 
(B)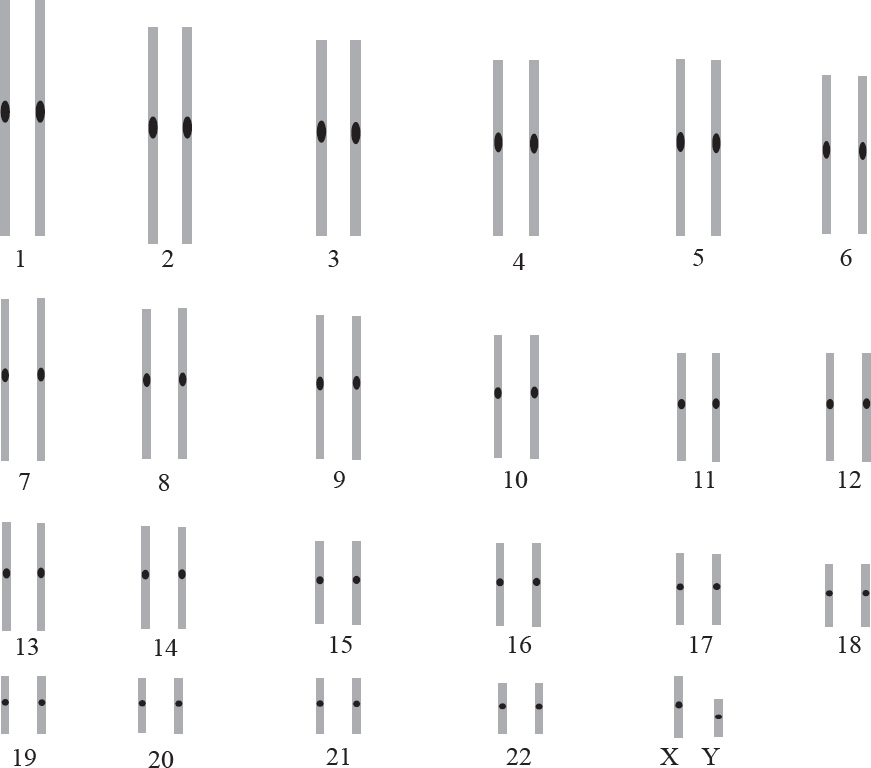 
(C)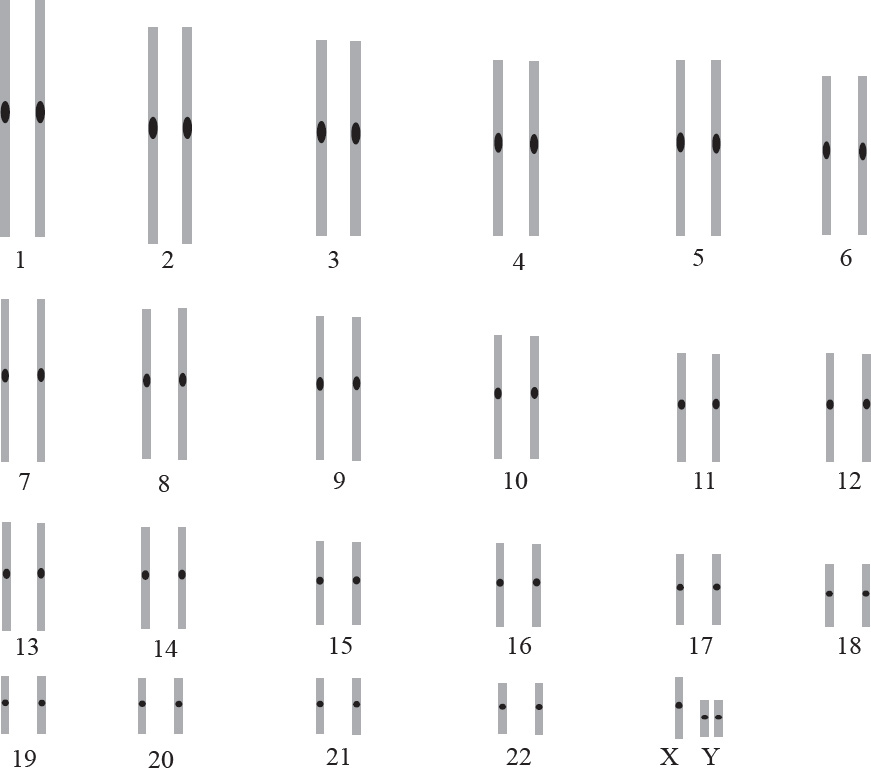 
(D)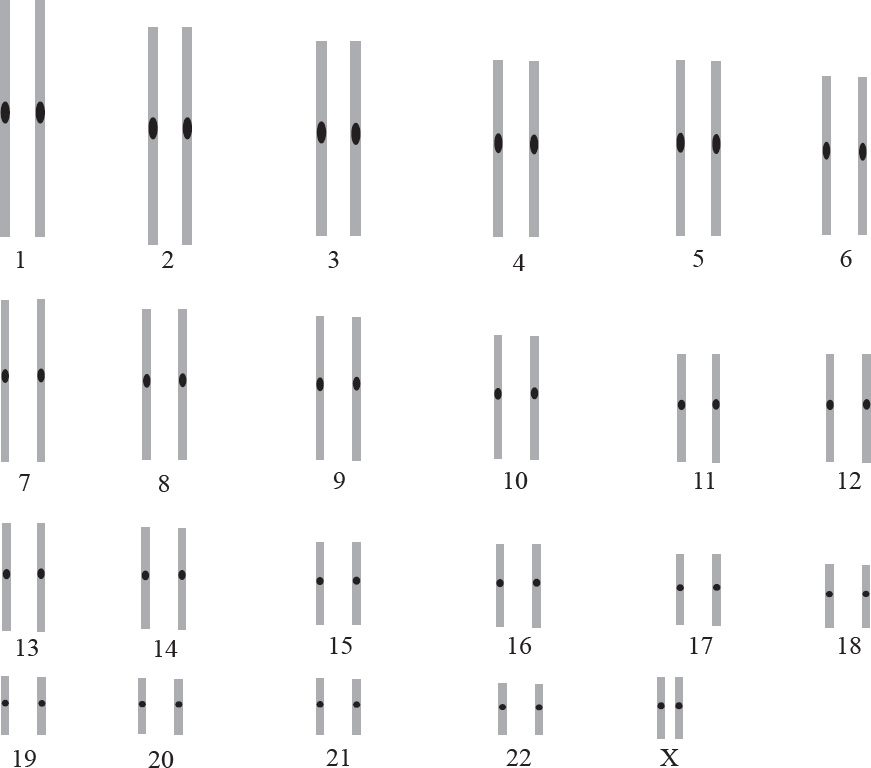 
2. The restriction enzyme BAMHI predictably cuts the following sequence at the locations designated by the star.
5′ G*GATCC 3′
3′ CCTAG*G 5′
Which of the following DNA helix sequences would be cut into 3 pieces by BAMHI?
(A) 5′ GGATCC 3′
3′ CCTAGG 5′
(B) 5′ GGATCCGGATCCGGATCC 3′
3′ CCTAGGCCTAGGCCTAGG 5′
(C) 3′ GGATCC 5′
5′ CCTAGG 3′
(D) 5′ AAGGATCCGGATCCAA 3′
3′ TTCCTAGGCCTAGGTT 5′
Questions 3–5 refer to the following passage.
In a species of peas, green (G) peas are known to be classically dominant over yellow (g) peas. A true breeding green plant and a true breeding yellow plant are crossed. The resulting F1 generation is evaluated, and its phenotype distribution is shown in Table 1. Two green pea producing members of the F1 generation are then crossed, and the resulting F2 generation phenotype distribution is shown in Table 2.
Table 1. F1 Generation
| Type of Peas Produced | # of Plants |
| Green | 1432 |
| Yellow | 1 |
Table 2. F2 Generation
| Type of Peas Produced | # of Plants |
| Green | 1196 |
| Yellow | 374 |
3. You are given a green pea plant with an unknown genotype. You want to figure out the genotype by crossing it with another plant and looking at the offspring. Which of the following plants would be the most helpful in determining the genotype of the original plant?
(A) Any green pea plant
(B) A homozygous green plant
(C) A yellow pea plant
(D) None of the above
4. Which of the following best explains the single yellow pea producing plant in the F1 generation?
(A) Crossing over occurred in the parental pea plant to provide genetic variation.
(B) The law of independent assortment: one homologous pair separated independently of the others during gamete production in the parental pea plant.
(C) Nondisjunction in one of the parental pea plants caused an imbalance in chromosome distribution.
(D) A DNA polymerase error occurred during DNA replication just prior to meiosis.
5. If a yellow plant from the F2 generation is crossed with a green plant from the F1 generation, what would be the green-to-yellow phenotype ratio?
(A) 1:1
(B) 3:1
(C) 1:0
(D) 1:3
6. According to the information in the figure, a sensory neuron would likely be oriented in which of the following ways?

(A) Dendrite connecting to the anterior pituitary in the brain and axon connecting to a skeletal muscle cell
(B) Dendrite connecting to the photoreceptors in the eye and axon connected to a smooth muscle cell
(C) Dendrite connecting to the photoreceptors in the eye and axon connecting to the brain
(D) Dendrite connecting to the anterior pituitary in the brain and axon connecting to the spinal cord
7. Cells in the body are related to each other by many types of intercellular connections such as desmosomes, adherens junctions, tight junctions, and gap junctions. In the intestines, tight junctions are necessary because an absolute barrier must be maintained to separate the lumen of the gut from the rest of the body. Gap junctions are found in cardiac muscle where these direct connections from cell to cell allow action potentials to be passed. Adherens junctions and desmosomes are similar types of connections between cells that are flexible and mechanically pull on each other. Desmosomes attach to the intermediate filaments in the cytoskeleton.
Compact bone contains rings of osteons, each of which contains a central canal housing the blood vessels, which can only be accessed by those osteocytes adjacent to it. What type of cell junctions would best help distant osteocytes exchange gases with the central canal?
(A) Desmosomes
(B) Gap junctions
(C) Tight junctions
(D) Adherens junctions
Questions 8–10 refer to the following passage.
The somatic cells in a newly identified sexually reproducing species are found to be octoploidy (8n), and each cell contains 32 chromosomes, slightly fewer than the 46 chromosomes in a somatic human cell. The cell cycle of this species is similar to ours, and gametes are made during meiosis. Two gametes will come together at fertilization to create octoploidy offspring.
8. How many chromosome segments are present during the G1 phase?
(A) 64
(B) 32
(C) 16
(D) 8
9. How many unique, non-homologous chromosomes are present in this species?
(A) 4
(B) 8
(C) 16
(D) 32
10. How many chromosome segments will be in each gamete if you count each homologous member individually?
(A) 32
(B) 16
(C) 8
(D) 4
Questions 11–14 refer to the following passage.
A population of goldfish in a large, isolated pond were studied in 1958 and again in 2008. The fishes’ pigment level varied from pale white-orange fish to dark brown-orange fish. The color of each of the fish was recorded and recorded in the figure below.
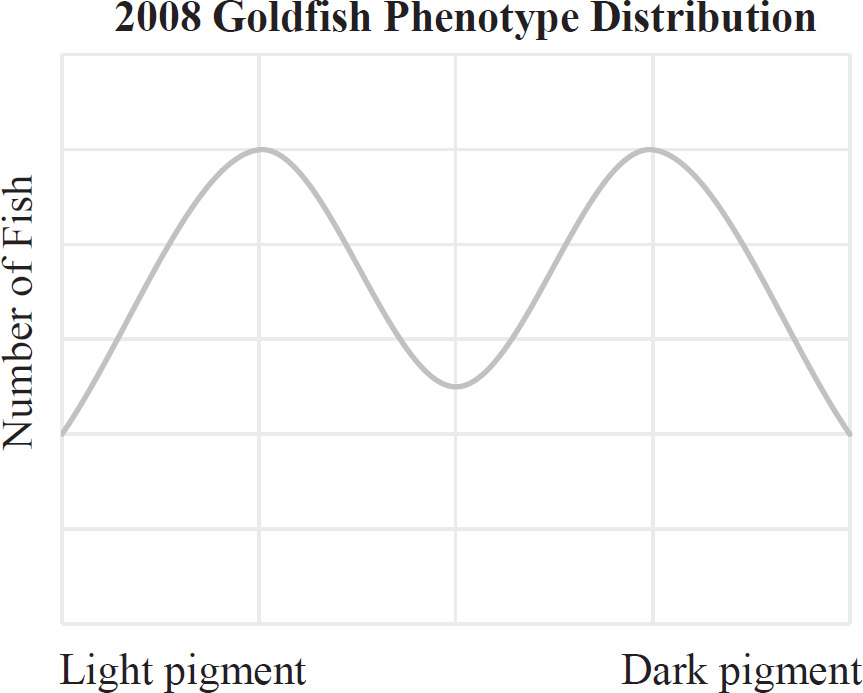
11. Which of the following best describes the fish population in the year 1958?
(A) All medium-orange in color
(B) Mostly medium-orange fish with some nearly white and some nearly brown
(C) Mostly white fish and brown fish with a few orange fish
(D) Equal numbers of orange fish, white fish, and brown fish
12. Which of the following theories is supported by the evidence?
(A) The pigment trait in fish demonstrates incomplete dominance.
(B) The pigment trait in fish demonstrates classical dominance.
(C) The pigment trait in fish demonstrates codominance.
(D) None of the above
13. If a dark colored, poisonous fish of similar appearance to these fish is added to the ecosystem in 2008, what will the graph likely look like in 50 years?
(A) 
(B) 
(C) 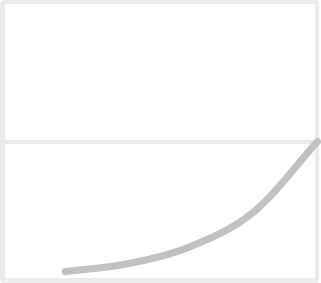
(D) 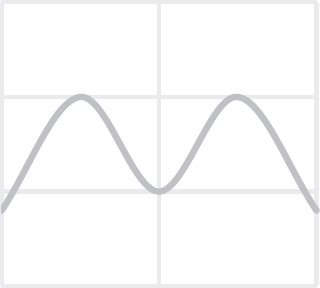
14. Which addition to the pond is NOT a likely contributor to the change between 1958 and 2008?
(A) A poisonous fish with a medium orange pigment
(B) Runoff from fields that makes the water dark and murky
(C) A light-orange water grass that grows in the pond
(D) Predatory birds that can easily see medium-orange pigment
Questions 15–16 refer to the following figure.
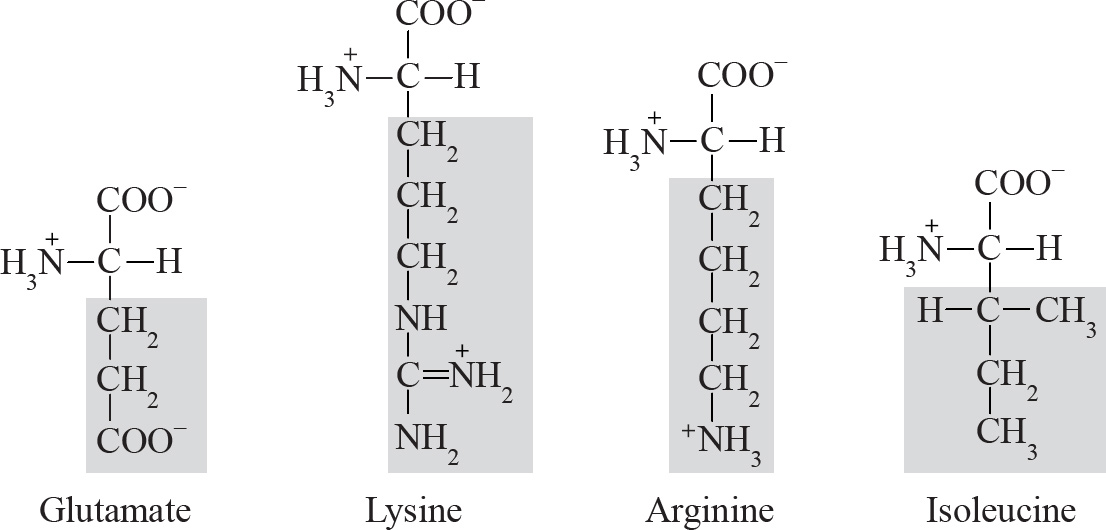
15. Which of the following amino acids would likely be found in the transmembrane domain?
(A) Lysine
(B) Arginine
(C) Isoleucine
(D) Glutamate
16. An enzyme’s active site contains arginine residues. Which residues will likely be found on the corresponding substrate region?
(A) Arginine
(B) Lysine
(C) Glutamate
(D) Isoleucine
Questions 17–19 refer to the following passage.
The extracellular environment of the human body is typically abundant in sodium and calcium. In skeletal muscle cells, the sarcoplasmic reticulum is a specialized organelle that actively sequesters calcium from the cytosol. It stockpiles the calcium until a motor neuron triggers its release through an action potential, which opens voltage-gated calcium channels in the sarcoplasmic reticulum membrane. The calcium is necessary for the calcium-mediated functions of the protein troponin. In the presence of calcium, troponin removes tropomyosin from the myosin binding sites on actin filaments. This attachment is essential for sarcomere and muscle contraction.
17. Which of following statements best describes the uptake of calcium by skeletal muscles and the sarcoplasmic reticulum?
(A) Calcium is passively taken from the extracellular environment and the cytosol.
(B) Calcium is actively taken up from the extracellular environment and the cytosol.
(C) Calcium is passively taken up from the extracellular environment and actively taken up from the cytosol.
(D) Calcium is actively taken up from the extracellular environment and passively taken up from the cytosol.
18. Which statement best summarizes the role of calcium in skeletal muscle contraction?
(A) It prevents muscle contractions during action potentials.
(B) It changes the hypertonic nature of the cytosol to allow action potentials.
(C) It amplifies the physical contraction of the sarcomere.
(D) It connects the electrical neuronal signal to the actual physical contraction.
19. If a muscle cell fails to contract, which of the following could be a reason?
(A) The cell is lacking tropomyosin
(B) Too much calcium in the sarcoplasmic reticulum
(C) The cell is lacking troponin
(D) Too many action potentials reaching the cell
20. Dehydration synthesis is a key part of the creation of many macromolecules. It is best described as:
(A) Loss of a water molecule in order to make something else
(B) Water rushing out of a cell during the process of osmosis
(C) Life moving out of the ocean and becoming complex
(D) Kidneys filtering hydrophilic compounds during urine formation
21. A person rings a small bell every time a dog is fed. Over time, the dog begins to associate the jingle of the bell with eating and begins to salivate at the sound of the bell even if no food has been presented. This is an example of classical conditioning, a type of learning.
Imagine that you adopt a cat and discover that it has a strange fear of cotton balls. If this is a classically conditioned response, then which of the following could explain it?
(A) The cat has eaten cotton balls and become sick before.
(B) The cat sees cotton balls at the vet when it gets vaccinated.
(C) The cotton balls resemble a type of toy it had as a kitten.
(D) The cat is confused because it has never seen a cotton ball before.
22. A molecule of ADP is dephosphorylated once and then phosphorylated twice. What molecule will result?
(A) AMP
(B) ADP
(C) ATP
(D) AUP
Questions 23–25 refer to the following figure.
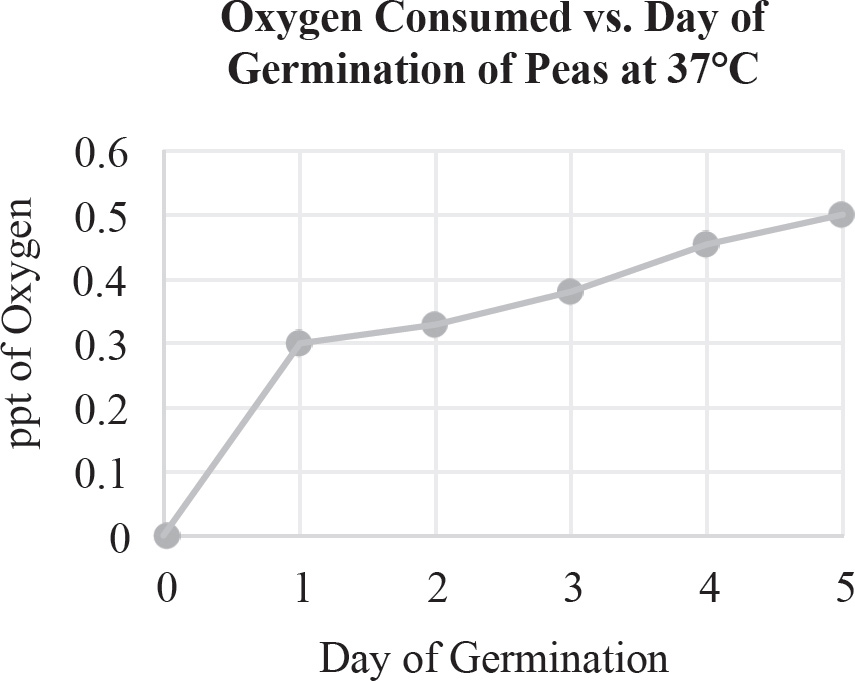
23. In the above figure, ______ is the dependent variable and _______ is the independent variable.
(A) ppt of oxygen consumed; day of germination
(B) ppt of oxygen consumed; 37°C
(C) day of germination; ppt of oxygen consumed
(D) 37°C; day of germination
24. In the above figure, which process is likely occurring?
(A) Photosynthesis
(B) Cellular Respiration
(C) Fermentation
(D) All of the above
25. If the trend continues, what will be the oxygen consumed on day 7?
(A) 0.55
(B) 0.6
(C) 0.75
(D) 0.8
26. There are two broad categories of how a cell dies. The first is called apoptosis, which occurs when the cell chooses to die and carries out a programmed cell death. The cell carefully disposes of itself and implodes in a way that causes the least amount of problems for neighboring cells. The other form of death is called necrosis. Necrosis is a messy form of cell death that is not carefully controlled. The cell often ruptures and causes inflammation and danger to the surrounding cells. Which of the following situations does NOT likely end with apoptosis?
(A) An error occurs during DNA replication causing severe chromosome breakage.
(B) A killer T-cell identifies a cell infected with a virus.
(C) A virus-infected cell reaches the lytic phase.
(D) A cell undergoes uncontrolled mitosis.
27. A neuron experiences two refractory periods after firing an action potential. One is called an absolute refractory period, which means that a neuron cannot fire under any circumstances because the voltage-gated sodium channels are inactive. The second type is a relative refractory period, which means that the cell is experiencing a state of hyperpolarization, making it unlikely to reach action potential threshold.
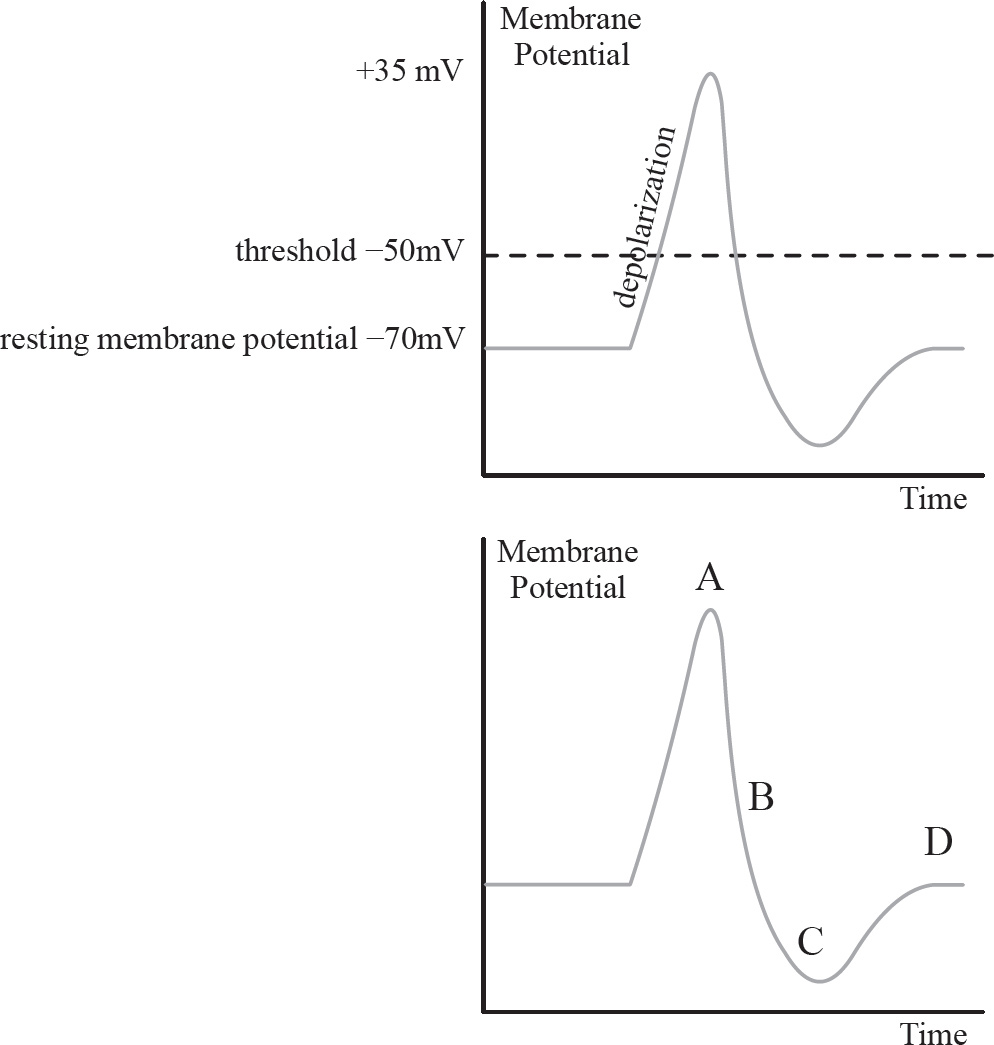
In the figure above, which of the following letters marks a relative refractory period?
(A) A
(B) B
(C) C
(D) D
28. During spermatogenesis, a single 2n spermatogonium undergoes meiosis to become:
(A) Two spermatids with 1n chromosomes
(B) Four spermatids with 1n chromosomes
(C) Two spermatids with 2n chromosomes
(D) Four spermatids with 2n chromosomes
29. Which is NOT a true statement about acid rain?
I. It increases the pH in the water and can harm aquatic plants.
II. High levels of H+ can harm fish hatchlings.
III. It can change the composition of the soil.
(A) I only
(B) I and II
(C) II only
(D) I and III
30. The Krebs cycle produces which of the following electron carriers?
(A) NADPH
(B) NADH
(C) FADH
(D) NAD+
31. Plant cells are well-known for having a structural dependence upon their large central vacuole. As this is a water-dependent structure, plants have developed many strategies for maintaining a state of hydration. One of these is a thick, waxy skin called a cuticle, which prevents water escaping from the plant surface. If a plant is misted with an enzyme designed to eat away the waxy cuticle, all of the following would be predicted outcomes EXCEPT
(A) the plant would not stand up as tall
(B) the plant would dry out more quickly
(C) the plant would grow more roots
(D) the plant would transport sugars more quickly
Questions 32–34 refer to the following passage.
Radiometric dating is a scientific technique based on predictable radioactive decay. The age of a rock or other substance that contains trace amounts of radioactive isotopes can be estimated by measuring how much of the original radioactive isotope is present and how much of the decayed version is present. Because the rate of decay occurs in an even, predictable manner, the original creation date of the rock can be estimated.
32. How does carbon-14 differ from the typical carbon, carbon-12?
(A) It formed a long time ago.
(B) It has shared electrons.
(C) It has more neutrons.
(D) It has half the atomic mass.
33. Which of the following assumptions does radiometric dating NOT make?
(A) The rock has not been in the presence of a strong magnetic field.
(B) The rock formed at the same time that the radioactive isotope began decaying.
(C) Neither the original isotope nor the decay product has escaped from the rock.
(D) The rate of decay is predictable and has not greatly changed over time.
34. Which of the following research questions could NOT be answered using radiometric dating?
(A) How many years ago did the planet Venus form?
(B) When did the last large dinosaurs go extinct?
(C) How old is the deepest layer in the Grand Canyon?
(D) How long have mastodon bones been buried in the permafrost?
Questions 35–37 refer to the following passage.
Diabetes mellitus is a disease characterized by an inability of the cells to properly produce (type I) or respond (type II) to insulin, a hormone produced by the pancreas in response to high levels of blood glucose. Without insulin, glucose accumulates in the blood. In situations of low blood glucose, another pancreatic enzyme, glucagon, is released, which triggers the process of gluconeogenesis shown on the right side of the pathway below. The stimulators, activators, or inhibitors of each step are shown with + or – signs.
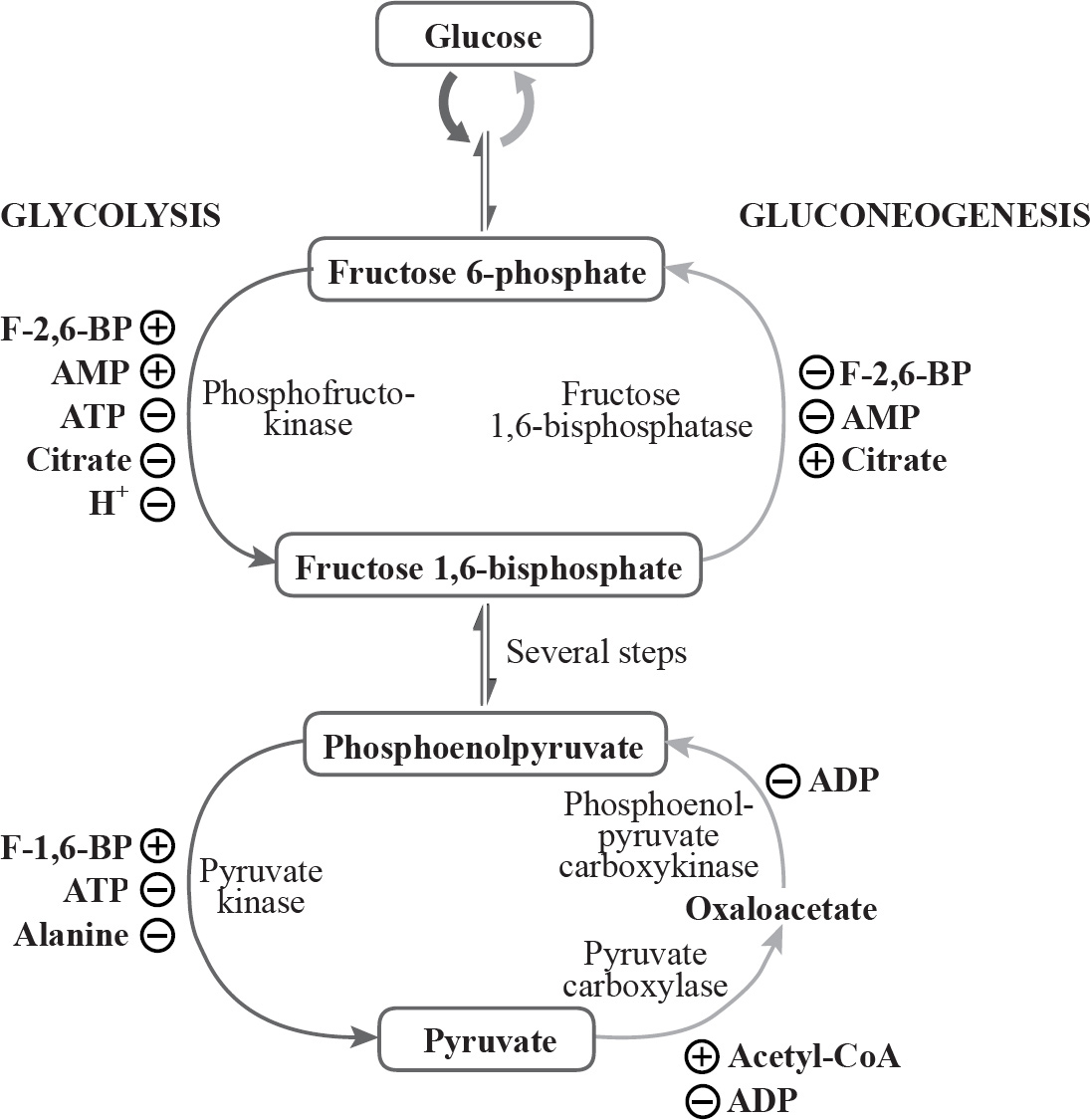
35. Which of the following conditions would lead to increased production of fructose 1,6-bisphosphate?
(A) High ATP and high citrate
(B) High AMP and high citrate
(C) High AMP and high F-2,6-BP
(D) High ATP and high F-2,6-BP
36. Patients with type I diabetes often require insulin injections. Which of the following situations would most require an insulin injection?
(A) After eating a stalk of celery
(B) After eating a cookie
(C) After skipping breakfast
(D) After drinking a lot of water
37. Which of the following situations likely stimulates gluconeogenesis?
(A) High levels of insulin
(B) High levels of F-2,6-BP
(C) High levels of glucagon
(D) High levels of ADP
Questions 38–39 refer to the following passage.
Embryogenesis is a carefully timed and well-organized process. As a single-celled zygote divides and grows into hundreds and thousands of cells, a process called differentiation occurs wherein certain areas of the embryo become specialized to become different types of tissue. As differentiation continues, the level of specificity increases, and the cell potency decreases until highly specialized unique tissues and organs develop. The figure below shows 12 stages of development of human embryos.
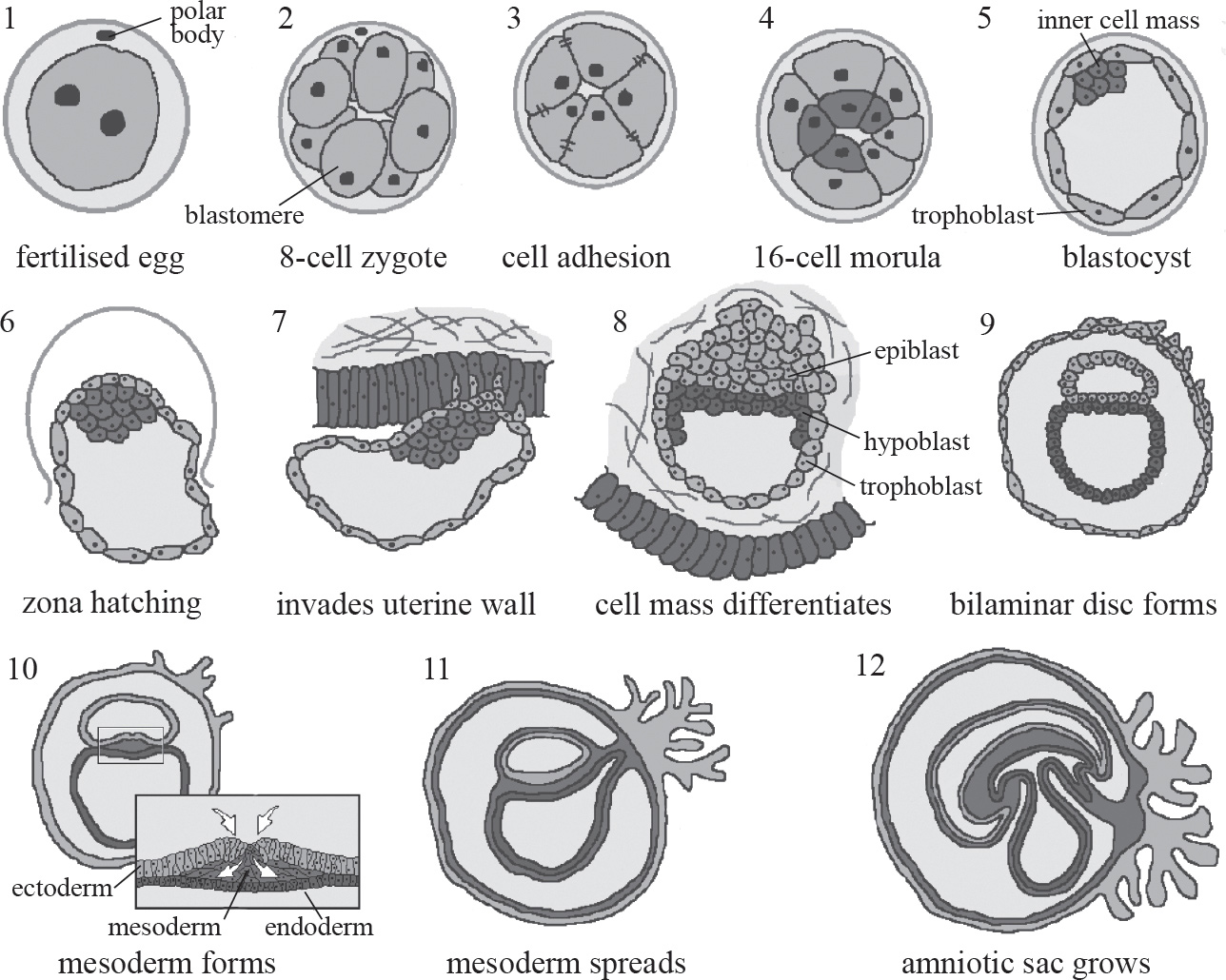
38. The inner cell mass is what eventually forms the embryo. During development, the embryo differentiates into various types of cell layers. Which of the following is not one of them?
(A) mesoderm
(B) hypoblast
(C) blastomere
(D) endoderm
39. A totipotent embryonic cell has the most cell potency. Which of the following is most likely to be totipotent?
(A) 8-cell zygote
(B) inner cell mass
(C) mesoderm
(D) digestive tract
40. The following diagram demonstrates the ecological succession that occurs in an environment over time as it is colonized by different species.
Why does it take 75 years for a beech-maple to occur in the figure above?
(A) Beech seeds have a very long period of dormancy prior to germination.
(B) It takes an average of 75 years for conifer trees to become extinct.
(C) Agriculture was the predominant industry, and hardwood trees were removed.
(D) Maple trees grow better in a pine forest than they do in a grassland.
41. During labor, pressure on the cervix and oxytocin form a positive feedback loop as shown below.
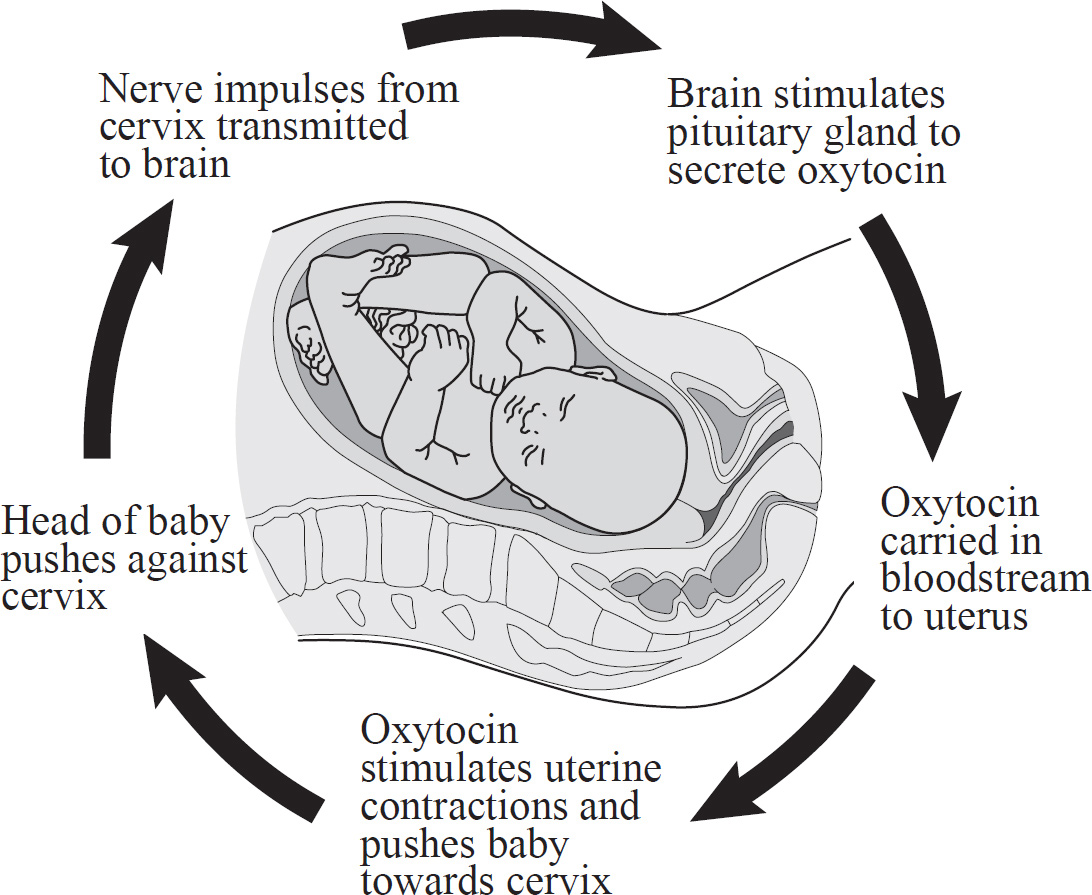
Which of the following other pathways also demonstrate positive feedback?
(A) Glycolysis leads to the production of ATP. ATP, in turn, turns off the enzyme phosphofructokinase, which catalyzes a key phosphorylation step in glycolysis.
(B) The anterior pituitary gland in the brain releases adrenocorticotropic hormone (ACTH). ACTH then causes the adrenal cortex to release glucocorticoids. Glucocorticoids then prevent the pituitary from releasing more ACTH.
(C) Lutenizing hormone triggers ovulation and the formation of the corpus luteum, which is a hormone-producing structure formed during ovulation. The corpus luteum secretes progesterone, which inhibits LH. The drop in LH causes the degradation of the corpus luteum.
(D) When a tissue is injured, it releases chemicals that activate platelets. Activated platelets themselves then release chemicals that activate more platelets. These activate platelets then release chemicals to activate more platelets.
42. The figure below is a typical chemical synapse. The table below shows the measurements of 4 different synapses. The presynaptic membrane potential and the postsynaptic membrane potential are measured before the neurotransmitter is released and after the neurotransmitter is received.
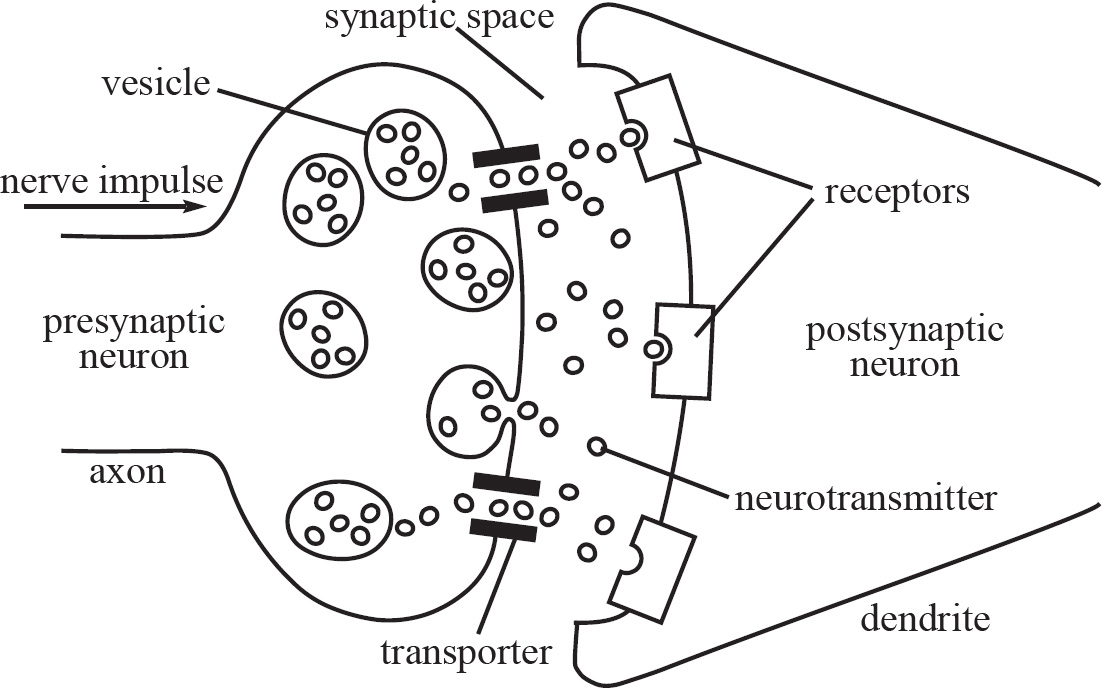
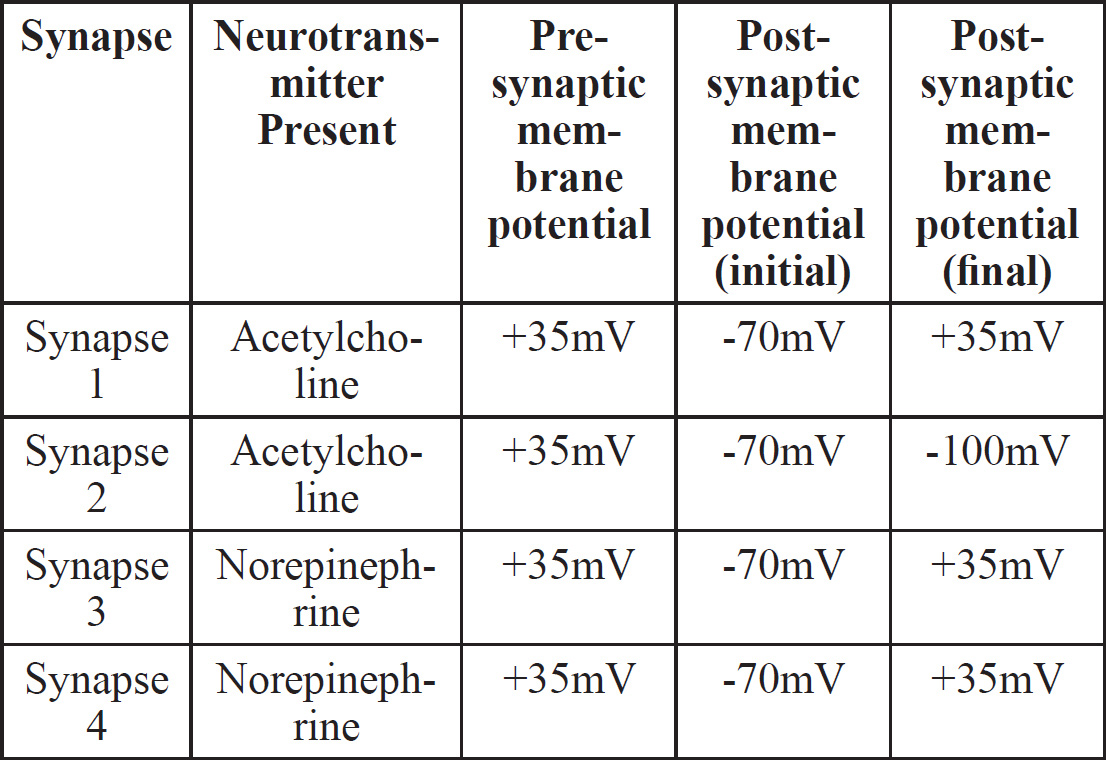
What is the best explanation for the varying behavior of acetylcholine on the postsynaptic membrane potential in synapse 1 and synapse 2?
(A) There might be acetylcholinesterase enzyme within synapse 1.
(B) The receptors on the postsynaptic cell determine the effect of the neurotransmitter.
(C) The action potential at the presynaptic cell is not always large enough to pass on.
(D) All of the above
Questions 43–46 refer to the following passage.
The blood of two patients is mixed independently with 3 different antibodies (A, B, and C) that are known to react to 3 diseases (1, 2, and 3 respectively). The results of the antibody binding are shown in the figure below.
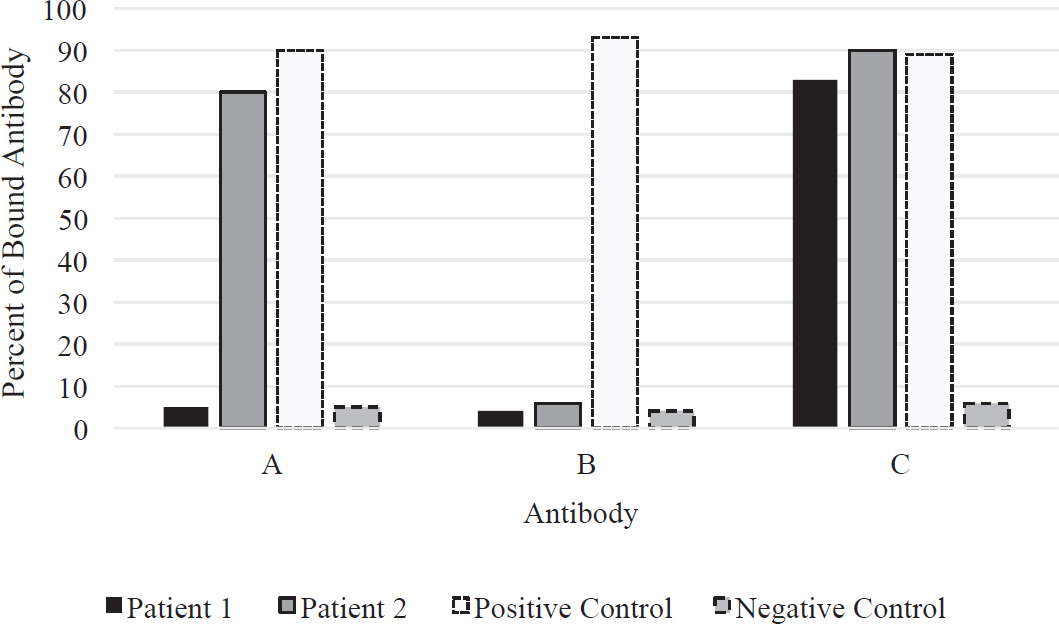
43. Which question is best answered by this experiment?
(A) Has either patient 1 or 2 been infected in the past with disease 1, 2, or 3?
(B) Do either of the patients have antibodies A, B, or C in their blood?
(C) Are either patient 1 or patient 2 currently infected with disease 1, 2, or 3?
(D) Has either patient 1 or patient 2 ever made antibodies A, B, or C in the past?
44. The low bars for antibody B represent each of the following EXCEPT:
(A) Neither patient currently has disease 2 in their blood
(B) A low level of non-specific binding occurs by antibody B
(C) Neither patient currently has antibody B in their blood
(D) Antibody B did not specifically bind to the patient’ blood
45. What is likely in the positive control?
(A) Blood containing diseases 1, 2, and 3
(B) Blood containing antibodies A, B, and C
(C) Blood without disease 1, 2, and 3
(D) Blood without antibodies A, B, and C
46. If this study was done a year from now, which graph would likely result for antibody B?
(A)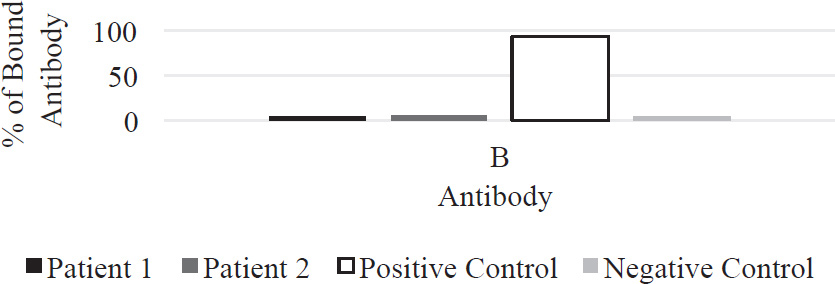 
(B) 
(C) 
(D) This is impossible to predict with the information given
47. The following graph demonstrates 3 different strategies for survival.
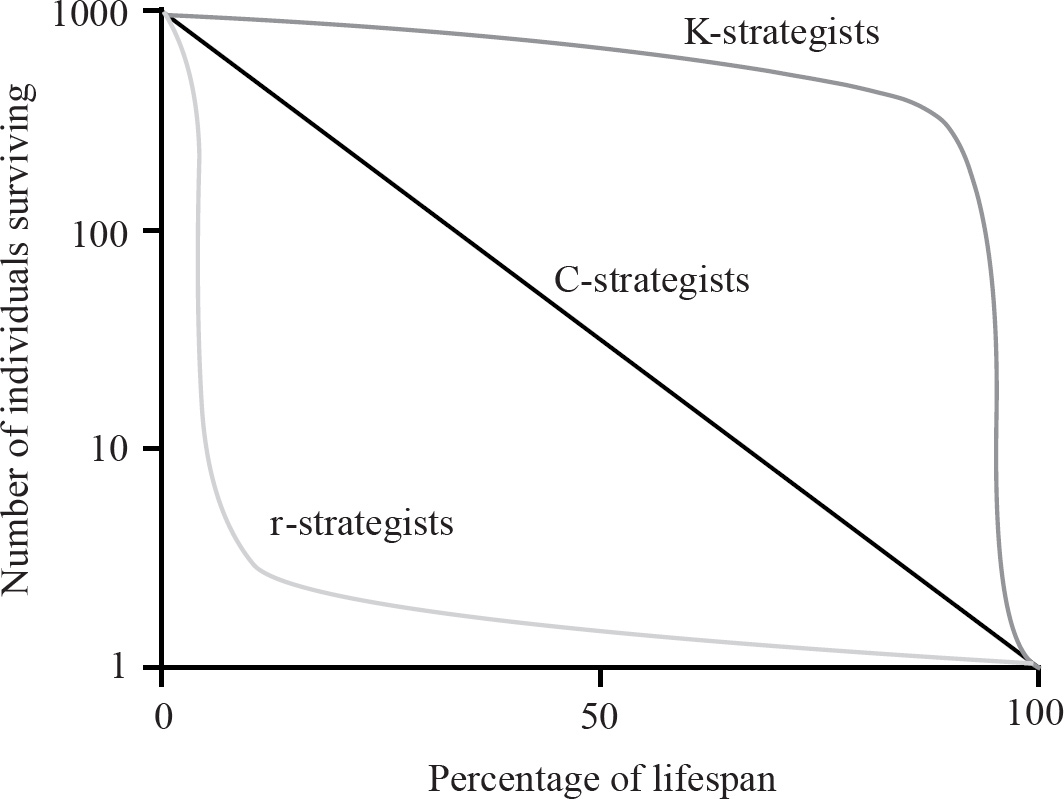
Which of the following are examples of r-strategists?
I. Human
II. Spider
III. Beluga whale
(A) I only
(B) II only
(C) I and III
(D) I, II, and III
Questions 48–49 refer to the following figure.
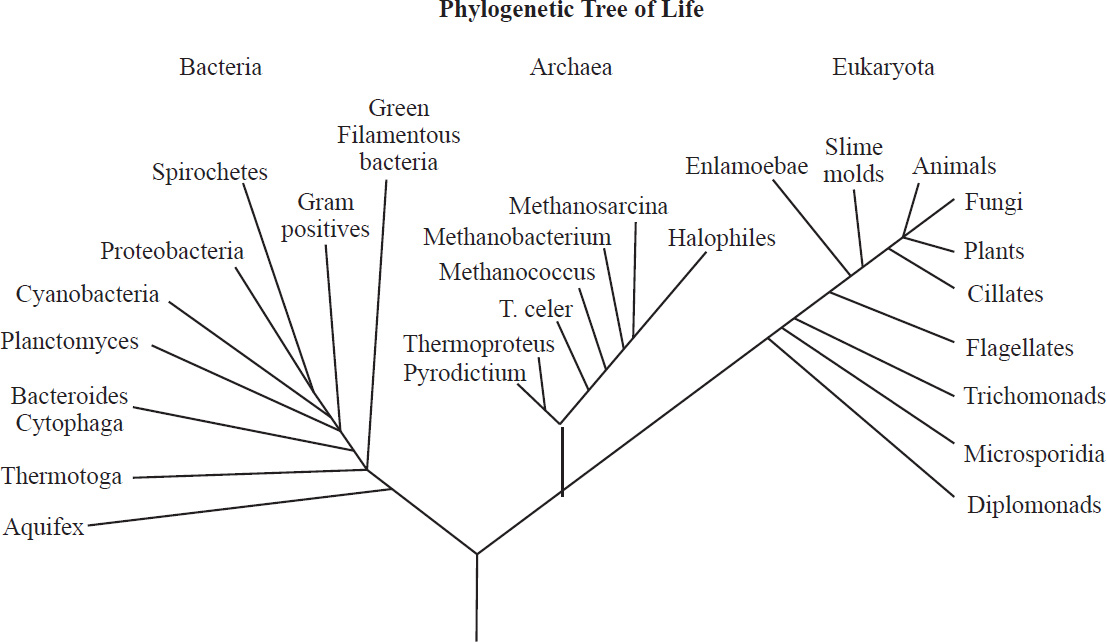
48. Which of the following are the most closely related?
(A) Aquifex and diplomonads
(B) Animals and fungi
(C) Halophiles and entamoebae
(D) There is not enough information given.
49. A planctomyces is dividing by budding, which of the following will occur?
(A) A mitotic spindle will pull chromosome segments to opposite ends of the cell.
(B) Enzymes will unwind the helix and copy the entire bacterial genome.
(C) The nuclear envelope will break down and then reform after DNA replication.
(D) The mitochondria will replicate and be divided between the two cells.
50. Which of the following would make the Calvin cycle unnecessary?
(A) If the light-dependent reactions made sugar and ATP
(B) If plants could make ATP in their electron transport chain
(C) If plants could use ATP to power cellular processes
(D) If NADPH could be created by photosystem I
51. Without phototropism how would plant growth be affected?
(A) Plants would never point towards the sun.
(B) Plants would not be able to change direction of growth.
(C) Plants would not be able to perform photosynthesis.
(D) The direction of plant growth would not be related to the sun.
52. Which of the following does NOT affect higher order protein structure?
(A) Ester linkages between amino acids
(B) Amino acid sequence order
(C) Hydrogen bonding between R groups on the same polypeptide
(D) Hydrophobic interactions between side groups on different polypeptides
Questions 53–54 refer to the following passage.
The following study was carried out to examine sexual selection of beetles. Chemicals believed to be pheromones were isolated from males of certain species of butterfly. An environment was created where a pheromone-containing droplet was applied to one side of the box and a control drop was applied to the other side. Female butterflies were introduced to the center of the box and given the opportunity to go to either side. The results are shown in the table below.
| Preferred Side | # of Butterflies |
| Male chemical side | 2760 |
| Control side | 2240 |
53. What is the best testable null hypothesis for the above experiment?
(A) Female butterflies prefer the scent of chemicals produced by males of their own species over the control chemicals.
(B) Female butterflies prefer the scent of the control chemicals over chemicals produced by males of their own species.
(C) Female butterflies cannot sense the chemicals produced by males of their own species.
(D) Female butterflies have no preference for either of the chemicals.
54. A chi-squared analysis would be performed in this experiment to make which of the following conclusions?
(A) To calculate that there is 1 degree of freedom
(B) To determine if the null hypothesis can be rejected
(C) To determine the standard deviation between two samples
(D) To prove that the working hypothesis is correct
Questions 55–56 refer to the following passage.
Two DNA sequences are shown below.
Sequence 1
5′ GATTCCTACATCAG 3′
3′ CTAAGGATGTAGTC 5′
Sequence 2
5′ CGGCGAGACGCGGC 3′
3′ GCCGCTCTGCGCCG 5′
55. If the mRNA sequence transcribed from one of the above sequences is 5′ GAUUCCUACAUCAG 3′, what is the sequence of the coding strand of DNA?
(A) 3′ CTAAGGATGTAGTC 5′
(B) 5′ GACTACATCCTTAG 3′
(C) 5′ GATTCCTACATCAG 3′
(D) 3′ CTGATGTAGGAATC 5′
56. Which of the following best describes the relationship between sequence 1 and sequence 2?
(A) Sequence 1 is less likely to be a coding sequence.
(B) Sequence 2 is more likely to degrade over time.
(C) Sequence 1 has more hydrogen bonds between base pairs.
(D) Sequence 2 has a higher melting temperature.
57. Microvilli provide what benefit to organisms?
(A) Aiding in locomotion in unicellular organisms
(B) Increasing surface area, which increases absorption efficiency
(C) Secreting hormones necessary to maintain homeostasis
(D) Clearing dust and pathogens from mucus membranes
58. Blood pressure is determined by the volume of the blood and the peripheral resistance of the blood vessels. Blood volume is dependent upon hydration level and osmotic pressure within the blood. Peripheral resistance refers to the volume of blood vessel, which is dependent upon constriction and dilation of arteries as blood flow is maximized and minimized in response to the body’s needs.
Which of the following would BOTH help to raise blood pressure?
(A) Constriction of blood vessels and decreasing salt intake
(B) Dilation of blood vessels and decreasing salt intake
(C) Constriction of blood vessels and increasing salt intake
(D) Dilation of blood vessels and increasing salt intake
Questions 59–61 refer to the following passage.
Hydrogen peroxide (H2O2) is broken down into oxygen and water by the enzyme horseradish peroxidase (HRP). This reaction can be measured with an oxygen-sensitive color indicator, and the color change can be measured on a spectrophotometer. The rate of oxygen formation is measured with a constant HRP concentration and 5 different concentrations of H2O2 at room temperature. A follow-up experiment adds a known inhibitor of HRP (sodium azide). The results of both studies are shown in Figure 1.
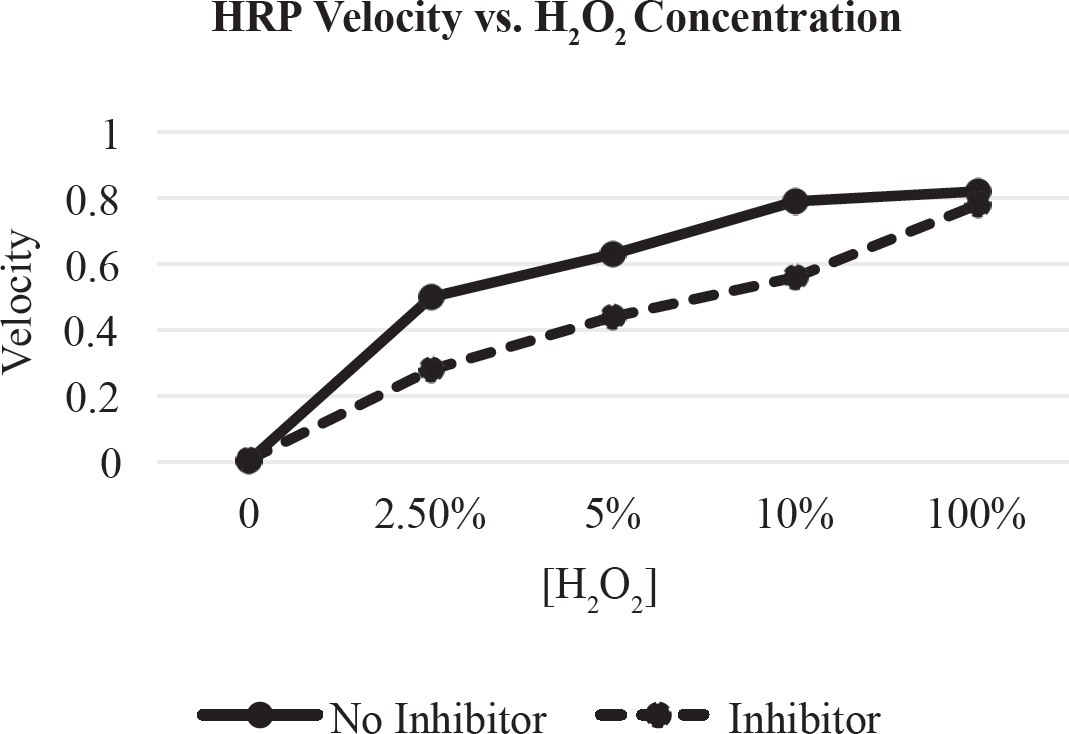
59. What is the substrate in this reaction?
(A) H2O2
(B) HRP
(C) water
(D) oxygen
60. Which of the following hypotheses about why the no-inhibitor graph plateaus after 10% H2O2 is supported?
(A) Water begins to act as an inhibitor.
(B) All the HRP is fully engaged.
(C) There is no more H2O2 available.
(D) The temperature is no longer optimal.
61. Which statement about sodium azide is supported by the data?
(A) It performs best at room temperature.
(B) It binds to the active site on HRP.
(C) It binds to an allosteric site on HRP.
(D) It has no effect on the reaction.
Questions 62–63 refer to the following passage.
Interactions between biotic and abiotic factors are what makes an ecosystem stable. Some interactions are obvious, but others occur underground beyond our sight. Mycorrhizae are a type of symbiotic relationship between fungi and plant roots. Special growth patterns allow the two species to easily share water, nutrients, and sugars. The plant is better at making sugar than the fungi, and the fungi have lots of tiny little strands, which provide the plant a huge surface area.
62. How might the fungi be helping the plant?
(A) Preventing the plant from touching the soil
(B) Giving it more capacity to suck up water
(C) Decreasing the osmotic pressure in the root
(D) Assisting in the light-independent reactions of photosynthesis
63. If the plant devised a way to block sugar transport to the fungi, the relationship could best be described as
(A) mutualistic
(B) parasitic
(C) commensal
(D) individual
Directions: Part B consists of questions requiring numeric answers. Calculate the correct answer for each question.
64. A species of butterfly comes in two colors: blue ((B) and white (b). In a population of 500 butterflies at Hardy-Weinberg equilibrium, there are 300 blue butterflies. What is the frequency of the recessive allele? Answers should be rounded to 1 decimal place.

65. In the following mRNA, what it the length of the 5′ untranslated region in bases?
5′ UAACGAUCAUAUAUGACGCGUAUCUAG 3′

66. A homozygous, dominant male and a homozygous, recessive female are crossed. One of the male offspring is backcrossed to the female. What percentage of their offspring will be heterozygous?

Questions 67–68 refer to the following figure.
67. What is the change in Gibbs free energy for the reaction?

68. What is the activation energy for the equation?
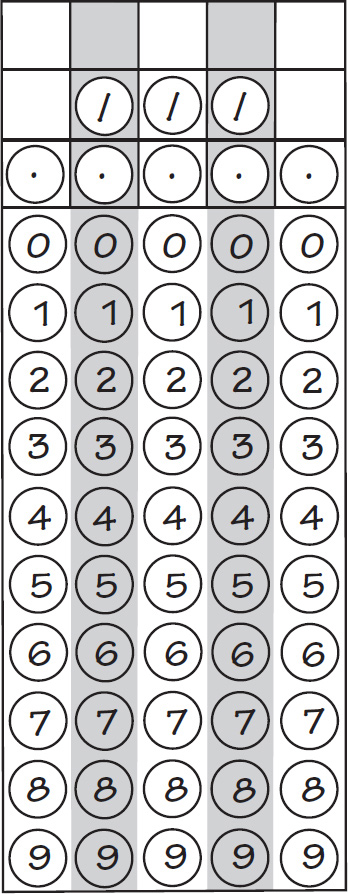

69. To determine whether pillbugs prefer wet or dry conditions, you create a choice chamber that measures 400 cm2. Half of the chamber is covered with wet soil and half of the chamber is covered in dry soil. You place 10 pillbugs at the center line of the choice chamber. After 15 minutes, you count how many pillbugs are on each side. After 3 trials, you find that an average of 8.8 pill bugs choose the wet side and an average of 1.2 pill bugs choose dry side.
What is the chi-square value for this experiment (round to one decimal place)?

STOP
END OF SECTION I
IF YOU FINISH BEFORE TIME IS CALLED, YOU MAY CHECK YOUR WORK ON THIS SECTION. DO NOT GO ON TO SECTION II UNTIL YOU ARE TOLD TO DO SO.
BIOLOGY
SECTION II
8 Questions
Planning Time—10 minutes
Writing Time—80 minutes
Directions: Questions 1 and 2 are long free-response questions that should require about 22 minutes each to answer and are worth 10 points each. Questions 3 through 8 are short free-response questions that should require about 6 minutes each to answer. Questions 3 through 5 are worth 4 points each, and questions 6 through 8 are worth 3 points each.
Read each question carefully and completely. Write your response in the space provided following each question. Only material written in the space provided will be scored. Answers must be written out in paragraph form. Outlines, bulleted lists, or diagrams alone are not acceptable.
1. A bacterial transformation is a way to introduce new DNA into bacteria. Using special equipment, the bacteria are made permeable and will uptake plasmid DNA from their surroundings. This plasmid DNA can be expressed within them in the same way as their own bacterial chromosome.
You are given three types of bacteria (A, B, and C), unlimited plates of agar, the antibiotics tetracycline and ampicillin, and plasmids with either the tetracycline or ampicillin genes for antibiotic resistance. Assume that you also have the proper equipment to grow bacteria and carry out bacterial transformations.
(a) Design an experiment to determine the antibiotic resistance of each type of bacteria. Be sure to use proper controls.
• Describe your methods.
• Show mock data in a graph or table.
• Justify your conclusion using the data as a reference.
(b) Design a follow-up experiment to test if antibiotic resistance can be genetically transferred.
• Describe your methods.
• Show mock data in a graph or table.
• Justify your conclusion using the data as a reference.
2. Homeostasis is the steady-state that the human body strives to maintain.
(a) Explain why homeostasis is important in the body and include at least THREE physiological variables that must be maintained.
(b) Contrast homeostasis in a unicellular organism with homeostasis in a complex organism.
(c) Calcium is a very important ion in the body. The hormones parathyroid hormone (PTC) and calcitonin are released when calcium levels are low and high, respectively, to help the body return to its homeostatic state. Describe the likely effects of each PTH and calcitonin in the body with respect to:
• intestinal absorption rates (collection of calcium from the diet)
• kidney reabsorption rates (retaining calcium in the blood during blood filtration instead of excreting it in urine)
• the production of osteoblasts (bone builders) and osteoclasts (bone destroyers)
3. Describe the general process of photosynthesis, including reactants, products, and the different objectives of the light-dependent and the light-independent reactions.
4. The vertebrate immune system consists of two parts: innate immune response and cell-mediated specific immune response. Briefly explain how each of them would work together to combat a first-time exposure infection.
5. Explain why a universal energy molecule is essential in cells. Give examples of energetically unfavorable reactions/processes and explain how they can be powered.
6. The following graph shows the shapes of a certain protein found in the cytosol.
Explain how two shape outcomes can be possible. Be sure to mention DNA replication, transcription, and translation in your response.
7. Describe the process of natural selection and give an example. Explain how genetic drift differs from natural selection.
8. Draw a food energy pyramid and explain why 90% is estimated to be lost at each ascending level.
STOP
END OF EXAM
Practice Test 3: Answers and Explanations
PRACTICE TEST 3 ANSWER KEY
1. A
2. D
3. C
4. D
5. A
6. C
7. B
8. B
9. A
10. B
11. B
12. D
13. C
14. A
15. C
16. C
17. C
18. D
19. C
20. A
21. B
22. C
23. A
24. B
25. B
26. C
27. C
28. B
29. A
30. B
31. D
32. C
33. A
34. A
35. C
36. B
37. C
38. C
39. A
40. D
41. D
42. B
43. C
44. C
45. A
46. D
47. B
48. B
49. B
50. A
51. D
52. A
53. D
54. B
55. C
56. D
57. B
58. C
59. A
60. B
61. B
62. B
63. C
64. 0.2
65. 12
66. 50%
67. −30
68. 50
69. 5.8
PRACTICE TEST 3 EXPLANATIONS
Section I: Multiple Choice
1. A Since Klinefelter’s is a disease in males, this means the person will have a Y chromosome. Because it involves the creation of a Barr body, this also means that the person will have an extra X chromosome. The answers show karyotypes, which display a person’s chromosomes. The 23rd pair is the sex chromosomes. Choice (A) is the only one with a Y and two X chromosomes.
2. D If BAMHI cuts between the guanines in the sequences 5′ GGATCC 3′, then each answer choice should be checked for that sequence. Choice (A) has one cut site, resulting in two chunks. Choice (B) has three cut sites, resulting in four chunks. Choice (D) has two sites, resulting in three chunks. Choice (C) has zero cut sites because the cut sequence is reversed from the correct 5′ → 3′ direction.
3. C Green pea plants can result from either a homozygous dominant (GG) or a heterozygous (Gg) phenotype, so your mystery pea plant could be either. You need to figure out which it is. If you cross it with any green plant (A), you won’t know what genotype you are crossing it with. If you cross it with a homozygous dominant plant (B), you will always get green plants no matter what your original mystery plant was. If you cross it with a yellow plant, then you will know it is a homozygous recessive. If your original plant is a homozygous dominant, you will only get green plants. If your original plant is a heterozygote, then you should get yellow plants 50% of the time.
4. D The passage says the parental plants crossed are true breeding, so they are each homozygous. That cross should result in only green plants in the F1 generation. Crossing over (A) will not give variation because the parental plants are homozygous. Their two copies are identical and crossing over would just swap identical genes. The law of independent assortment (B) explains mixing of maternal and paternal versions of the different chromosomes. The two homozygous dominant pea color alleles are on the two copies of the same chromosome. A true breeding plant would also be unlikely to have gene alleles on other chromosomes that impact the color. Choice (D) is correct because mutation can occur at any time and result in changes to phenotype.
5. A A yellow plant must be homozygous recessive (gg). All plants in the F1 generation are heterozygotes (Gg) because the parents are a homozygous dominant (GG) and a homozygous recessive (gg). As shown in the Punnett square below, the ration of green : yellow would be 1:1.
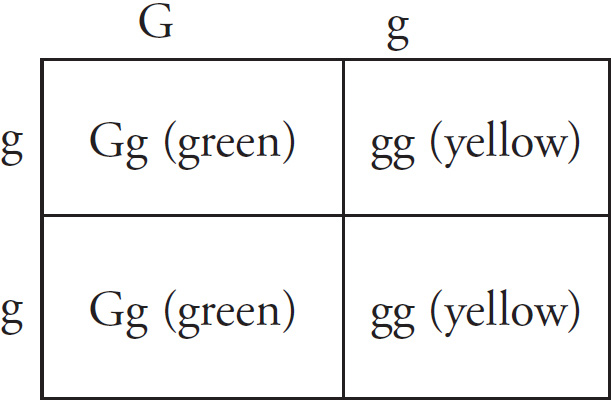
6. C The figure shows that a sensory neuron collects information from the periphery and brings the signal to the brain. You know that neurons receive information through their dendrites and pass it on with their axons. Choice (A) represents a neuron receiving signal from the brain and passing it to a muscle (This is a motor neuron.). Choice (B) is a neuron that collects a stimulus and sends it to a muscle cell (This type of neuron does not exist.). Choice (C) is a neuron that gets a stimulus and passes it to the brain. Choice (D) is a neuron going from brain to spinal cord, which are both part of the central nervous system (This is an interneuron.).
7. B Gas exchange across several cells must involve passing of things from cell to cell. Gap junctions are described as allowing action potentials to be passed from cell to cell. The other types of junctions all create a barrier or tension.
8. B G1 is before DNA replication occurs. This means that there should be 32 unreplicated chromosomes. After replication there would be 64 segments since each chromosome would have a sister chromatid.
9. A The passage says that humans have 46 chromosomes total. Humans are diploid and have two copies of each of the non-homologous chromosomes. Thus, humans have 23 unique, non-homologous chromosomes. If the species has 32 chromosomes and they are octoploidy, then they have 8 copies of each. So, they only have 4 unique chromosomes.
10. B The passage says that two gametes come together to make an octoploidy (8n) species. Therefore, each gamete should be tetraploidy (4n). If they have four non-homologous chromosomes, then 4 × 4 = 16.
11. B The peak in 1958 is in the middle, corresponding to medium-orange fish. However, there are also some fish on each extreme (light or dark).
12. D These graphs of phenotype tell us nothing about the mode of inheritance.
13. C If a dark-colored, poisonous fish existed, predators would soon learn to avoid dark-colored fish. The dark-colored fish would have an advantage. With no other information known about this ecosystem, the assumption is that the dark fish will become more prevalent. Choice (C) is the only one that has the right side of the graph (shown above to represent the dark fish) increased.
14. A From 1958 to 2008, the graph shifted away from medium-orange and moved towards the extremes (light and dark). Choice (A) would make less orange fish be eaten, which would promote more medium-orange fish. Choices (B) and (C) would cause less light and dark fish to be eaten, respectively. This would promote the light and dark fish. Choice (D) would cause the medium-orange fish to be eaten more often and thus promote the light and dark extremes shown in 2008.
15. C Transmembrane domains are in the hydrophobic space and must not be polar. Of the amino acids shown above, glutamate is negatively charged and lysine and arginine are positively charged. Isoleucine (C) is the only non-charged choice.
16. C Enzyme active sites are the places where the substrate binds. If the active site contains positively charged residues, the substrate should have a negative charge because the active sites and substrate would attract each other.
17. C If calcium is prevalent outside the cell, then it will passively flow into the cell (as long as a calcium channel is open) because it wants to move down its concentration gradient. The passage states that the sarcoplasmic reticulum actively takes up calcium.
18. D Calcium links the action potential to contraction of the sarcomere (D). It stimulates contraction, it does not prevent it (A). It is not the effect on hypertonicity that causes the action potentials (B); action potentials are calcium specific. Calcium just relays the action potential message, it does not amplify the signal (C).
19. C Without troponin, the calcium could not move tropomyosin off the myosin binding sites, and then there would be no contraction. If there were no tropomyosin, then the contraction could occur at any time, regardless of calcium. An excess of calcium in the sarcoplasmic reticulum (B) or action potentials (D) would also lead to contraction, not a lack thereof.
20. A Dehydration is loss of water. Synthesis is building something. This is a common type of reaction during the polymerization of macromolecules.
21. B Classical conditioning pairs together two things that are not directly related. In this instance, the cat has paired together cotton balls with something else that causes it to have fear. Choice (A) would be a direct response to cotton balls, so this is not classical conditioning. Choice (C) would also be a direct relation to a bad experience with the cotton balls. Choice (D) would not be a type of learning if it has never seen them before.
22. C ADP has two phosphates. If it is dephosphorylated, then it would be AMP, which has one phosphate. If it gets two added, then it would be ADP and then ATP.
23. A The x-axis is typically the independent variable, which is something set by the experimenter. The dependent variable is the thing that is measured as data. So germination is the independent variable, and ppt of oxygen is the dependent variable. The temperature is a constant in this instance as every point on the graph is meant to show germination at that given temperature. Often, temperature will be the independent variable in an experiment.
24. B The graph shows that oxygen consumed starts at 0 and then increases. In photosynthesis, oxygen is released. In fermentation, there is no oxygen present. This must be cellular respiration, which requires oxygen.
25. B The trend over the 4 points that seem to fall on a line averages 0.1 every 2 days. This would increase it to 0.6 on day 7.
26. C The question is asking for the situation that is NOT apoptosis. Choices (A), (B), and (D) are all situations where the body can step in and initiate apoptosis to eliminate the bad cell. Choice (C) will lead to a messy death by necrosis when the cell is lysed by the virus.
27. C The relative period is when the cell is hyperpolarized. This means it is even more negative than usual and further from the threshold position.
28. B Spermatogonia (2n) undergo meiosis to become four haploid (1n) cells. Meiosis replicates the DNA and then splits it twice to result in 4 cells each with half the DNA.
29. A Acid rain has high H+ and thus a low pH. Therefore, it would NOT increase the water pH (I). However, these high H+ levels can harm fish (II) and change soil composition (III).
30. B The Krebs cycle reduces NAD+ to NADH. Therefore, NADH is produced. It also produces the electron carrier FADH2.
31. D The waxy cuticle helps prevent water loss. Water is important in the plant for maintaining a turgid structure. The plant would not likely stand as tall (A), it would dry out rapidly (B), and it would grow more roots to seek out more water (C). This would not likely cause increased sugar transport (D).
32. C An isotope is an element with a differing number of neutrons. Carbon-14 has two more neutrons.
33. A Radiometric dating assumes that the isotope started decaying when the rock was formed (B). It also assumes the full amount of the isotope or the decay product is there, so scientists can compare the levels of each (C). The measurement of decay is assumed to predictable, which is why it can be used like a stopwatch (D). The presence of a magnetic field is not relevant, and anything on Earth is always in the presence of a strong magnetic field.
34. A In order to use radiometric dating, a sample of something that contains radioisotopes that have been decaying is necessary. We have no samples of planet Venus to test, therefore the formation cannot be measured this way.
35. C ATP inhibits its production since it inhibits phosphofructokinase. This eliminates choices (A) and (D). Citrate stimulates turning fructose 1,6-bisphosphate into fructose 6-phosphate, which eliminates (B). Choice (C) includes two things that stimulate phosphofructokinase and lead to production of fructose 1,6-bisphosphate.
36. B Insulin is needed when blood glucose gets too high. Eating a cookie would increase blood sugar more than eating celery, fasting, or drinking water.
37. C Gluconeogenesis is the formation of glucose. If the body had high insulin then it would already have lots of glucose, so (A) is eliminated, but if the body had high glucagon, it would need glucose (C). High ADP would inhibit gluconeogenesis as shown in the figure (D). High F-2,6-BP would also inhibit gluconeogenesis (B).
38. C The blastomere is shown prior to the inner cell mass; therefore, it cannot be a differentiation of the inner cell mass. The hypoblast comes from the inner cell mass, and the endoderm and mesoderm also come from it.
39. A Potency is the potential to become many things. The most potent cells are the least differentiated or the least specialized. The more specialized they become, the more they lose their potency. The 8-cell zygote is the least specialized and has the most potency.
40. D Few species can grow in barren areas, but once they are colonized with plants, then more species can grow and live there. It takes a long time for the beech and maple trees to grow because first species first need to change the environment, allowing other species to colonize. The environment was not hospitable until it was a pine forest, which took a long time to reach as well.
41. D Positive feedback is when a process creates an end product whose production stimulates the process to create even more of the end product. In Choice (D), the platelets lead to more platelet creation. Choice (A) is negative feedback. Choice (B) is negative feedback. Choice (C) is negative feedback.
42. B Each of these synapses changed from −70mV, but they changed in different directions. Synapse 1 depolarized to +35mV, and synapse 2 hyperpolarized to −100mV. If there were not enough action potentials this doesn’t explain one being depolarized and the other hyperpolarized and (C) is eliminated. Acetylcholinesterase would degrade acetylcholine, and without acetylcholine as a neurotransmitter the post-synaptic membrane potential would be unchanged before and after the action potential so (A) is eliminated.
43. C The experiment shows how patients’ blood reacts with antibodies. Therefore, the patients’ blood is being tested for the antigens that react with those antibodies. Choices (A), (B), and (D) would only work if the patients’ blood were being tested for antibodies.
44. C Low bars for patients 1 and 2 show they don’t have disease 2 antigens in their blood (A). The reason they have bars at all is because of non-specific binding that is considered a background level (B). This is the same thing as saying antibody B did not bind specifically to the patients’ blood (D). Since the test is not for antibodies in the blood, choice (C) is not something that is represented by the low bars.
45. A A positive control is something that shows that the experimental procedure is working as it should. Since this test is for the antigen, then the positive control should have the antigen.
46. D We do not know what antigens will be in the patients’ blood a year from now. It is not possible to predict if they will get those diseases and be unable to clear them from their blood sometime in the next year.
47. B Humans and beluga whales have few young. They take care of them, ensuring most survive to adulthood. Spiders have many young, but they are abandoned and many of them don’t make it. Even if you don’t know how whales reproduce, you should know that spiders lose lots of their young and humans don’t. This makes (II) valid and (I) not. The only option left is (B).
48. B Animals and fungi might seem different, but in the grand scheme of life, they are not so distant at all. They branched away from each other most recently on the phylogenetic tree.
49. B Planctomyces are on the bacterial branch of the tree. Bacteria are prokaryotes and do not have segments of chromosomes because they have one circular chromosome. They do not need a mitotic spindle during division (A). They also do not have a nucleus (C) or mitochondria (D).
50. A The Calvin cycle converts the ATP and NADPH into sugar. If the light-dependent reactions made sugar and ATP, then the Calvin cycle would be unnecessary. Plants already do make ATP from the gradient of their electron transport chain (B). They also use ATP in their cellular processes (C). NADPH is made by photosystem I, and it still needs the Calvin cycle.
51. D Phototropism causes plant growth to change based on the sun. Without it, plants would grow randomly. They would sometimes point toward the sun (A), but this behavior would be random and would not be due to phototropism. Plants would still be able to perform photosynthesis (C) and they could even change their direction, but it would not be due to the sun. For example, instead, they might display thermotropism or hydrotropism (B).
52. A Higher order protein structure depends on the original amino acid sequence (B), the interactions between the ride chains of the amino acids (C), and the interactions between other polypeptides (D). Ester linkages are not the linkages between proteins.
53. D The null hypothesis would be that butterflies show no preference for either of the sides and the presence of the pheromone has no effect. The working hypothesis is that the females prefer the pheromones. If females cannot sense the chemicals, that would explain a possible reason the null hypothesis turns out to be true, but that is not the null hypothesis.
54. B Chi-squared analysis can only prove something is unlikely; it cannot prove something is correct. There is one degree of freedom, but this is a fact and is not the conclusion of a chi-squared analysis. Standard deviation is a different statistical test.
55. C The coding strand should always be the same as the mRNA except for the thymine in DNA and the uracil in RNA. mRNA is the complement of the template strand of DNA, not the coding strand.
56. D Although the previous question showed that part of sequence 1 is a coding sequence, this does not mean it is more likely to be one than sequence 2. The two sequences differ by the composition of their sequences. Sequence 2 has more guanines and cytosines. The G-C bond has 3 hydrogen bonds between it rather than the 2 hydrogen bonds that the A-C bond has. This gives us a higher melting temperature because it is more difficult to break the strands apart.
57. B The microvilli in the small intestine increase the surface area of the small intestine so that it can absorb nutrients more effectively. Cilia clear dust and pathogens. Cilia and flagella help unicellular organisms to move. The small intestine secretes some hormones. However, these hormones are not necessarily for homeostatis, and their production is not the function of the microvilli.
58. C Blood pressure is increased when the space gets smaller (constriction of blood vessels) or when the volume gets bigger. When the blood gets too salty, it takes on water to dilute the salt and the volume increases. So, constricting vessels and increasing salt intake will increase blood pressure.
59. A The substrate is H2O2 because that is what the HRP enzyme acts upon. Water and oxygen are the products.
60. B The substrate increases from 10% to 100%, so we know the substrate is still available. However, it is the enzyme that remained constant. The temperature remains constant. There is also no evidence that the water product has become an inhibitor.
61. B Sodium azide is an inhibitor. The graph shows that more substrate is needed, but eventually it can reach the same speeds as the uninhibited graph. This means that it is a competitive inhibitor that competes with the substrate for the active site (B) and not an allosteric site (C). We don’t know if it performs best at room temperature (A). We also know that it has an effect on the reaction (D).
62. B The fungi are increasing the surface area, which would increase the water absorption of the roots. Maybe they prevent the plant from touching soil, but this would not necessarily help the plant in an obvious way.
63. C The plant does not give anything to the fungi, but the fungi still help the plant get water. This would be a commensal relationship because the fungi are not harmed but they do help the plant collect water.
64. 0.2 Blue is dominant because it is shown with an uppercase letter. White is recessive because it has a lowercase letter. The population has 200 white butterflies (bb). This makes the frequency of recessive homozygotes 200/500. If q2 = 0.4, then q = 0.2.
65. 12 The 5′ untranslated region of an mRNA is the region of mRNA where the ribosome docks before the AUG start codon. On a real mRNA there may be several AUGs, and the official starting AUG is preceded by a special sequence. In this question, there is only one AUG, so that must be the start codon. Count the bases before the AUG: UAA CGA UCA UAU = 12.
66. 50% Let’s use the same letters as the butterfly example above, but you could use any letters to represent your dominant and your recessive alleles. The following Punnett square shows the possible offspring from the initial cross, and the second square shows the next cross between the male offspring being backcrossed to the original female. There are 2/4 offspring from the second cross that are heterozygotes, or 50%.
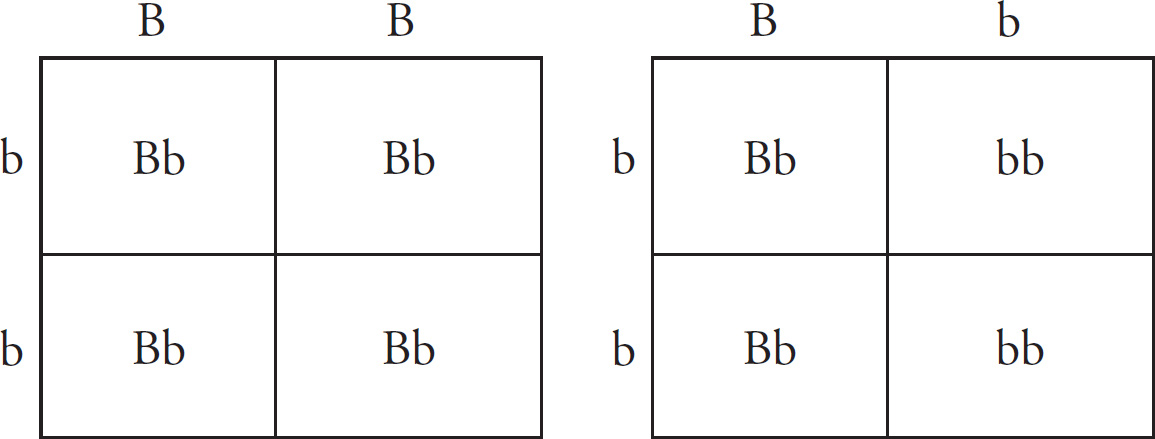
67. −30 The change in free energy is the following formula: free energy of the products – free energy of reactants.
20 − 50 = −30
68. 50 The activation energy is the energy required to reach the transition state from the reactants.
100 − 50 = 50
69. 5.8 Chi-square is calculated by the sum of (observed – expected)2/(expected) for each group in the experiment. In this experiment there are two groups: bugs on dry-side and bugs on wet-side. The expected are based on a null hypothesis. In this instance, the null hypothesis is that the bugs have no preference. The expected will be 5 bugs on each side.
Wet group: (observed – expected)2/(expected): (8.8 − 5)2/(5) = (3.8)2/(5) = 2.888
Dry group: (observed – expected)2/(expected): (1.2 − 5)2/(5) = (-3.8)2/(5) = 2.888
Add the two groups: 2.9 + 2.9 = 5.8
Section II: Free Response
On the following pages you’ll find two types of aids for grading your essays: checklists and sample responses. Checklists like these are used to grade your essays on the actual test. They’re actually quite simple to use.
For each item you mentioned, give yourself the appropriate number of points (1 point, 2 points, and so on). Remember that you can only get a maximum of 10 points for each essay. As you evaluate your work, don’t be kind to yourself just because you like your own essay. If the checklist mentions an explanation of structure and function and you failed to give both, then do not give yourself a point. The testing board is very particular about this. You need to mention precisely the things they’ve listed, in the way they’ve listed them, in order to gain points.
What if something you’ve mentioned doesn’t appear on the list? Provided you know it’s valid, give yourself a point. If it was something you pulled out of your hat at the last minute, odds are it’s not directly applicable to the question. However, because you might come up with details even more specific than those contained in the checklist, it’s not unlikely that the example or structure you’ve cited is perfectly valid. If so, go ahead and give yourself the point. Remember that the testing board hires college professors and high school teachers to read your essays, so they’ll undoubtedly recognize any legitimate information you slip into the essay.
The second part of the answer key involves short paragraphs. These are not templates; the testing board does not expect you to write this way. They are simply additional tools to help illustrate the things you need to squeeze into your essay in order to rack up the points. They explain in some detail how the various parts of the checklist relate to one another and may give you an idea about how best to integrate them into your own essays come test time.
If you find that you didn’t do too well on the essay portion, be sure to read carefully the chapter on the free-response questions (Chapter 14). You can use the chapters themselves as checklists. If you find it too difficult to grade your own essays, see if your teacher or a classmate will help you out. Good luck!
Long-Form Free-Response Checklist 1
(10 points total)
I. Experiment describing antibiotic resistance
a. Describe your methods
i. Clearly describe methods including controls (1 point)
ii. Designed experiment would correctly answer the question (1 point)
b. Show mock data in a graph or table
i. Include proper labels (1 point)
ii. Include data with an appropriate trend (1 point)
c. Justify your conclusion by referencing the graph (1 point)
II. Experiment testing if resistance can be transferred
a. Describe your methods
i. Clearly describe methods including controls (1 point)
ii. Designed experiment would correctly answer the question (1 point)
b. Show mock data in a graph or table
i. Include proper labels (1 point)
ii. Include data with an appropriate trend (1 point)
c. Justify your conclusion by referencing the graph (1 point)
Example of a 10-Point Response
To test antibiotic resistance in each of the three types of bacteria, each type of bacteria should be grown in the presence of each antibiotic (ampicillin and tetracycline). They should also be grown in plates without antibiotic as a control. This means there should be three conditions of growth for each type of antibiotic. It would also be best to do this with replicates to rule out random problems that may occur.
This shows that bacteria A is resistant to ampicillin because it still grows when ampicillin is there. Bacteria B is resistant to tetracycline because it still grows when tetracycline is there. Bacteria C grows in all conditions, so it is resistant to both ampicillin and tetracycline.
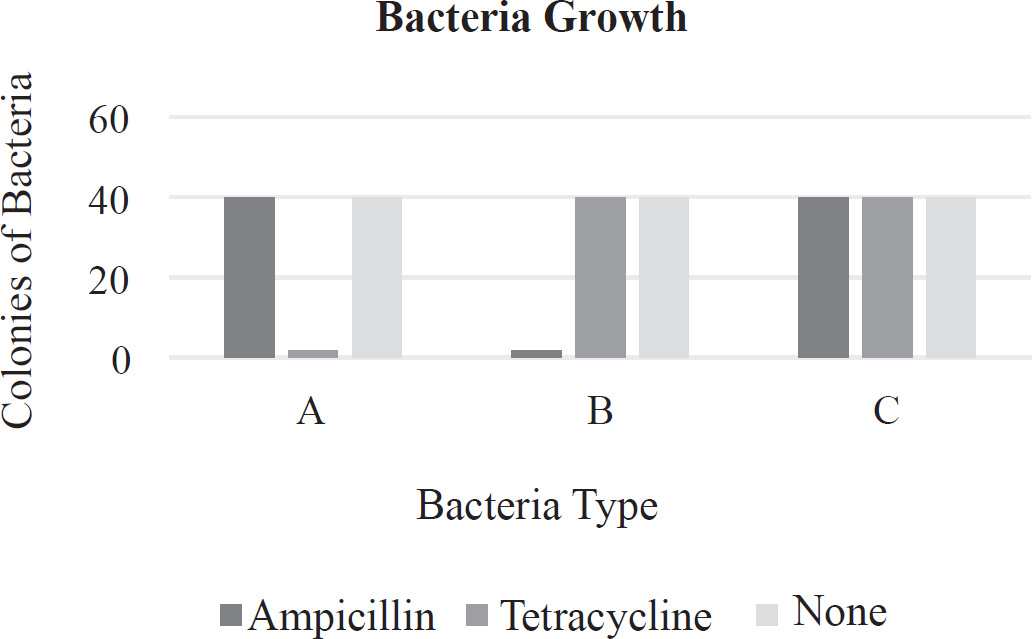
To determine if bacterial resistance can be transferred, bacteria that is sensitive to an antibiotic would have to be transformed with an antibiotic resistance gene and then grown in the presence of the antibiotic. If it grows, then the resistance can be transferred. Bacteria A can’t grow with tetracycline, so it is transformed with the gene for tetracycline resistance. Similarly, bacteria B is transformed with the gene for ampicillin resistance. Each one is also transformed with an empty plasmid as a control to show that the transformation process did not harm them.
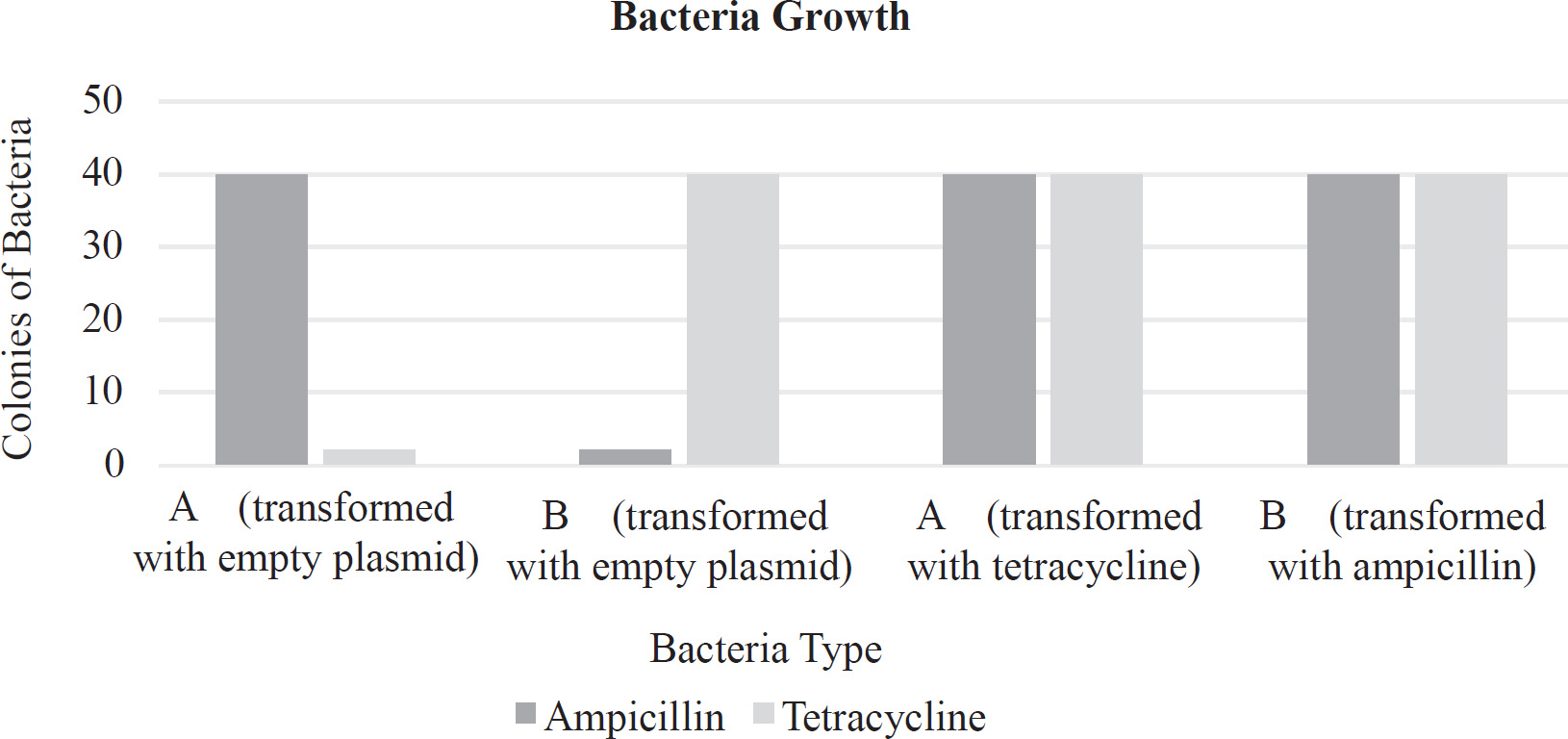
This shows that antibiotic resistance can be transferred. Bacteria A used to be sensitive to tetracycline, but after a transformation with tetracycline, it can grow in the presence of tetracycline. Similarly, bacteria B used to be sensitive to ampicillin and now the bacteria is resistant to it and can grow with ampicillin present.
Long-Form Free-Response Checklist 2
10 points total
I. Explain the importance of homeostasis (2 points)
a. Give 3 examples (3 points)
II. Contrast homeostasis in a simple unicellular organism and a complex organism (2 points)
III. Describe the effects of PTH and calcitonin on:
a. intestinal absorption rates (1 point)
b. kidney reabsorption rates (1 point)
c. production of osteoblasts/osteoclasts (1 point)
Example of a 10-Point Response
Homeostasis is important in the body because our body systems are built to function within certain parameters. If the pH or the temperature become too low or too high, then our enzymes will stop working, our membranes won’t function well, and molecules will not form the right shapes anymore. If osmolarity is not maintained, then cells can explode or shrink. Some physiological variables that must be maintained are blood pressure, temperature, pH, blood osmolarity, glucose levels, oxygen levels, sodium levels, etc.
Homeostasis is important in both types of organisms. Blood pressure and blood osmolarity levels are not present in unicellular organisms, but they still need to have the right pressure and osmolarity in their cytoplasm. Other things, such as pH and temperature, are still important in unicellular organisms. Homeostasis is more difficult to maintain in a complex organism because there are more spaces and compartments to maintain. There are also systems that are at play far away from each other, and the body needs to coordinate them to maintain homeostasis.
PTH is released when calcium is low. If calcium is low, intestinal absorption of calcium will increase. Kidney reabsorption will also increase. The production of osteoblasts will decrease, and the number of osteoclasts will increase because the osteoclasts are necessary to break down bone to get more calcium.
Calcitonin will be released when calcium is high. This means that the intestinal absorption will decrease because the body does not need more calcium. The kidney reabsorption rate will decrease as well. More osteoblasts will be made, and less osteoclasts will be made because bone can be built and not broken down.
Short-Form Free-Response Checklist 3
3 points total
I. Describe the reactants and products (1 point)
II. Describe the light-dependent reactions (1 point)
III. Describe the light-independent reactions (1 point)
Example of a 3-Point Response
The reactants of photosynthesis are water, sunlight, and carbon dioxide, and the products are oxygen and sugar. The process is divided into two parts: light-dependent and light-independent reactions. In the light-dependent reactions, energy from the sun is absorbed by chlorophyll and energizes electrons. These are then passed to the electron transport chain, where they are passed along and hydrogens are pumped. The hydrogens come from splitting a water molecule, releasing oxygen. At the end of the chain, NADPH is formed, and the hydrogens can form ATP by chemiosmosis with an ATP synthase. In the light-independent reactions, the NADPH and ATP from the light-dependent reactions and CO2 are used in the Calvin-Benson-Bassham cycle to make sugar. The plant can either use this directly or it can break it down for cellular respiraton.
Short-Form Free-Response Checklist 4
4 points total
I. Describe innate response to a first-time exposure. (2 points)
II. Describe the specific immune response to a first-time exposure (2 points)
Example of a 4-Point Response
The innate immune response does not try to identify what the pathogen is. It just fights anything that seems foreign. The innate response begins with the barriers of the body such as the skin, the mucus membranes, and the acid in the stomach. These things would prevent something from getting inside our bodies. However, should something infect us, then It would cause inflammation, and certain cells like macrophages would try to eat the pathogen.
The specific immune response comes in later. B-cells will make antibodies that can bind to the pathogen. This marks the pathogen for destruction by other immune cells. Other cells, called T-cells, help identify cells that are infected with pathogens and destroy them. After the infection, a militia of memory cells remains to be ready in case the pathogen returns.
Short-Form Free-Response Checklist 5
3 points total
I. Explain why a universal molecule is needed (1 point)
II. Give examples of energetically unfavorable reactions (1 point)
III. Explain how they can be powered (1 point)
Example of a 3-Point Response
A universal energy molecule is needed because many reactions in the body need energy. It makes it easier to have one common source of energy for many reactions instead of multiple sources. This makes it easier to make energy and supply it. It also makes it easy to measure the amount of energy in a cell, because only one type of molecule needs to be counted.
Reactions that are energetically unfavorable are reactions that build things or make things. DNA replication requires energy. Building proteins requires energy. Active transport requires energy. Cell movement requires energy.
Reactions that are unfavorable can be coupled to the hydrolysis of ATP. When a phosphate is removed from ATP, this is very favorable, and the excess energy from this reaction can be used to power an unfavorable reaction.
Short-Form Free-Response Checklist 6
4 points total
I. Describe how two shapes are possible (1 point)
II. Mention how the process of DNA replication could lead to two different shapes of protein (1 point)
III. Mention how the process of transcription could lead to two different shapes of protein (1 point)
IV. Mention how the process of translation could lead to two different shapes of protein (1 point)
Sample of a 4-Point Response
Two shape outcomes can be possible if a normal protein is simply misfolded or if the protein was made incorrectly, which causes it to fold in an abnormal way. The normal shape is one outcome, and the abnormal shape is another outcome. The shape of the protein depends on its amino sequence and the way it folds up with itself and with other proteins.
DNA replication is not always perfect. DNA polymerase can make errors, which are called mutations. These mutations can be missing nucleotides, or added nucleotides, or the wrong nucleotides.
These same types of mistake can be made by RNA polymerase during the process of transcription. Additionally, a mistake made during DNA replication would also be carried over through transcription. These mistakes technically would not cause the problem in the protein later.
During translation, mistakes in DNA replication or transcription can cause the wrong amino acid to be inserted, or no amino acid to be inserted, a stop codon to be inserted, or the translation reading frame to be shifted entirely. Translation itself can also cause problems if the ribosome happens to make a mistake.
Short-Form Free-Response Checklist 7
3 points total
I. Describe natural selection (2 points)
II. Explain how genetic drift differs from natural selection (1 point)
Example of a 3-Point Response
Natural selection is the process whereby traits that help individuals reproduce better than others (called evolutionary fitness) will become more prevalent over time because they are passed on to future generations at a greater rate. An example of this is if faster individuals became more prevalent over time because they can outrun predators and live to reproduce more than slower individuals. Slower individuals would not reproduce, and the next generation would have more faster individuals than slow individuals, thereby causing the next generation to be faster, and so on.
Genetic drift is different because it is all just random luck. If a tornado comes through and kills 90% of a population, the individuals that survive are not necessarily the fastest since nobody can outrun a tornado, but their genes are going to be passed on to the next generation even if they are slowpokes. Genetic drift is when certain traits become more prevalent because of random luck and not because they natural selected because they make an individual reproduce better.
Short-Form Free-Response Checklist 8
4 points total
Draw a food energy pyramid (2 points)
Explain why only 10% is passed on at each level (2 points)
Example of a 4-Point Response
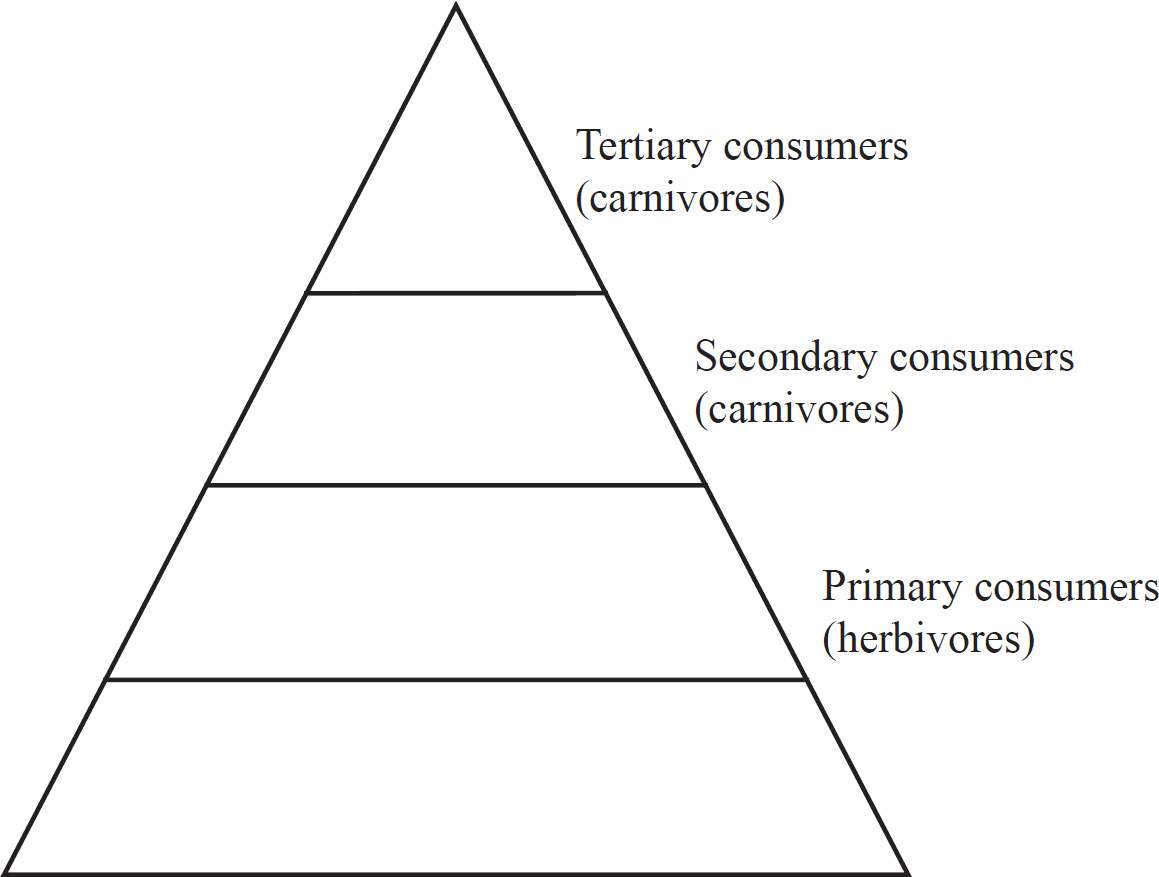
An energy pyramid shows that of all the energy stored in the living things in an ecosystem, most of it is in the producers. There are lots of producers, and they make their own energy from photosynthesis. Primary consumers eat these plants, but some of the plant energy that they eat gets used in the daily life of the primary consumer (walking around, communicating, digestion, etc.). It is estimated that 90% of the energy consumed by eating the level below is lost to the environment through normal activities. When a secondary consumer eats a primary consumer, it does not get the full amount of producer-energy that the primary consumer originally ate; it only gets 10% of it. The same thing happens at each level, which is why the lower level will always have more energy than the level above it. Only if a primary consumer were to eat a producer and immediately be eaten by a secondary consumer would the secondary consumer be consuming a greater amount of energy that was eaten from the producer level.
Practice Test 4
Click here to download a PDF of Practice Test 4
The Exam
AP® Biology Exam
SECTION I: Multiple-Choice Questions
DO NOT OPEN THIS BOOKLET UNTIL YOU ARE TOLD TO DO SO.
At a Glance
Total Time
1 hour and 30 minutes
Number of Questions
69
Percent of Total Score
50%
Writing Instrument
Pencil required
Instructions
Section I of this examination contains 69 multiple-choice questions. These are broken down into Part A (63 multiple-choice questions) and Part B (6 grid-in questions).
Indicate all of your answers to the multiple-choice questions on the answer sheet. No credit will be given for anything written in this exam booklet, but you may use the booklet for notes or scratch work. After you have decided which of the suggested answers is best, completely fill in the corresponding oval on the answer sheet. Give only one answer to each question. If you change an answer, be sure that the previous mark is erased completely. Here is a sample question and answer.
Sample Question
Chicago is a
(A) state
(B) city
(C) country
(D) continent
Sample Answer

Use your time effectively, working as quickly as you can without losing accuracy. Do not spend too much time on any one question. Go on to other questions and come back to the ones you have not answered if you have time. It is not expected that everyone will know the answers to all the multiple-choice questions.
About Guessing
Many candidates wonder whether or not to guess the answers to questions about which they are not certain. Multiple choice scores are based on the number of questions answered correctly. Points are not deducted for incorrect answers, and no points are awarded for unanswered questions. Because points are not deducted for incorrect answers, you are encouraged to answer all multiple-choice questions. On any questions you do not know the answer to, you should eliminate as many choices as you can, and then select the best answer among the remaining choices.
BIOLOGY
SECTION I
69 Questions
Time—90 minutes
Directions: Each of the questions or incomplete statements below is followed by four suggested answers or completions. Select the one that is best in each case and then fill in the corresponding oval on the answer sheet.
Questions 1–3 refer to the following passage and figure.
During photosynthesis, chlorophyll pigment absorbs sunlight that energizes its electrons. There are two types of chlorophyll pigment found in plants: chlorophyll a and chlorophyll b. Each pigment absorbs light at different optimal wavelengths, and unabsorbed light is reflected outward. To energize its electrons, it is important that light of absorbable wavelengths reaches the plant for photosynthesis to occur. The graph below demonstrates the absorbed spectrum of light for chlorophyll a, chlorophyll b, and another type of pigment found in plants called carotenoid.
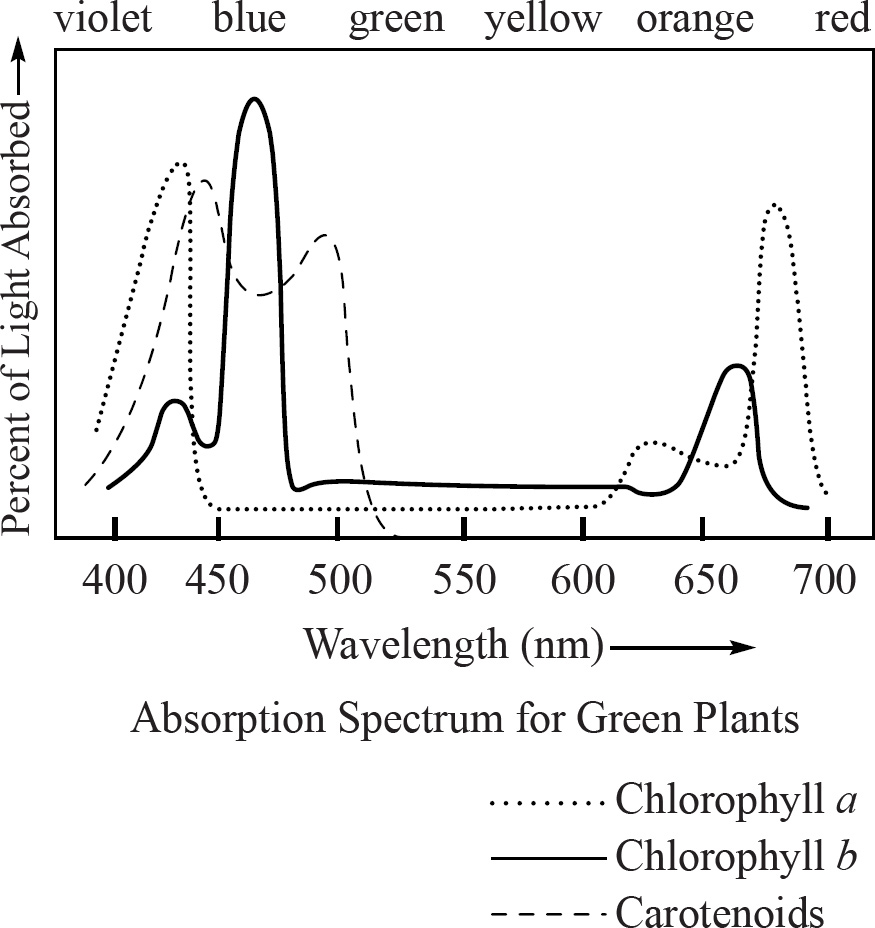
1. Approximately which color wavelengths would a plant containing chlorophyll a appear as to humans?
(A) 400–450 nm
(B) 450–600 nm
(C) 675–700 nm
(D) None of the above, because humans cannot see the visible light spectrum.
2. If carotenoids were capable of energizing electrons for photosynthesis in the same manner as chlorophyll, in approximately which wavelengths of light would they best perform photosynthesis?
(A) less than 450 nm
(B) 450–500 nm
(C) 500–700 nm
(D) greater than 700 nm
3. The following graph shows the height of a new species of plant when it is grown under lights of different wavelengths. What color does this plant likely appear to humans?
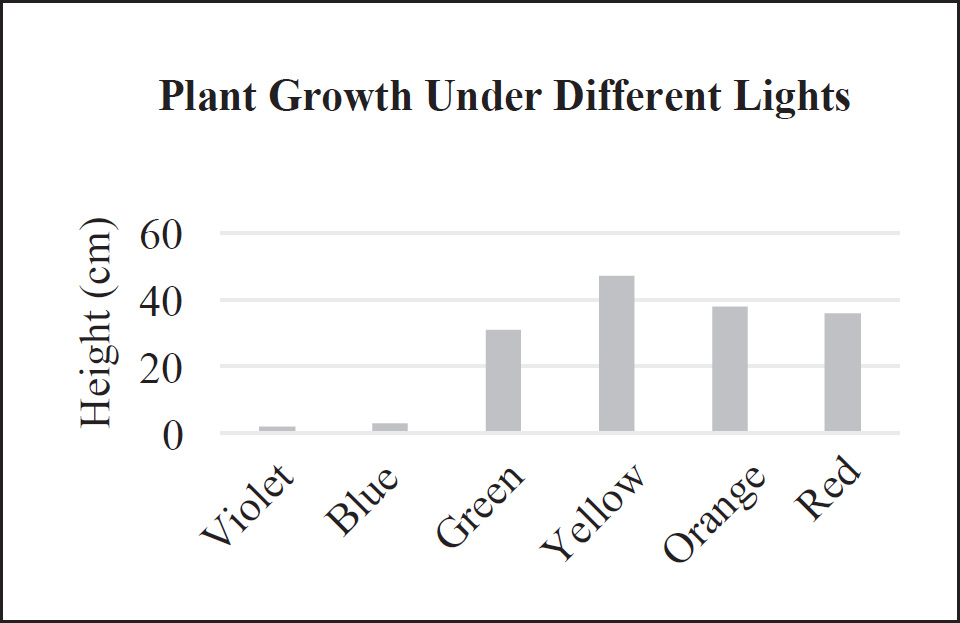
(A) Violet
(B) Green
(C) Yellow
(D) Red
Questions 4–5 refer to the following passage and figure.
The following table identifies the stages of embryogenesis in humans. As the embryo develops, the cells differentiate, and their potency, or potential to become many types of cells, is reduced.
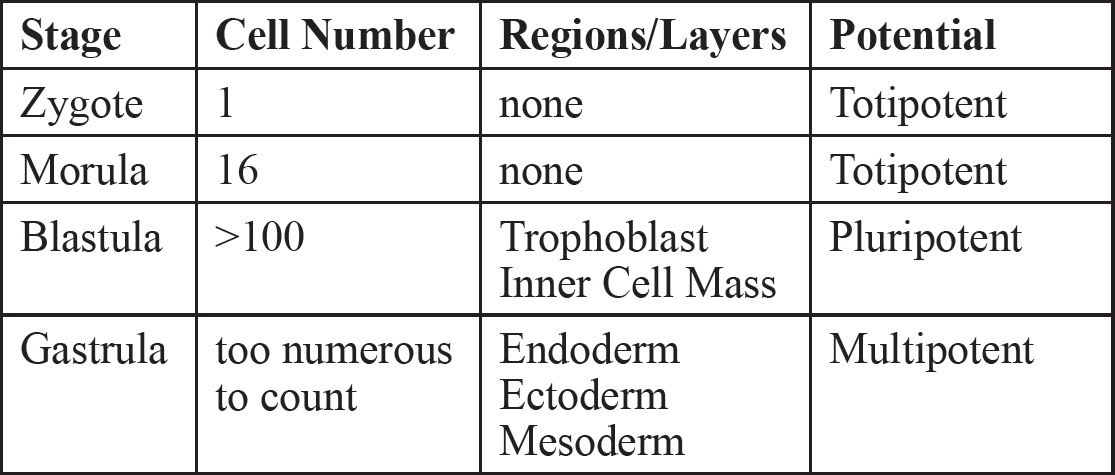
4. Which of the following could be derived from a totipotent stem cell?
I. Morula
II. Trophoblast
III. Endoderm
(A) I only
(B) II only
(C) II and III
(D) I and II and III
5. If a cell in a morula becomes a cell of the trophoblast, it has become
(A) more differentiated and less specialized
(B) less differentiated and more specialized
(C) more differentiated and more specialized
(D) less differentiated and less specialized
Questions 6–8 refer to the following passage and figure.
Four genes in the TNS family are identified as being involved in the viral entry of a particular virus. To evaluate this claim, the expression of each gene is reduced using RNA interference. Afterwards, viral infection is attempted and the viral load is assessed. A non-targeting (NT) siRNA is used as a control.
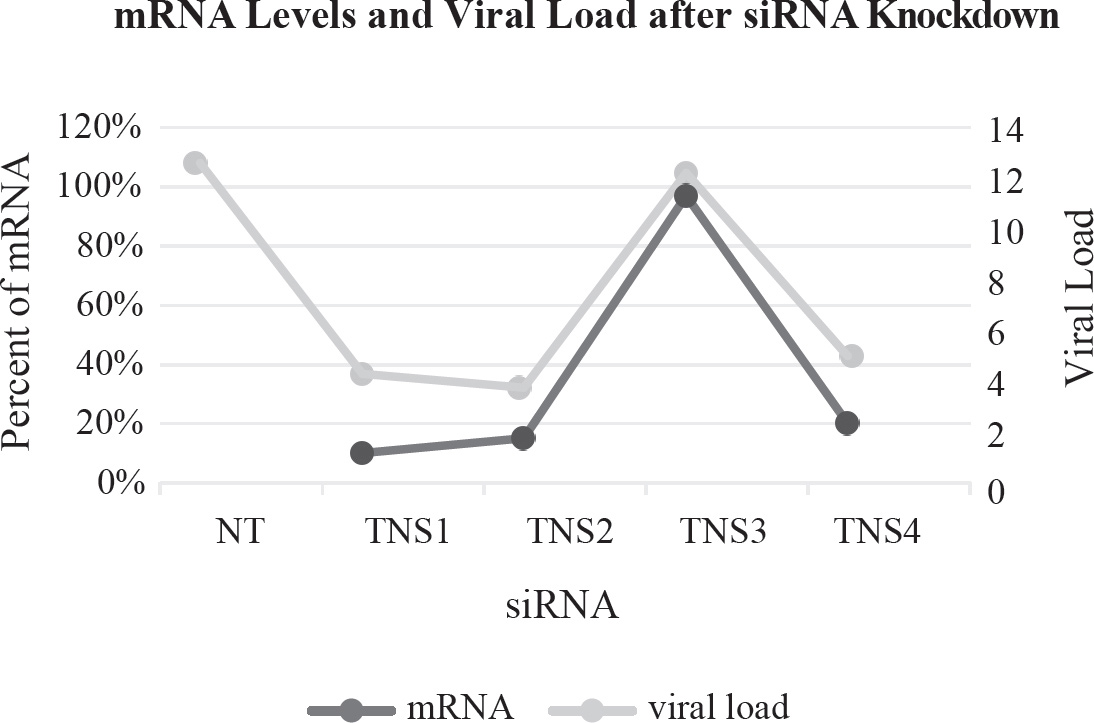
6. Which likely describes why there is no mRNA data for the NT siRNA treated sample?
(A) NT siRNA is designed to NOT reduce expression of any specific gene.
(B) NT siRNA is designed to reduce the expression of ALL genes equally.
(C) NT siRNA reduces the expression to levels below the detectable threshold.
(D) NT siRNA enhances viral infection, and looking at mRNA is not necessary.
7. Which of the following conclusions is correct?
I. TNS 1 is involved in viral infection.
II. TNS 2 is involved in viral infection.
III. TNS 3 is NOT involved in viral infection.
(A) I only
(B) III only
(C) I and II
(D) I and II and III
8. Which of the following would NOT have been helpful as a control in this experiment?
(A) An siRNA for a gene known to be involved in viral infection
(B) An siRNA for a gene known to make very little mRNA
(C) An siRNA for a gene known to NOT be involved with viral infection
(D) A second siRNA that targets each of the genes being studied
Questions 9–11 refer to the following passage.
Frederick Griffith showed in 1928 that when a heat-killed virulent strain of bacteria was mixed with a living non-virulent strain of bacteria, it resulted in an infection with living virulent bacteria. They called this process the transformation of the non-virulent bacteria.
In 1944, Avery, McCarty and MacLeod used laboratory techniques to separate the DNA, RNA, lipids, proteins, and carbohydrates present in a virulent bacterial strain and then they repeated Griffiths experiment to see which fraction was capable of transforming the non-virulent strain into a virulent strain. Since the DNA fraction was the only one that transformed the bacteria, they concluded that DNA must be the macromolecule that gives characteristics in bacteria.
9. Which of the following conclusions could be made in 1929?
(A) A heat-resistant molecule can be passed to living bacteria and change its characteristics.
(B) A DNA molecule cannot be killed by heat treatment when it is inside a bacterium.
(C) Heat-killed bacteria can come back to life when they are mixed with virulent bacteria.
(D) A virulent bacterial strain will always win out against a non-virulent strain of bacteria.
10. Which of the following, if true, would most negate Avery, McCarty, and MacLeod’s conclusion?
(A) If it was shown that the fraction of lipids did not contain lipids
(B) If it was shown that the RNA fraction also contained trace amounts of DNA
(C) If it was shown that the DNA fraction also contained bits of protein
(D) If it was shown that the DNA fraction was much larger than the RNA fraction
11. Which of the following best describes the process of transformation as it is used above?
(A) Acquisition of any DNA by bacteria causes it to acquire new traits.
(B) The traits in bacteria are determined by the DNA that is present.
(C) DNA is capable of giving any species the traits of another species.
(D) Bacteria are revived after being heat-killed if they get new DNA.
Questions 12–14 refer to the following passage and figures.
A DNA molecule is a double helix formed from two strands of DNA that base pair together with hydrogen bonds. Adenine preferentially binds with thymine, and cytosine preferentially pairs with guanine. The two strands form an anti-parallel conformation in the helix with the 3′hydroxyl end of one strand aligning with the 5′ phosphate end of the other strand and vice-versa.
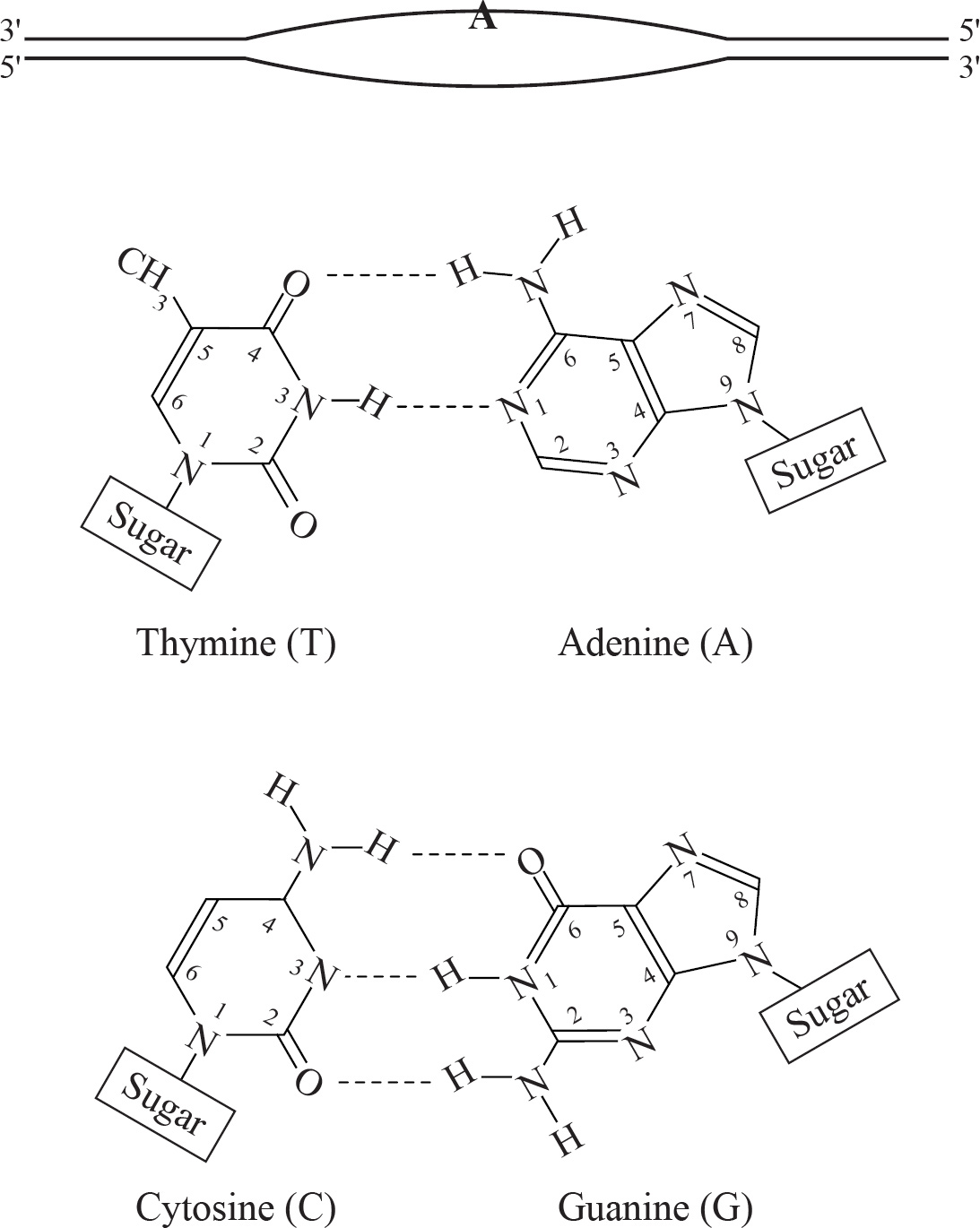
12. If a DNA dependent DNA polymerase were attached at site A, which statement best describes its movement?
(A) It will travel towards the right and build a copy of the top DNA strand.
(B) It will travel towards the left and build a copy of the top DNA strand.
(C) It will travel towards the right and build a complement of the top DNA strand.
(D) It will travel towards the left and build a complement of the top DNA strand.
13. The lagging strand cannot be built continuously during DNA replication for which of the following reasons?
(A) The DNA polymerase enzyme only travels in one direction, and when the helix opens in the opposite direction of the polymerase, there is limited space for the polymerase to travel, only building small segments at a time.
(B) The two strands of DNA are opening and closing asynchronously, and the polymerase must copy the limited small segments that are available to it before the helix closes.
(C) The lagging strand will not be used for the production of mRNA, and a continuous segment is only required for the coding strand that will code for the eventual mRNA.
(D) An RNA primer is needed during DNA replication, and the lagging strand is the last strand to receive an RNA primer during replication. The lagging strand must be built after a short delay.
14. As the base pairs align to form a double helix, which are easier to separate?
(A) A-T because it has 3 hydrogen bonds
(B) A-T because it has 2 hydrogen bonds
(C) G-C because it has 3 hydrogen bonds
(D) G-C because it has 2 hydrogen bonds
Questions 15–16 refer to the following figure.
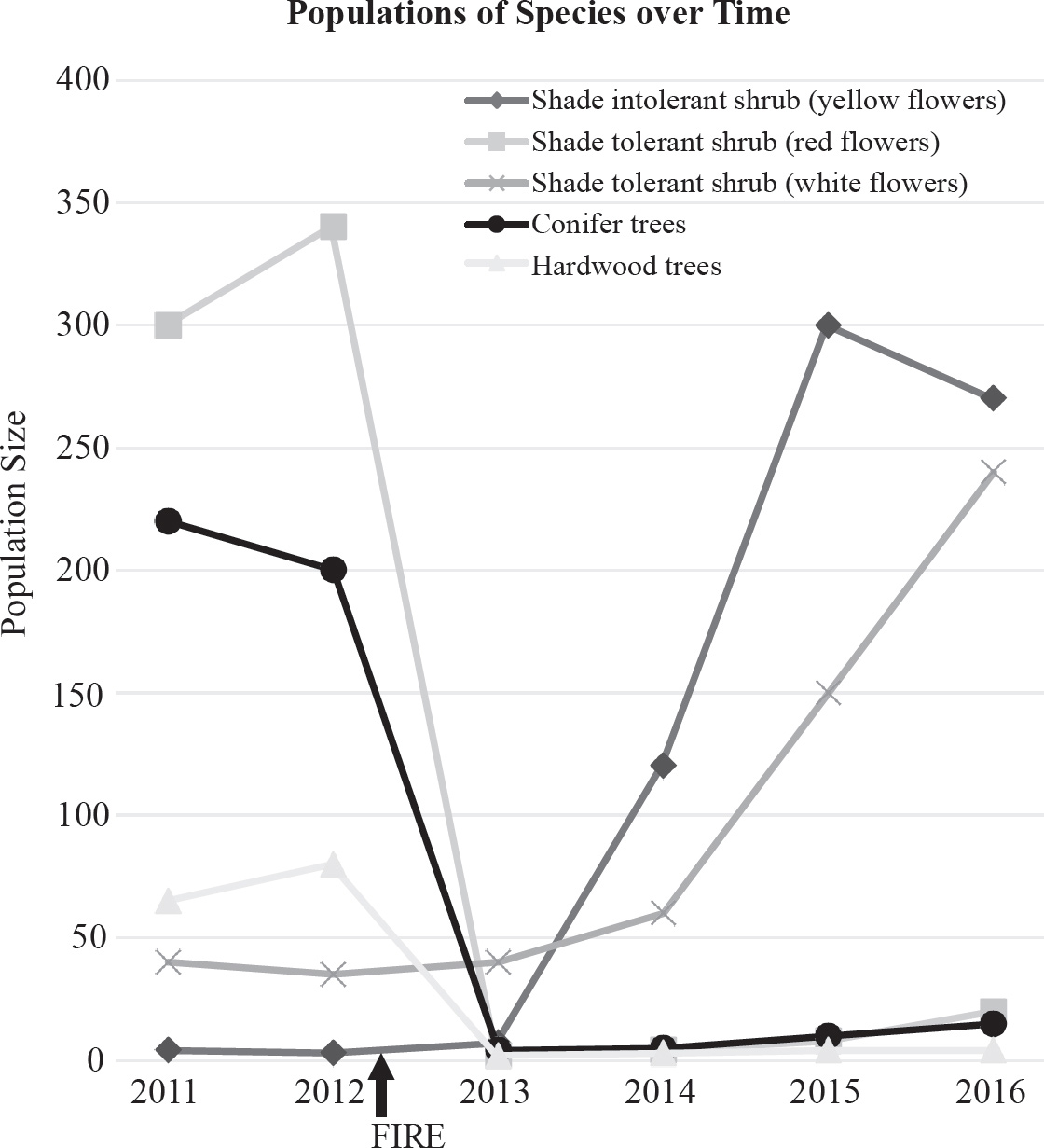
15. If the fire never occurred, which prediction about the population in 2016 is the most supported by the data?
(A) The white-flowered, shade tolerant shrubs would disappear from the region.
(B) The number of shade intolerant shrubs would increase, and the conifers would disappear.
(C) Conifers would claim the territory of all colors of the shade tolerant shrubs.
(D) Hardwood trees would increase in number, and conifers would decrease.
16. Which of the following could describe the growth patterns of the red- versus white-colored, shade tolerant shrubs?
I. Red flowers were selected for before the fire.
II. White flowers had increased fitness under the selective pressure of the fire.
III. White flowers are tolerant to fire and were always the most fit.
(A) I only
(B) II only
(C) I and II
(D) I and II and III
Questions 17–18 refer to the following figure.
| Fish | Urine Volume | Blood Osmolarity |
| Saltwater Fish | Low | Medium |
| Freshwater Fish | High | Medium |
17. What describes the urine output for the saltwater and the freshwater fish?
(A) The freshwater has a large volume of urine because freshwater is less dilute than the inside of the fish and the urine is necessary to change the osmolarity of the water surrounding the fish.
(B) The saltwater fish doesn’t need a large volume of urine because it is fine with a large amount of salt. It does not need to discard it as waste.
(C) The freshwater fish drinks a lot of water. It must urinate a lot so it does not rupture from too much water because it cannot control how much it drinks.
(D) The saltwater fish has less solute than the environment and constantly loses water to the environment. It needs low volume of urine to retain water.
18. Salmon spend some time in saltwater and some time in freshwater. Which of the following would you expect for the salmon at different phases of life?
(A) Saltwater salmon should have a lower blood osmolarity and a higher urine volume than freshwater salmon.
(B) Freshwater salmon should have a higher blood osmolarity and a higher urine volume than saltwater salmon.
(C) Saltwater salmon should have the same blood osmolarity and a higher urine volume than saltwater salmon.
(D) Freshwater salmon should have the same blood osmolarity and a higher urine volume than saltwater salmon.
Questions 19–20 refer to the following figure.
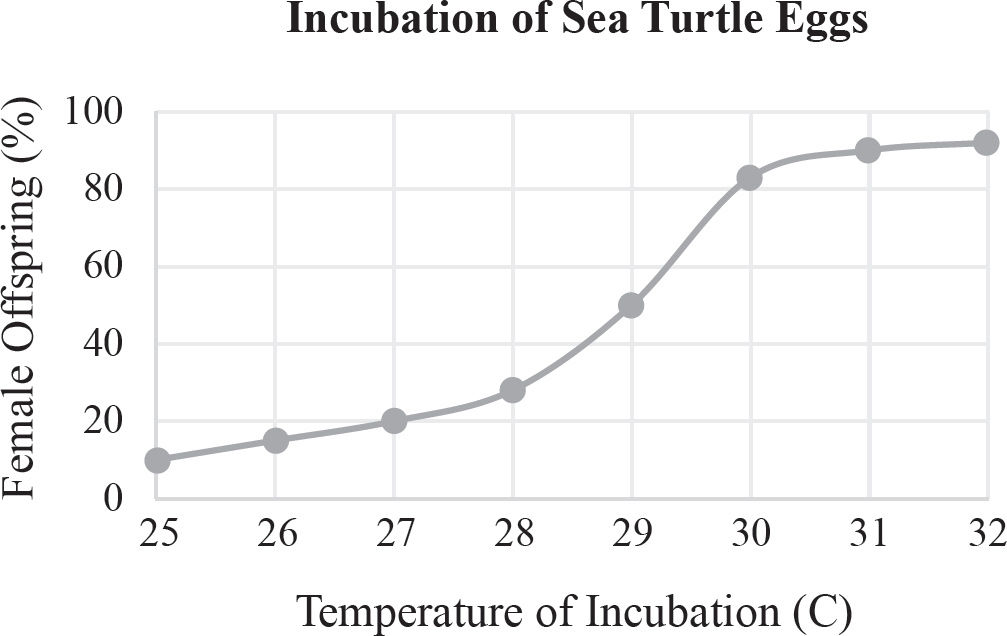
19. At which temperature would you expect 40 male turtles and 160 females turtles to be born?
(A) 26.5
(B) 38.2
(C) 29.7
(D) 31.4
20. Temperature is a key element in determining the activity of enzymes. If a particular enzyme must be active for female offspring to occur, which of the following enzyme activity versus temperature graphs represents that enzyme?
(A) 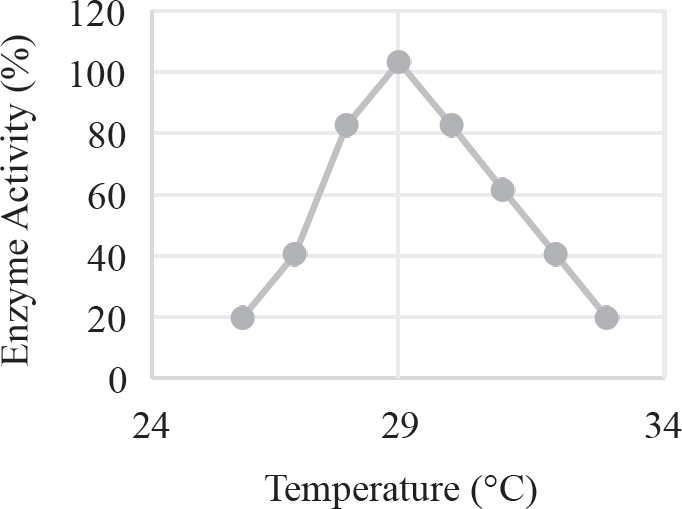
(B) 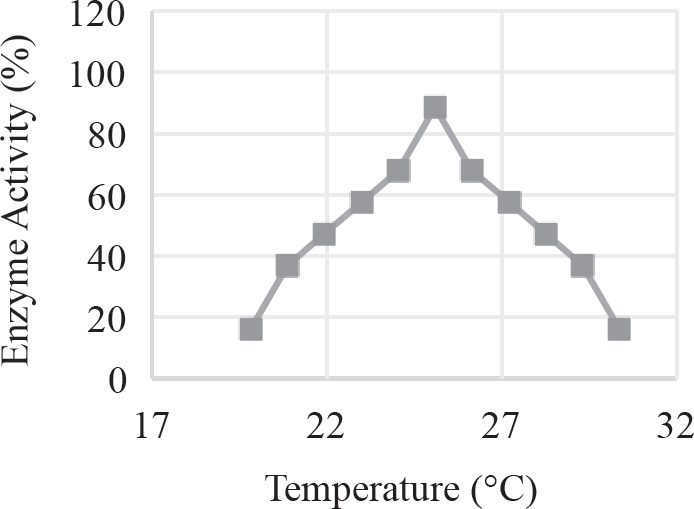
(C) 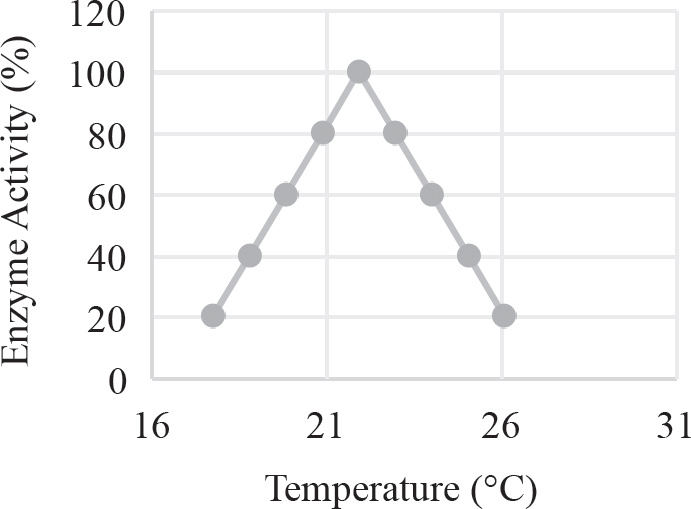
(D) 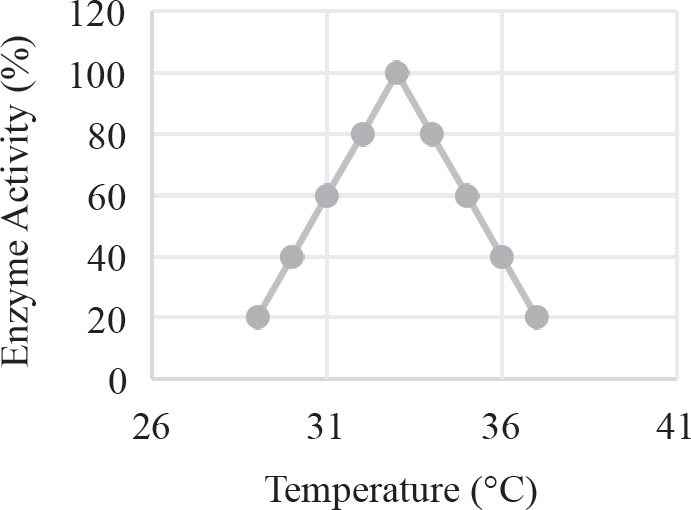
Questions 21–23 refer to the following passage and table.
Breast cancer is not a one-size-fits-all disease. Tumors in the breasts can occur due to many different factors. Depending on the characteristics of the tumor cells, the treatment and prognosis can vary greatly. The table below lists two (of many) known tumor markers. Each of these genes is often found to be overexpressed in breast cancer cells.
| Tumor Marker | Function | Drug |
| Estrogen Receptor (ER) | Docking site for the hormone estrogen, which causes changes in transcription that can lead to cell division | Tamoxifen |
| Human Epidermal Growth Factor Receptor 2 (HER2) | Promotes cell division and growth | Herceptin |
21. According to the passage, HER2 is found
(A) on tumor cells only
(B) on healthy cells, but it is found more often on tumor cells
(C) on healthy cells only
(D) on healthy cells, but is found less often on tumor cells
22. Which of the following is likely true for a cell with mutated ER leading to loss of function?
(A) It does not respond to estrogen as well.
(B) It binds to progesterone instead of estrogen.
(C) It divides more quickly than other cells.
(D) It is no longer under cell cycle control.
23. Which of the following is NOT a likely function of tamoxifen?
(A) Blocks the binding of estrogen and ER
(B) Increases the production of estrogen
(C) Decreases the production of ER
(D) Blocks the role of ER in the cell cycle
Questions 24–25 refer to the following passage and figure.
Humans make several types of antibodies called isotypes. There are five important isotypes: IgG, IgM, IgA, IgE, and IgD. The different isotypes are found in different parts of the body and on different types of immune cells. The following graph shows the prevalence of two isotypes at different points of an infection.
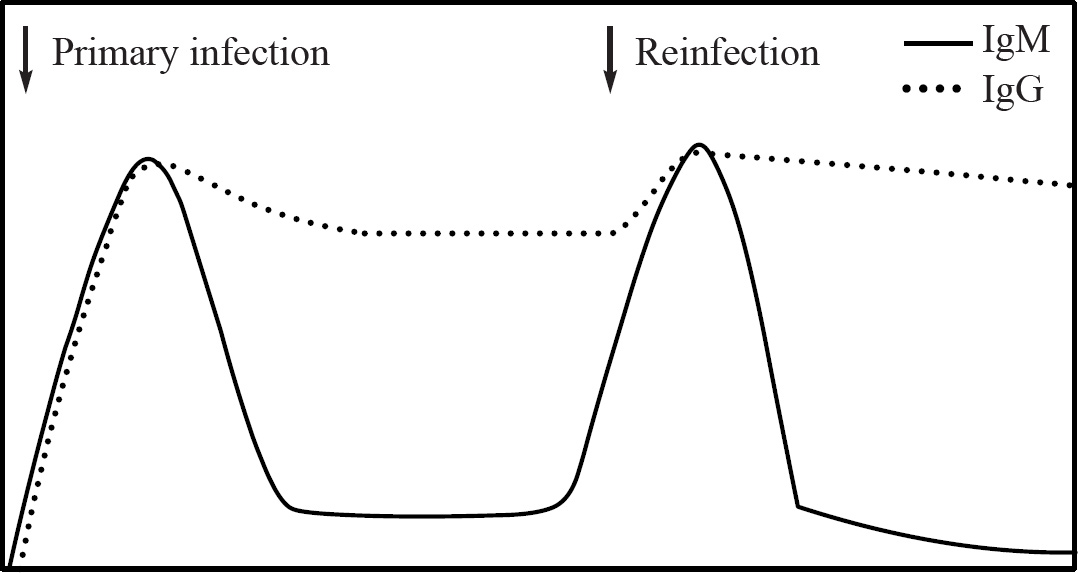
24. Which best describes the purpose of IgG demonstrated in the graph?
(A) It is stimulated by infection and is on memory B cells.
(B) It is stimulated by IgM levels during an infection.
(C) It becomes nonresponsive to antigen over time.
(D) It is formed from IgM that has switched isotypes.
25. An antibody monomer consists of a constant stem region and two arms that are each capable of binding to identical antigens (it is shaped like the letter Y). Different isotypes are polymers of identical monomers. For example, IgA is a dimer, and IgM is a pentamer. What is a likely benefit of IgM?
(A) It can cross-link several antigens together.
(B) It can cross membranes without assistance.
(C) It can be made without the addition of ATP.
(D) It can bind more than one type of antigen.
Questions 26–27 refer to the following figure.
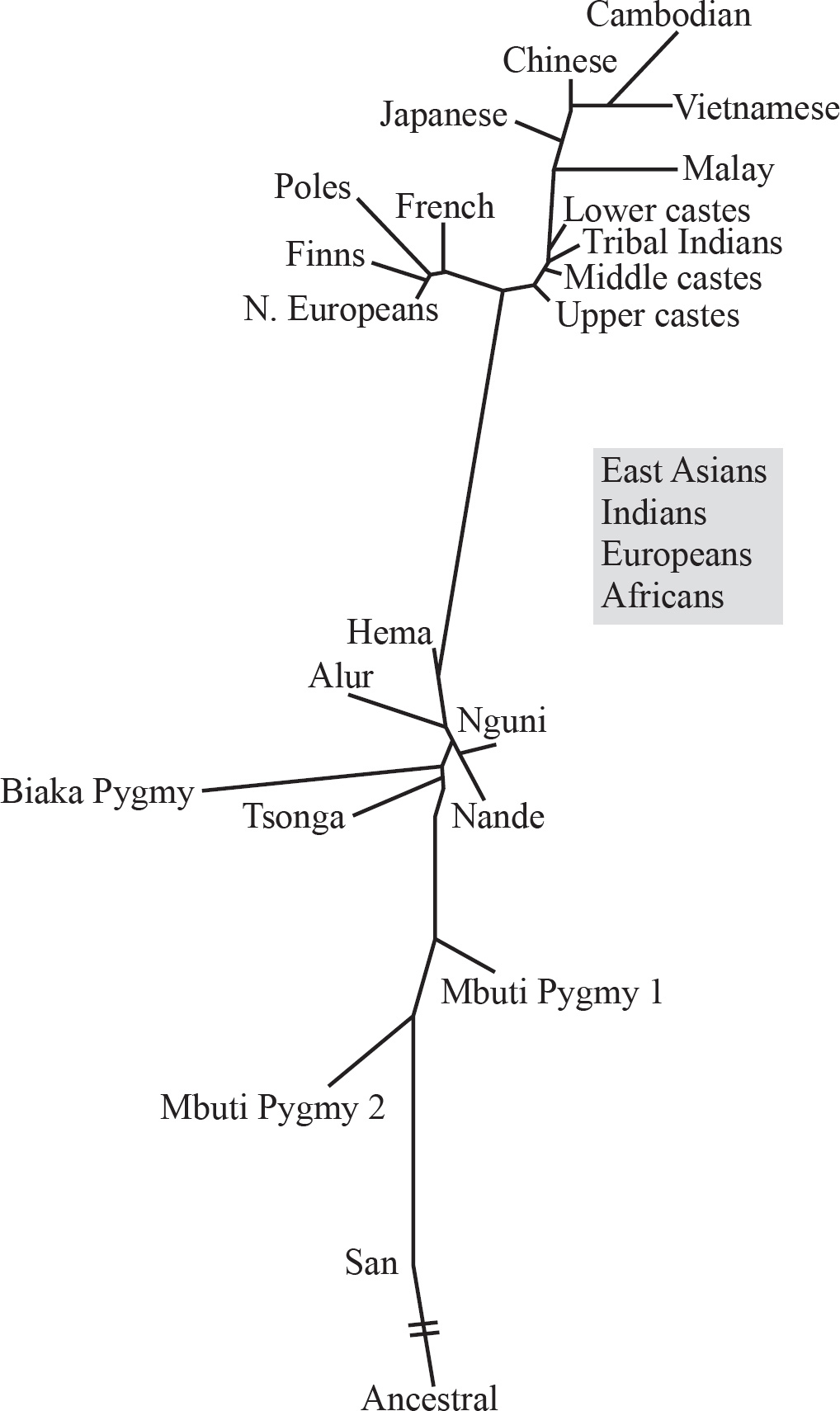
26. In the phylogenetic tree shown above, which of the following statements is true?
(A) Upper and lower castes do not share a common ancestor.
(B) The French are the common ancestor of Poles and Indians.
(C) Cambodians and Vietnamese have a common ancestor that Chinese do not share.
(D) Mbuti pygmy 1 and Mbuti pygmy are the closest related groups on the tree.
27. In the phylogenetic tree shown above, how many common ancestors do Tsonga and Japanese have?
(A) 0
(B) 4
(C) 6
(D) 12
Questions 28–31 refer to the following passage.
DNA and RNA polymerases build DNA and RNA, respectively. DNA polymerases typically have proofreading capabilities, whereas RNA polymerases typically do not. DNA-dependent DNA polymerases are used during DNA replication. DNA-dependent RNA polymerases are used during transcription.
28. Which of the following would require a RNA-dependent DNA polymerase?
(A) A bacterial strain that has a plasmid in addition to its genomic chromosome
(B) A virus that has a RNA genome and integrates into the host genome
(C) A plant cell that is being treated with RNA interference technology
(D) Any eukaryotic cell about to undergo post-transcriptional processing
29. What is the likely reason that DNA polymerases proofread and RNA polymerases don’t?
(A) Thymine is tough to base pair with, but uracil is easy to bond to.
(B) DNA requires base pairing to form a double helix and RNA is single stranded.
(C) DNA is passed from generation to generation, but RNA is only around for a short time.
(D) There is more than one type of RNA (mRNA, tRNA, rRNA), and that makes proofreading difficult.
30. Which of the following explains why RNA viruses are always changing?
(A) Their genome gets changed due to many mistakes during replication.
(B) RNA viruses are single-stranded and they can take many shapes.
(C) RNA does not need to travel in a capsid and gets exposed to chemicals.
(D) There are thousands of types of RNA viruses in the world.
31. What strand would likely be created from the following RNA sequence by an RNA-dependent-RNA polymerase enzyme?
5′ AUGUUUAGCGCUGGAUAC 3′
(A) 5′ GUAUCCAGCGCUAAACAU 3′
(B) 5′ AUGUUUAGCGCUGGAUAC 3′
(C) 5′ TACAAATCGCGACCTATG 3′
(D) 5′ CAUAGGUCGCAUUUGUA 3′
Questions 32–34 refer to the following passage.
Atmospheric nitrogen (N2) does not react well with other things; however, ammonia (NH3) is a form of nitrogen that can be readily used by many organisms. The process of turning N2 into NH3 (nitrogen fixation) is an essential process, but it is not one that many organisms are capable of. Bacteria called diazotrophs perform much of the natural nitrogen fixation on the planet. Legume plants have developed a symbiotic relationship with nitrogen-fixing rhizobia bacteria. The bacteria inhabit special nodules along the legume roots and provide the plant with fixed-nitrogen to use in cellular processes.
32. Which of the following molecules would be LEAST affected by a viral pandemic affecting rhizobia?
(A) Proteins
(B) RNA
(C) DNA
(D) Carbohydrates
33. Which of the following facts would classify the relationship as mutualistic?
(A) If the legumes were harmed during the process of nitrogen fixation
(B) If the rhizobia were harmed during the process of legume colonization
(C) If the legumes received another benefit apart from fixed-nitrogen
(D) If the rhizobia received a benefit from the legumes they colonize
34. Nitrogen fixation can also be accomplished synthetically through a chemical process known as the Haber process. Which of the following experiments would BEST show if synthetically fixed nitrogen or naturally fixed nitrogen was better for legume plant growth?
(A) Two groups of plants are created, each containing a different species. They are measured to be identical in every possible way and are planted in two conditions. One type of soil includes rhizobia bacteria, and the other includes synthetically fixed nitrogen. After a specific length of time, the height and width of each plant are measured.
(B) Two groups of plants are created, containing three plants, each of a different species. They are measured to be identical in every possible way and are planted in two conditions. One type of soil includes rhizobia bacteria, and the other includes synthetically fixed nitrogen. After a specific length of time, the height and width of each plant are measured.
(C) Two large groups of plants are created, each containing equal distributions of three different species. They are measured to be identical several ways and are planted in two conditions. One type of soil includes rhizobia bacteria, and the other includes synthetically fixed nitrogen. After a specific length of time, the height and width of each plant are measured.
(D) Two large groups of plants are created, each containing equal distributions of three different species. They are measured to be identical in every possible way and are planted in two conditions. One type of soil includes rhizobia bacteria, and the other includes synthetically fixed nitrogen. After a specific length of time, the height and width of each plant are measured.
Questions 35–38 refer to the following passage.
Transcription is often a carefully regulated process, with many factors determining if a gene product should be expressed. In the example shown below, the trp operon in some bacteria is a group of genes that work together to code for production of tryptophan. The promoter for the genes that code for tryptophan synthesis (trpE, trpD, trpC, trpB, and trpA) is located before an operator site through which the RNA polymerase must pass to initiate transcription. When tryptophan is present, a repressor (encoded by trpR) binds to the operator site and prevents transcription.
35. Which of the following could be true if there was a mutation in the trpR gene?
(A) The repressor protein would not bind as well to the operator site.
(B) RNA polymerase will no longer bind to the promoter.
(C) Tryptophan levels will not increase transcription.
(D) Tryptophan will bind directly to the operator site.
36. The direct relationship between tryptophan and the trp operon depends on which of the following?
(A) Equal parts of the repressor protein and tryptophan proteins
(B) Constant transcription and translation of the trpR gene
(C) Ongoing synthesis of tryptophan to unlock the repressor
(D) More than one promoter for RNA polymerase to bind at
37. Which diagram best shows the system when tryptophan is present?
(A) 
(B) 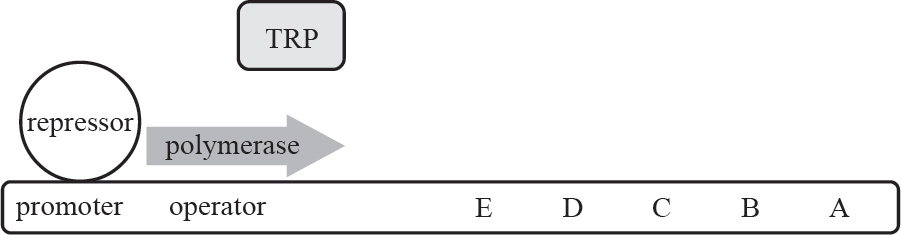
(C) 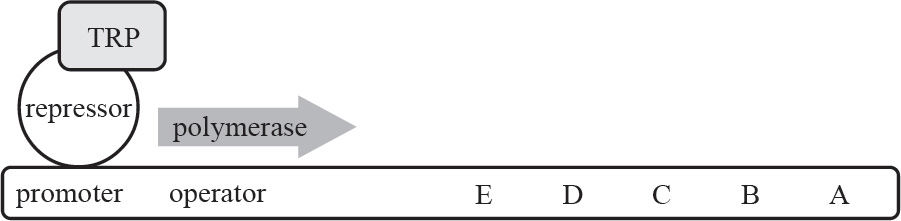
(D) 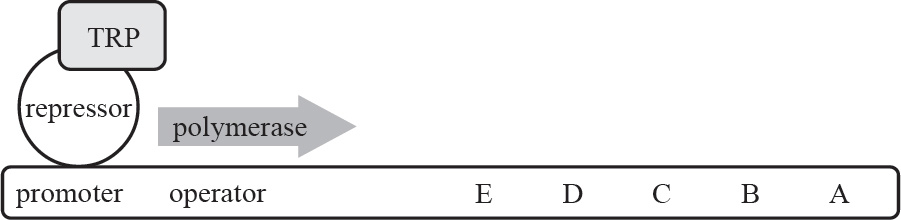
38. Which of the following graphs supports the information given on the trp operon?
(A) 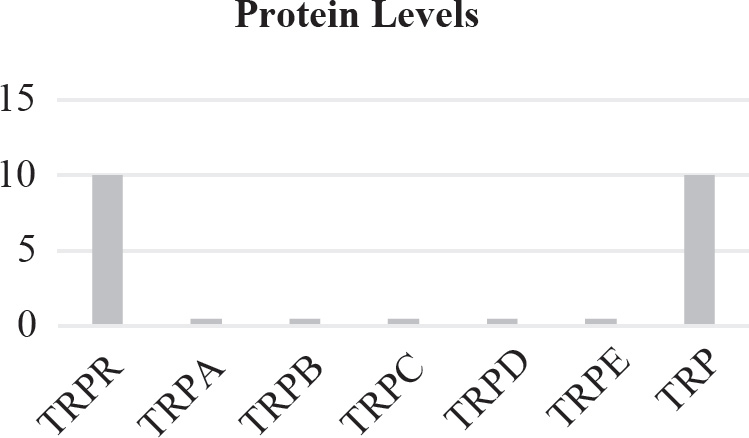
(B) 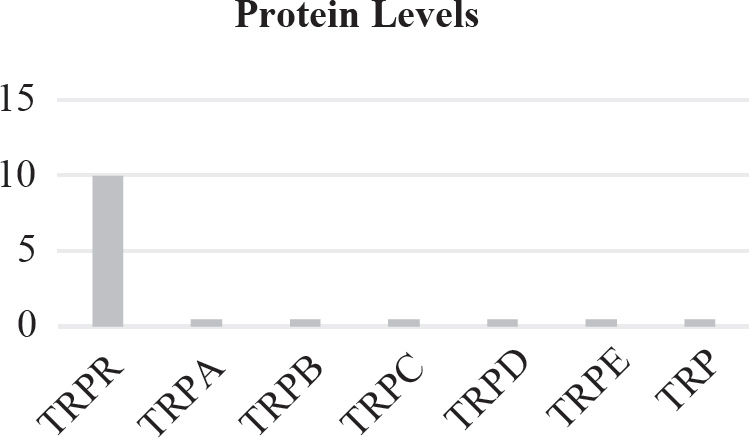
(C) 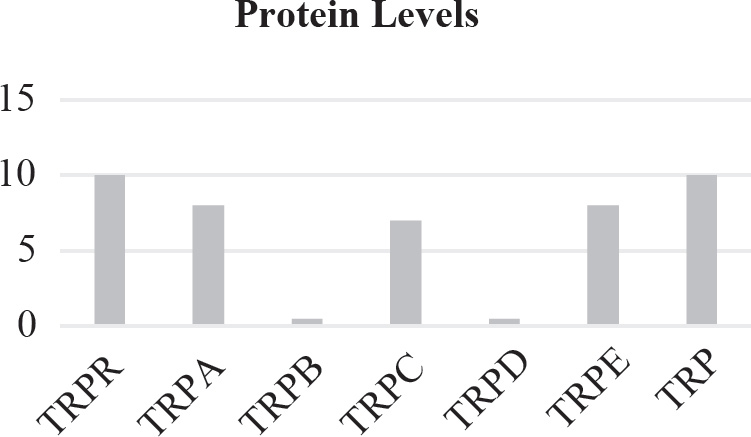
(D) 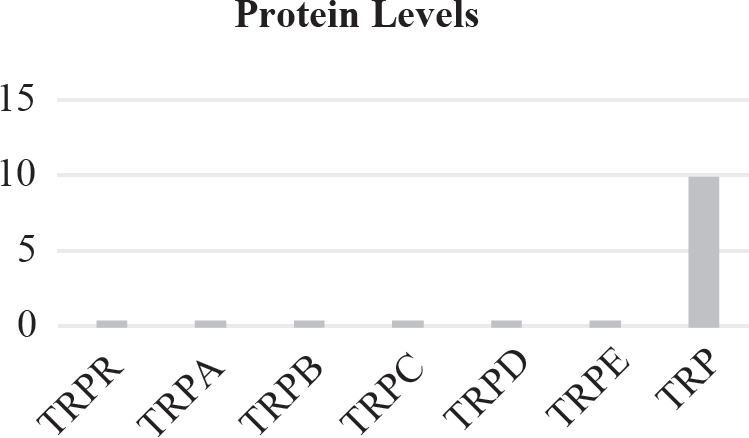
39. A virus is used to infect a plate of bacteria resistant to the antibiotic ampicillin and sensitive to the antibiotic tetracycline. New progeny viruses are collected and allowed to infect a culture of bacteria that is sensitive to both ampicillin and tetracycline. When ampicillin is added, some bacteria survive. What would be expected if tetracycline was added?
(A) They would survive because they acquired tetracycline resistance through viral transduction.
(B) They would survive because they failed to acquire tetracycline sensitivity through viral transduction.
(C) They would die because they acquired tetracycline sensitivity through viral transduction.
(D) They would die because they failed to acquire tetracycline resistance through viral transduction.
Questions 40–42 refer to the following passage.
The secretory pathway is a shipping route from the endoplasmic reticulum to the cell membrane. Things destined for secretion pass though the Golgi apparatus, which is an organelle that packages proteins it receives from the endoplasmic reticulum into transport vesicles. These proteins might be destined for incorporation in the membrane or they might be secreted from the cell via either constitutive or regulated secretion. Constitutively secreted proteins arrive at the membrane and are shipped without the requirement of further shipping signals. Regulated secretion requires the appropriate signal for the release of the protein from the cell. The Golgi also ships non-secreted proteins that will be incorporated into the intracellular lysosome, which is sometimes thought of as the stomach of the cell. It is the site of destruction of unwanted contents within the cell.
40. Which of the following is a benefit of regulated secretion?
(A) The proteins can be released more quickly after translation.
(B) The proteins can be built into larger complexes.
(C) The response to a stimulus can be limited and specific.
(D) The proteins can pass through the cell membrane.
41. Which of the following best depicts a vesicle with a protein destined for the cell membrane?
(A) 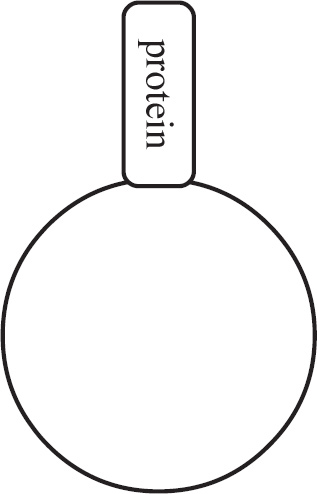
(B) 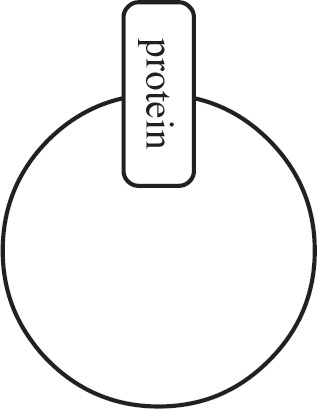
(C) 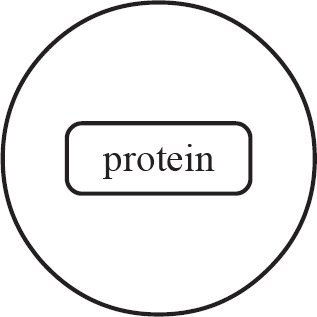
(D) 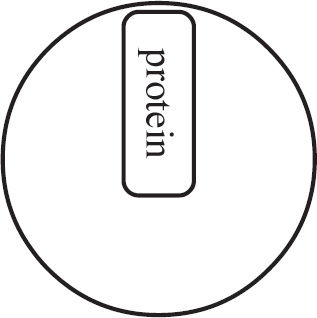
42. A lysosomal protein should have which of the following characteristics?
(A) Resists lysosomal enzymatic degradation
(B) Maintains stability for extended periods of time
(C) Contains both intracellular and extracellular domains
(D) Attaches to the intracellular side of the cell membrane
Questions 43–45 refer to the following passage.
The membrane potential of a cell is the difference in the charge on the intracellular side of the plasma membrane and the charge on the extracellular side. Most cells in the human body have a resting membrane potential of approximately −70mV. This is established primarily by the sodium-potassium ATPase that pumps three sodium out of and two potassium into the cell. The membrane potential of a cell is particularly important in neurons where action potentials are generated via a rush of Na+ ions down their concentration gradient.
43. What type of transport does the sodium potassium pump perform?
(A) Simple diffusion
(B) Facilitated diffusion
(C) Passive transport
(D) Active transport
44. When a neuron fires an action potential, how does the resting membrane potential change?
(A) It becomes more negative moving from the axon to the soma.
(B) It becomes more positive moving from the axon to the soma.
(C) It becomes more negative moving from the soma to the axon.
(D) It becomes more positive moving from the soma to the axon.
45. The following are all likely consequences of a failing sodium potassium pump EXCEPT?
(A) Action potentials would not be possible.
(B) The inside of the cell would become less negative.
(C) There would be more potassium outside the cell.
(D) There would be less sodium inside the cell.
Questions 46–47 refer to the following passage and figure.
The table below shows Watson-Crick base pairing (white) and wobble pairing (shaded) for RNA. There is even another nucleotide base that appears in tRNA anticodons. The wobble pairing can be seen between the nucleotide in the third position on the anticodon and the nucleotide in the 3′ most position on an mRNA codon. C: cytosine; A adenine; G: guanine; U: uracil; I: inosine.
| C | G |
| A | U |
| G | U |
| I | U |
| I | A |
| I | C |
46. Which of the following is the result of wobble pairing?
(A) There are more codons than there are anticodons.
(B) There are more amino acids than there are nucleotide bases.
(C) There are more inosines in mRNA than there are in tRNA.
(D) Some codons will only have one possible anticodon.
47. If the mRNA and the tRNA are oriented in an antiparallel direction during translation, what position on the tRNA is the wobble position?
(A) Always on the 5′ end of the anticodon
(B) Always on the 3′ end of the anticodon
(C) Sometimes on the 5′ end and sometimes on the 3′ end
(D) Neither, tRNA is not linear like mRNA
Questions 48–50 refer to the following passage and figure.
Table 1 shows different types of receptors found in the body, and the graph that follows shows the response to various stimuli of several unknown types of receptors.
| Stimulum | Type | Example |
| Light | Photoreceptors | Eye |
| Movement | Mechanoreceptors | Ear |
| Pressure | Baroreceptors | Blood vessels |
| Temperature | Thermoreceptor | Skin |
| Chemical | Promotes cell division and growth | Nose |
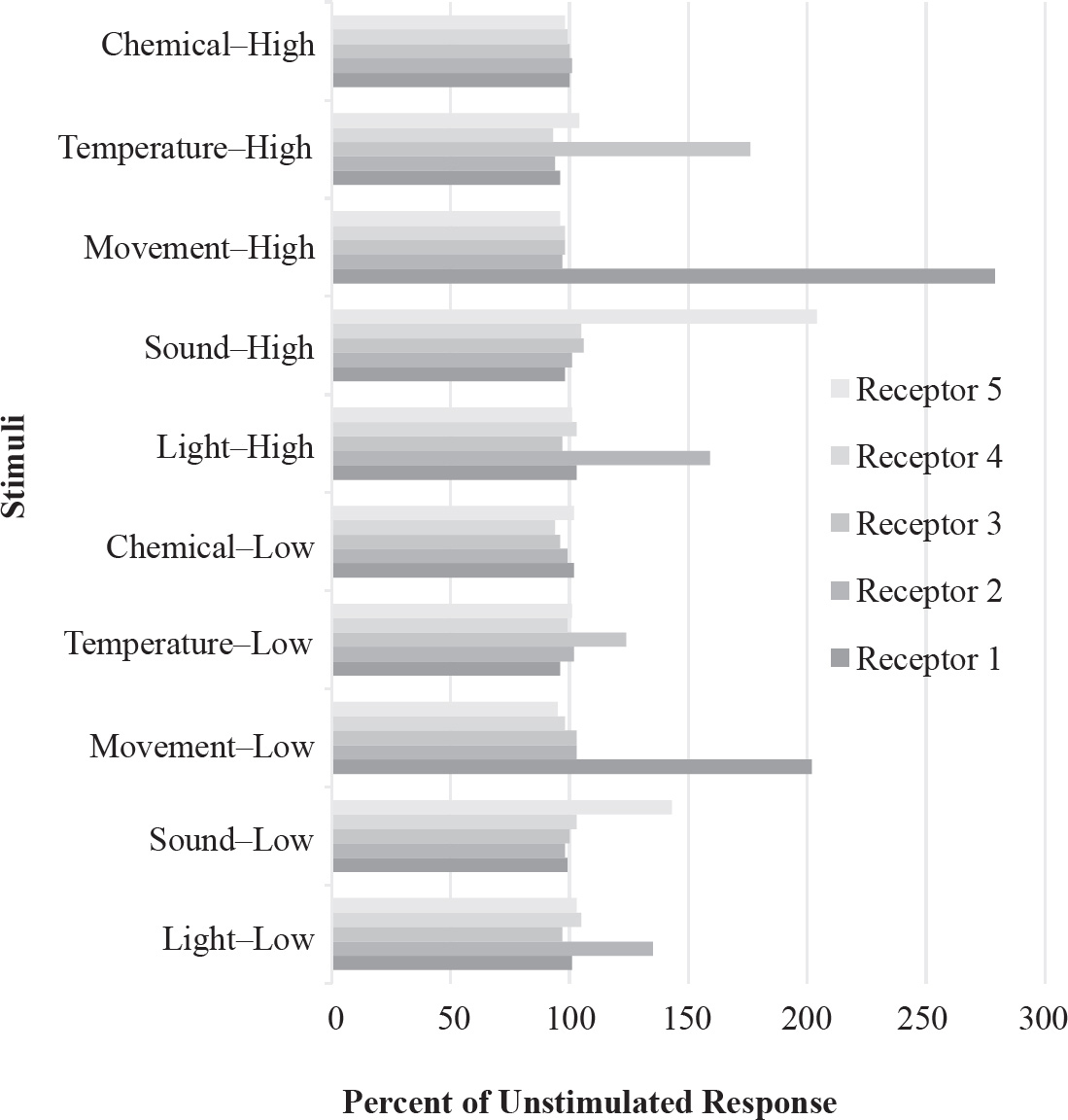
48. What information is required to compile the above results?
(A) The total response to all levels of stimuli
(B) The exact magnitude of each stimulus
(C) The magnitude of response in the absence of a stimulus
(D) The largest response of each receptor to a stimulus
49. Receptor 4 could be which type of receptor?
(A) Movement
(B) Sound
(C) Pressure
(D) Temperature
50. Where in the body would you likely find Receptor 1?
(A) The walls of the aorta
(B) The retina of the eye
(C) The inner ear
(D) The mammary glands
Questions 51–53 refer to the following passage and figure.
The following graph represents the amount of antidiuretic hormone released by the posterior pituitary to regulate blood volume and the corresponding overall blood osmolarity at the same time.
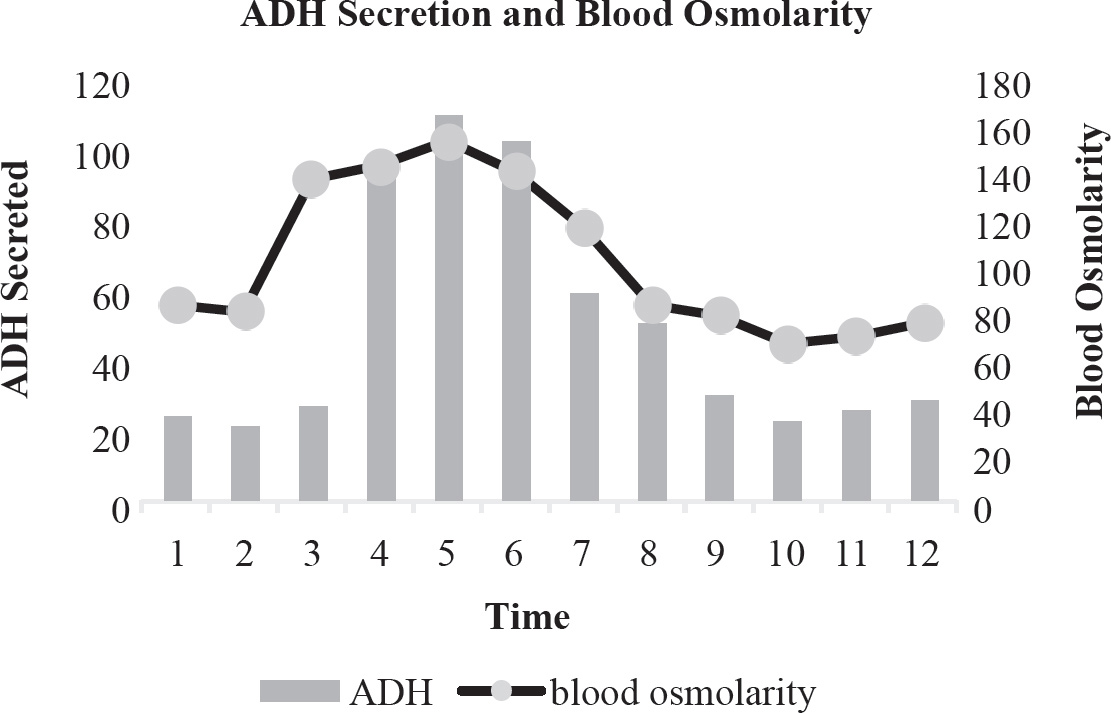
51. Which of the following likely occurred at 2 hours?
(A) The person drank a very large glass of water.
(B) The person drank a beverage isotonic to their blood.
(C) The person drank a beverage hypotonic to their blood.
(D) The person drank a beverage hypertonic to their blood.
52. Which best describes the relationship between blood osmolarity and ADH?
(A) Blood osmolarity increases upon ADH secretion.
(B) ADH increases upon increased blood osmolarity.
(C) ADH has a longer effect than blood osmolarity.
(D) Blood osmolarity has a longer effect than ADH.
53. Caffeine is an inhibitor of ADH. Which of the following would you expect to occur to the blood osmolarity if the person above had consumed caffeine?
(A) The osmolarity would increase higher than usual and then decrease.
(B) The osmolarity would increase the same amount, but would not decrease.
(C) The osmolarity would decrease and then increase.
(D) The osmolarity would decrease, but would not decrease.
Questions 54–55 refer to the following passage.
Mitochondria have an outer and an inner lipid bilayer. Within the inner membrane of the mitochondria sits the electron transport chain. There are four transmembrane proteins that span the membrane. Another protein, cytochrome C, is a peripheral membrane protein on the inner mitochondrial membrane, facing the intermembrane space. Within the membrane itself is a sixth member, a lipid molecule called ubiquinone. The members in the chain pass electrons sequentially, and the transmembrane segments pump hydrogen ions from the matrix into the intermembrane space.
54. Which of the following likely describe cytochrome C and ubiquinone?
(A) Both are mostly hydrophobic.
(B) Both are mostly hydrophilic.
(C) Ubiquinone is hydrophilic, and cytochrome C is mostly hydrophobic.
(D) Ubiquinone is hydrophobic, and cytochrome C is mostly hydrophilic.
55. If cytochrome C were labelled with a dye consisting of large hydrophilic molecules that were allowed to diffuse over time, which regions would be dyed?
(A) Extracellular Space
(B) Cytosol
(C) Intermembrane space
(D) Matrix
Questions 56–58 refer to the following passage and figures.
DDT is a powerful insecticide that was first shown to kill the Colorado Potato beetle, an invasive species in Europe. The pesticide has been shown to be effective at eradicating mosquitoes of the genus Anopheles, which transmits malaria. DDT breaks down over time into a similar chemical called DDE. Between the years 1986 and 2003, scientists sampled three River systems in the Sierra region of Ecuador, while also measuring the egg shell thickness of Andean Condors. They worked with epidemiologists at nearby health centers and obtained the data in Figure 2.
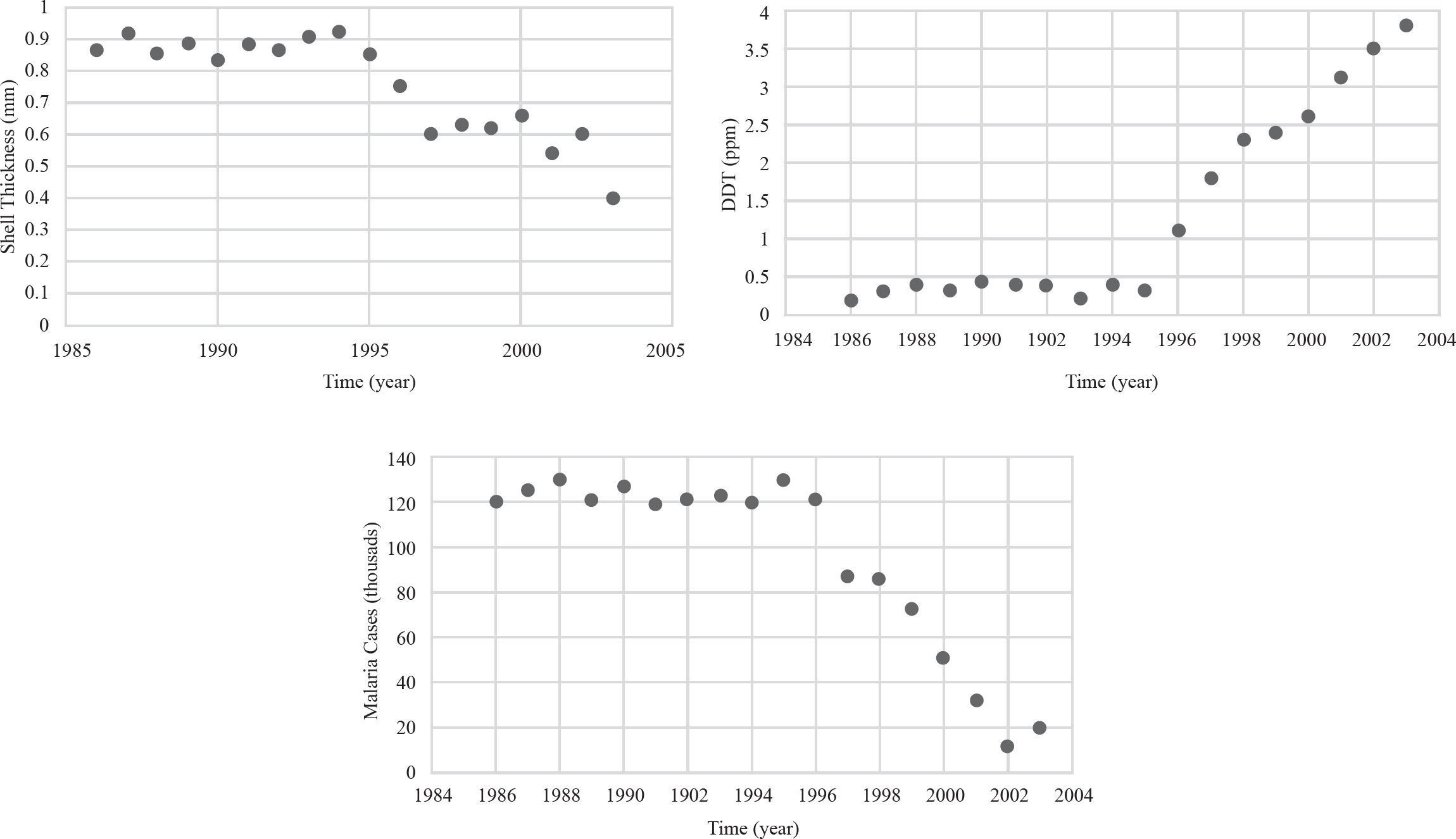
56. Which of the following is the most likely explanation for the data in the figures?
(A) DDT affects how calcium carbonate, which is responsible for the strength of eggshells, deposits onto biological membranes.
(B) Ecuadorian mosquitoes have a mutation that causes them to become stronger when exposed to DDT, which enabled them to infest Condor nests.
(C) Andean condors fish from the river systems, removing the predators of most mosquitoes.
(D) DDE has been shown to have strong effect on insect infestations, and Condors rapidly adapt to new food sources following exposure to the pesticide.
57. In the year 2007, 60,000 malaria cases were reported. What was the likely source of these numbers?
(A) Andean condors began to nest in the Costa region of Ecuador to escape the challenges to the species.
(B) Due to an atypically warm and humid summer, there were more mosquitoes than usual.
(C) Scientists introduced an additional pesticide to the region to protect the Andean Condors.
(D) Mosquitoes resistant to DDT emerged in the early 2000s and were naturally selected over time.
58. What concentration of DDE would decrease eggshell thickness by 20%?
(A) 0.2 ppm
(B) 1.1 ppm
(C) 2.6 ppm
(D) 3.2 ppm
Questions 59–60 refer to the following passage.
Many species of fish are famous for their shoaling or schooling behavior. Shoaling is intentional grouping behavior, but each fish still moves independently within the shoal. In schooling behavior, the fish are grouped and they swim in the same direction and change direction in a coordinated manner.
59. Some people believe that schooling helps fish conserve energy, similarly to birds flying in formation. This is thought to be due the physical forces acting on the entire school rather than just upon a single fish. Which of the following would DISPROVE this theory?
(A) Fish in the school take longer to fatigue than solitary fish.
(B) The combined water resistance of a school of fish is less than that of the combined water resistances of solitary fish.
(C) Fish swimming away from a predator survive longer in school formation than they did in shoal formation.
(D) The heart rates of fish in the school are higher than those in the shoal after swimming the same distance.
60. If extraterrestrials came to Earth, which human behavior is most like schooling?
(A) Many cars traveling on a winding highway in the same direction
(B) A large group of people gathering to watch a parade
(C) A group of school children on a playground
(D) A couple in a ballroom dancing competition
Questions 61–63 refer to the following passage.
The unit of contraction within skeletal muscle cells is called the sarcomere. A sarcomere contracts when the filamentous protein myosin stretches into a high-energy conformation and binds to the filamentous protein actin. When the myosin returns to its low-energy, relaxed conformation, actin is pulled, and the sarcomere contracts. The following steps relate ATP to each step of this process.
1—Myosin binds to actin (ADP is attached)
2—Myosin returns to low energy conformation (ADP is released)
3—Myosin releases actin (ATP binds)
4—Myosin stretches to high energy conformation (ATP is hydrolyzed)
61. What is bound to myosin when it is in its high-energy conformation?
I. Actin
II. ATP
III. ADP
(A) II only
(B) III only
(C) I and II
(D) I and III
62. If the cell runs out of ATP, what would be the state of the sarcomere?
(A) Myosin is bound to actin in the high-energy conformation.
(B) Myosin is alone in the high-energy conformation.
(C) Myosin is bound to actin in the low-energy conformation.
(D) Myosin is alone in the low-energy conformation.
63. A calcium ion is required for the binding of myosin to actin. If a calcium chelator, such as EDTA, is added to a muscle cell, which of the following graphs shows how it will affect muscle contraction?
(A) 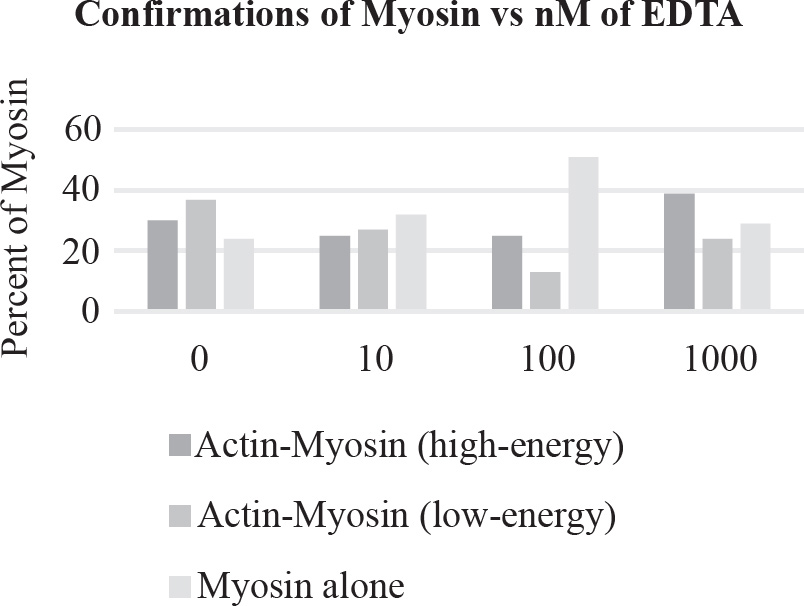
(B) 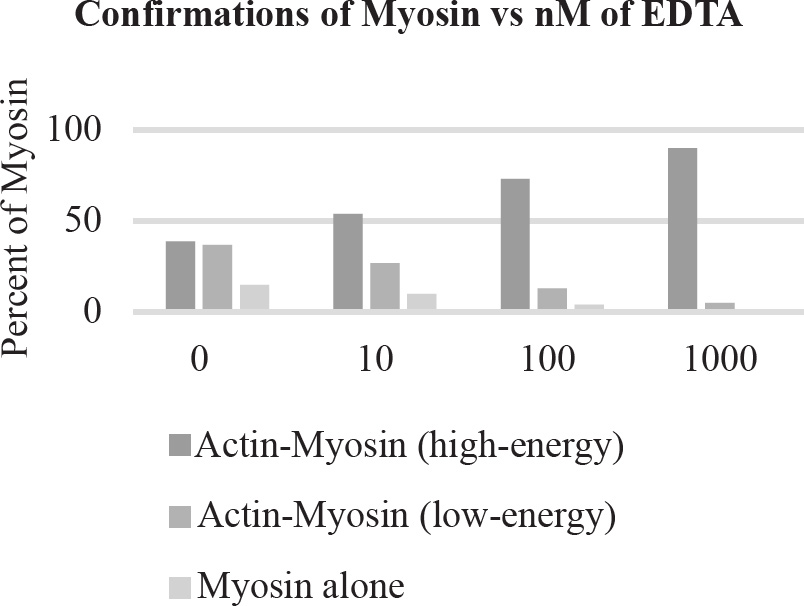
(C) 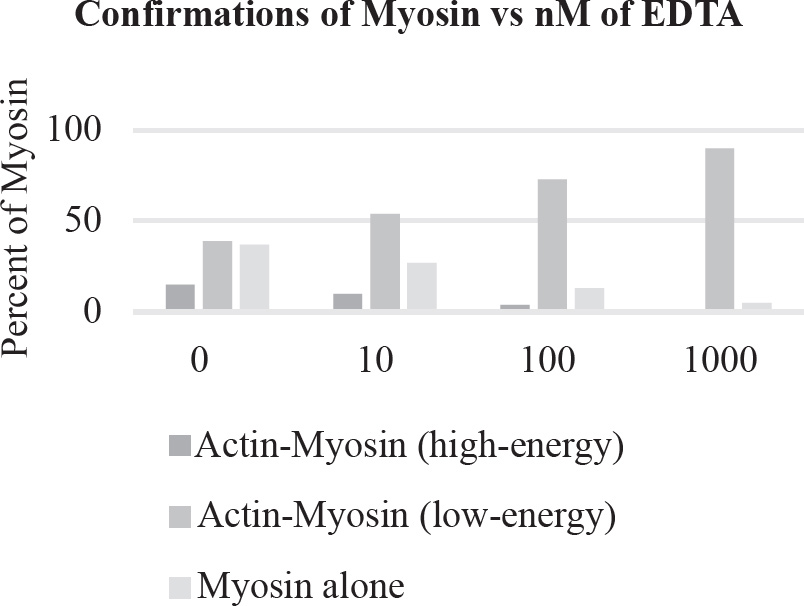
(D) 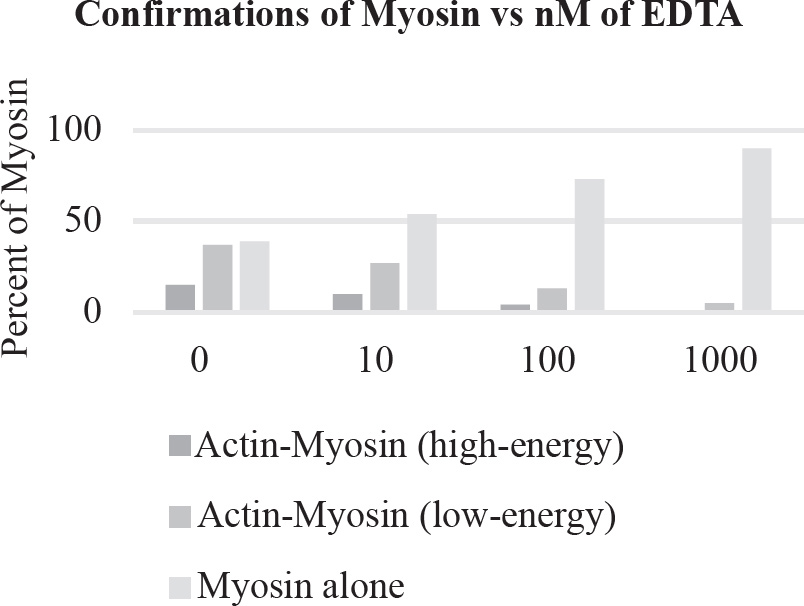
Directions: Part B consists of questions requiring numeric answers. Calculate the correct answer for each question.
64. You are given a large bottle of 20 mM NaCl solution and you need to make 10 mL of a 15 mM solution. What volume of the 20 mM NaCl will you need to make your diluted (mL) solution?

65. In order to measure your solution, you find an old graduated cylinder. However, the volume readings on the side of the cylinder are worn away, and all that exists are the lines. You have a ruler and you measure the height of the cylinder from base to the top line as 39.81 cm and you measure the diameter to be 2 cm. Calculate the volume reading in mL (cm3) of the lowest line on the cylinder, assuming the lines are evenly spaced. Use 3.14 to represent π in your calculation. Your answer should contain 4 digits.

66. In the following cladogram, a common ancestor (*) and species derived from it are illustrated. How many species have four or more common ancestors with Iguanodon?

Questions 67–68 refer to the following passage.
Blood type is determined by two unlinked genes. The first gene can code for protein A, protein B, or no proteins (called O). The second gene can code for the Rh protein (Rh+) or code for no Rh protein (Rh-). A person will form antibodies against any blood proteins that are different from his or her own.
A woman with the blood genotype A/A; Rh-/Rh- and a man with the blood genotype B/O; Rh+/Rh- have a child.
67. What percentage of their children are likely to have the A blood type? Give answer as a decimal.

68. What percentage of the children will be able to donate blood to their mother? Give answer as a decimal.
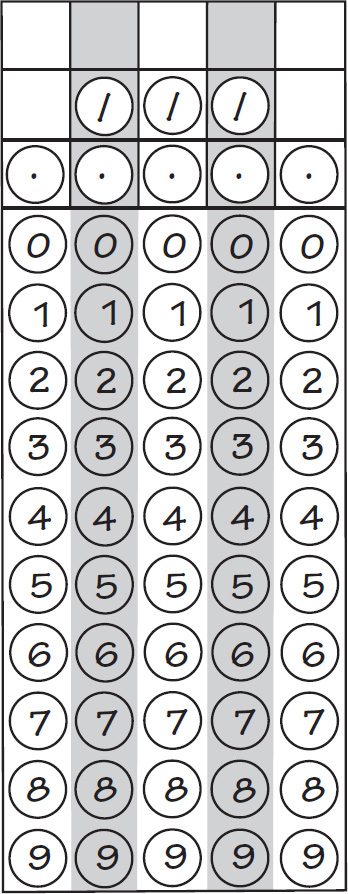

69. In a closed population, squirrel teeth can be either long or short. The long teeth allele is classically dominant. If there are 80 squirrels with long teeth and 40 squirrels with short teeth, what is the frequency of heterozygotes? Round to 2 decimal places.

STOP
END OF SECTION I
IF YOU FINISH BEFORE TIME IS CALLED, YOU MAY CHECK YOUR WORK ON THIS SECTION. DO NOT GO ON TO SECTION II UNTIL YOU ARE TOLD TO DO SO.
BIOLOGY
SECTION II
8 Questions
Planning Time—10 minutes
Writing Time—80 minutes
Directions: Questions 1 and 2 are long free-response questions that should require about 22 minutes each to answer and are worth 10 points each. Questions 3 through 8 are short free-response questions that should require about 6 minutes each to answer. Questions 3 through 5 are worth 4 points each, and questions 6 through 8 are worth 3 points each.
Read each question carefully and completely. Write your response in the space provided following each question. Only material written in the space provided will be scored. Answers must be written out in paragraph form. Outlines, bulleted lists, or diagrams alone are not acceptable.
1. There are four categories of macromolecules.
(a) Identify the four categories and provide one specific example of each.
(b) Describe the basic structure and function of each type. Be sure to include the individual building blocks and how they come together to form larger polymers and explain how their structure is important for their function.
2. While hiking along a looping forest trail in a mountainous region, you take pictures of the beautiful things around you. Later, you notice that a particular type of plant with tiny white flowers is very prevalent in your pictures at the beginning of the trail and the end of the trail, but that an identical looking plant with tiny red flowers is prevalent along the middle of your journey along the trail.
(a) Describe how you can determine if they are the same species.
(b) Discuss reasons why the red plants are only in the middle and the white plants are only at the beginning/end of the trail loop.
(c) Create three hypotheses for the distribution of the plants.
(d) Describe how you would test each hypothesis in an experiment (or separate experiments).
(e) Select one hypothesis that you will pretend is true and show example data (either in table or graph form) from your experiments that would lead you to that conclusion.
3. Describe three of the unique properties of water that make it essential to life and explain how each of these properties contributes.
4. There are two types of synapses in the body: physical and chemical. Physical synapses are direct physical connections between cells that allow them to pass information between them. Chemical synapses are connections where neither cell is in physical contact with the other cell, but information can still be passed from one cell to the other. Explain how a chemical synapse functions and where you might find one in the body. Be sure to include the parts of the two cells involved and how the information is sent and received. You may draw a synapse for your response with clear text descriptions.
5. Describe evolutionary fitness and explain how an adaptation could increase evolutionary fitness for a species even though it leads to an increased rate of predation for that species.
6. In the lungs, a large surface area is essential to maximize gas exchange (In humans, the surface area in the lungs could nearly cover a tennis court!), but the volume of the lungs must be kept relatively small so they fit into the chest cavity. Given the two cubes below, explain which of them would be better at gas exchange. Include the surface area-to-volume ratio in your explanation.
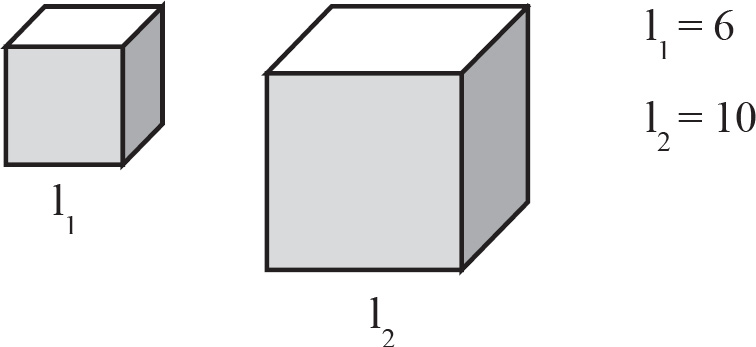
7. Identify examples of producers and different levels of consumers in the following food web. Describe the impact of an invasive species that competes with the phytoplankton and zooplankton and has no natural consumers. Using one species as an example, create a graph showing mock population levels before the arrival of the invasive species and the impact on population afterwards.
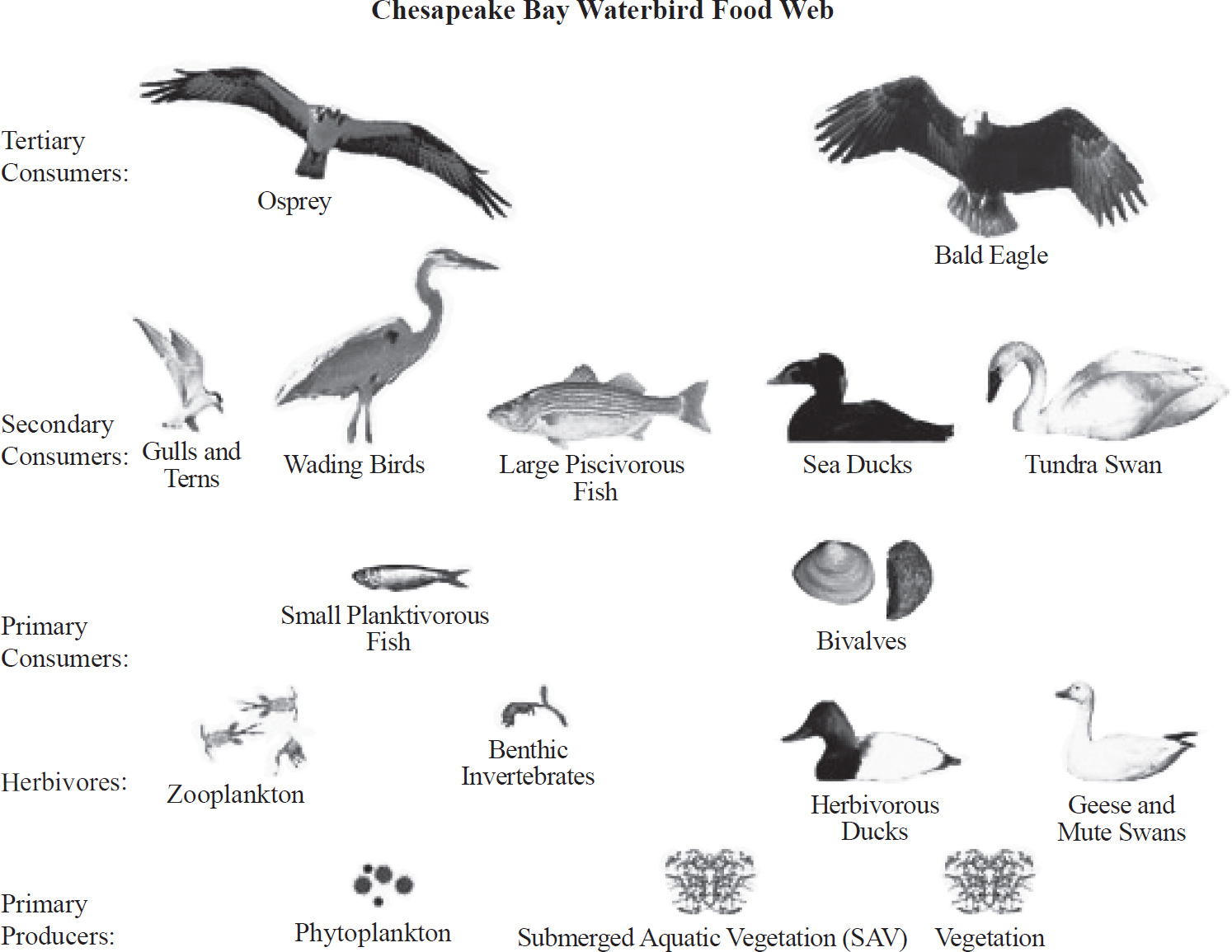
8. DNA errors can cause mutations to occur at any time. Describe the difference between germline and somatic cells in the human body and explain the difference in repercussions of a mutation in a germline cell and a mutation in a somatic cell.
STOP
END OF EXAM
Practice Test 4: Answers and Explanations
PRACTICE TEST 4 ANSWER KEY
1. B
2. B
3. A
4. D
5. C
6. A
7. C
8. B
9. A
10. C
11. B
12. C
13. A
14. B
15. D
16. C
17. D
18. D
19. C
20. D
21. B
22. A
23. B
24. A
25. A
26. C
27. B
28. B
29. C
30. A
31. A
32. D
33. D
34. D
35. A
36. B
37. A
38. A
39. D
40. C
41. B
42. A
43. D
44. D
45. D
46. A
47. A
48. C
49. C
50. C
51. D
52. B
53. B
54. D
55. C
56. A
57. D
58. B
59. D
60. A
61. D
62. C
63. D
64. 7.5
65. 25.00
66. 6
67. 0.5
68. 0.25
69. 0.44
PRACTICE TEST 4 EXPLANATIONS
Section I: Multiple Choice
1. B If it absorbs the light from 400–450 nm and 625–675 nm, then it is reflecting the other wavelengths. The reflected wavelength is the color the plant appears to us.
2. B The best wavelengths would be the ones that the pigment absorbs. Carotenoids absorb at 450–500 nm, so that is where they would best perform photosynthesis.
3. A If the plant absorbs green to red and does not absorb violet to blue, then it reflects violet to blue. Of the choices, it would appear violet.
4. D A morula is totipotent, but it still is derived from a totipotent stem cell. In fact, because the original cells of a zygote are totipotent, all cell types come from a totipotent stem cell.
5. C Trophoblast is more specialized, which means it is more differentiated than a morula.
6. A siRNA is used to reduce the mRNA of something. A non-targeting RNA has no specific target. It is included as a control to show that the process of the RNAi does not cause a problem. Instead, the problems to viral infection are from the particular gene that is reduced by the siRNA and not just a general effect. Choice (B) can’t be correct because the NT siRNA did not reduce the viral infection; if all genes were reduced, the infection should have been reduced. Choice (C) is possible, but because the axis is a percentage, it seems likely that some number could be plotted, even if it were small. Choice (D) is incorrect because NT siRNA does not increase viral infection much.
7. C It is likely that TNS 1, TNS2, and TNS4 are involved because when their mRNA was decreased the viral infection decreased. No conclusion can be made about TNS3 because the siRNA did not successfully reduce the mRNA.
8. B Reducing a gene known to be involved in viral infection would be a good positive control to show that reducing a gene involved in infection reduces the infection rate. Therefore, (A) can be ruled out. An siRNA for a gene making little mRNA would not cause an impact, so it is the correct answer (B). siRNA for a gene not known to be involved would also be a good positive control because it would show the result that is expected when something is not involved in viral infection (C). A second siRNA would be helpful because it would confirm what the first siRNA shows (D). More replicates are always better.
9. A In 1929, scientists did not know that DNA was the thing being passed in the transformation, so (B) is incorrect. Griffith did not think the heat-killed bacteria were brought back to life, but he thought they transformed the non-virulent bacteria into virulent bacteria (C). Virulence is the presence of an ability, so it seems like it won out. However, this is just because virulence is the presence of a trait and non-virulence is the absence of a trait.
10. C They concluded that the DNA fraction caused the transformation. If the DNA fraction was not pure DNA, this would negate their conclusion (C). The lipid fraction did not cause the transformation, so it doesn’t matter much what it had (as long as it was not shown to have DNA, but this answer doesn’t say that). If the RNA had a trace of DNA (B), it would hinder their conclusion because it should have slightly transformed the bacteria. However, (C) is a better choice. The size of each fraction is irrelevant (D).
11. B In the process above, DNA gives new traits to bacteria. The bacterial traits are determined by the DNA present. Acquisition of DNA does not necessarily mean that new traits will be acquired (A). What if a bacteria was given the same DNA that it already has? DNA is not capable of giving any species traits. The example above is in the same species of bacteria (C). Bacteria are not revived by new DNA. The living bacteria were just transformed (D).
12. C DNA polymerase adds bases to the 3′ side of a chain. So, it will travel towards the 5′ end of the template, which would be the 3′ end of the growing chain. It always builds a complement rather than building a copy. The reason the DNA gets copied is because each strand is used to build a new partner strand, not a direct copy.
13. A The direction the polymerase goes is the opposite on each strand of the helix. Because the helix unwinds in one direction, only one of the strands can be continuously built. The strands are not opening asynchronously (B). DNA replication is not producing mRNA (C). RNA primer is required, but this is not what determined the timing or non-continuous nature of the lagging strand synthesis (D).
14. B The picture shows the number of bonds, showing that A-T has 2 hydrogen bonds, whereas G-C has 3. More bonds would make something stick better. A-T is easier to separate.
15. D To judge what would happen without the fire, we should look at the trend before the fire. The white-flower shrub population was constant, so (A) is incorrect. The shade intolerant shrub population was low without any obvious sign of increase, and the conifer population was slightly decreasing (B). The conifer population was shrinking, so it is unlikely conifers would claim territory (C). Hardwood tree population was increasing, and conifer population was decreasing, so (D) is correct.
16. C The red flowers had high numbers and were increasing before the fire, while the white flowers had fewer numbers and were stable. After the fire, the white flowers did not lose numbers; instead, their population greatly increased. It is possible red flowers were selected because there were more before the fire (I). The white flowers seemed to tolerate the fire, so that particular selective pressure gave them fitness over the red flowers killed in the fire (II). Since the white flowers did not have great numbers before the fire, they were not always the most fit. They were only more fit than red when the selective pressure of the fire arrived (III). Therefore, I and II are true.
17. D The saltwater fish likely loses water to the saltwater, and the freshwater fish likely takes on too much water through osmosis. The urine from the freshwater fish is not meant to increase the osmolarity of the lake (A); it is meant to get rid of excess water so the inside of the fish is not too diluted. The blood osmolarity in the saltwater fish is still medium, so it is not okay with large amounts of salt (B). The freshwater fish needs to get rid of the salt, but keep the water. The freshwater fish is not necessarily “drinking” the water (C); it gains it through osmosis because the inside of the fish is saltier than the freshwater. Choice (D) is correct.
18. D The charts shows the blood osmolarity of the freshwater and the saltwater fish to be about the same. Salmon should maintain a fairly constant blood osmolarity as well. Choices (A) and (B) can be eliminated. Freshwater salmon should have a higher urine volume (D).
19. C The percentage of female offspring would be 160/200 = 80%. This closely matches the 30 degrees on the graph.
20. D Below 29 degrees there are more males, and above 29 degrees there are more females. Therefore, the enzyme that determines female sex must be active at above 29 degrees. Choice (A) shows an enzyme most active at 29. However, its activity is the same at 27 as at 31, so this does not explain the differences in sex at these two temperatures. Choice (B) has a peak around 25 degrees, and the enzyme level is mostly gone starting around 29 degrees and above. Choice (C) is active at even lower temperatures. Choice (D) gets its activity starting around 29 degrees and is active at the hotter temperatures as well.
21. B The passage says the two genes are overexpressed in tumor cells. This means they are on normal cells, but there are more of them on tumor cells.
22. A The estrogen receptor is responsible for receiving estrogen and then causing changes within the cell. If the ER is mutated, then it should not bind estrogen well (A). If estrogen binding leads to cell division, then no estrogen binding would lead to less cell division.
23. B Tamoxifen is a drug that treats tumor cells with too many estrogen receptors. It must function to prevent estrogen binding (A), or decrease the number of ER (C), or block the downstream function of ER (D). It will not increase estrogen, because that would make the problem worse (B).
24. A IgG is shown to be elevated by the primary infection and remain elevated. It is high when IgM is not, so it is not likely stimulated by IgM (B). The graph shows nothing about nonresponsiveness. It also says nothing about switching isotypes. The IgG likely exists on the memory cells that linger around as a militia in case the pathogen returns.
25. A A pentamer of IgM can bind 10 of the same antigens, 2 on each of its 5 monomers. It cannot likely be made without ATP (C). It can only bind one kind of antigen (D). It cannot cross membranes unassisted because it is a large protein (B). It can cross-link several antigens (A).
26. C In the tree, the upper and lower castes do have a common ancestor (A). The French are not an ancestor of Indians or Poles at all (B). Cambodians and Vietnamese have a common ancestor that Chinese do not have because the lineage for Cambodians and Vietnamese branched off together from the lineage that formed Chinese. The two Mbuti pygmy branched off early, but they are not necessarily the most closely related.
27. B Tsonga and Japanese have 4 common ancestor nodes on the tree and the initial common ancestor.
28. B RNA-dependent DNA polymerase uses an RNA template and makes DNA. If an RNA virus needed to integrate into a DNA genome host, it would need to make DNA from RNA.
29. C DNA is permanent and gets passed from generation to generation. RNA is a transient, cheap copy of DNA used for a short time. Both thymine and uracil can base pair fine. Both structures require the sequence to be accurate, but it is less important for RNA to be perfect. All strands of RNA are built similarly and then they fold differently to form different RNA molecules (D).
30. A They change because they acquire errors. These changes are not by choice, but sometimes they are helpful and get naturally selected for.
31. A RNA polymerase makes RNA, so (C) can be eliminated because it has thymine. The sequence is never copied directly (B), but it is used as a template to build a complementary strand. The enzyme builds by adding to the 3′ side, so it would start on the 3′ side of the template and go towards the 5′ end of the template, which is the 3′ side of the growing strand.
32. D If there were less rhizobia bacteria fixing nitrogen, there would be less nitrogen. DNA and RNA would be affected by the lack of nitrogen because they have nitrogen in their nitrogenous bases, and proteins would be affected because the amino acids have an amine group that contains nitrogen. Carbs would be the least affected, but without DNA, RNA, and proteins things would be in bad shape.
33. D The relationship is mutualistic if both organisms are benefiting. If either is harmed, then it would not be be mutualistic, so (A) and (B) are incorrect. The legumes are already benefiting, so more benefit would not change things (C), but if the bacteria benefited, then the relationship would be mutualistic (D).
34. D To compare the two processes, each type of nitrogen should be used to grow different plants. Choice (A) has two groups of plants. One gets one type of nitrogen, and the other gets the other type. This doesn’t work because the two groups are not identical in all ways except the nitrogen. Choice (B) is okay, but it would be better with replicates of each plant type. Choice (C) has several of each species, but they are not shown to be identical in all the ways. Choice (D) has replicates, and the two groups are shown to be identical in all ways.
35. A A mutation in trpR could cause the repressor to change shape and not bind as well. The RNA polymerase binding would not change (B). The tryptophan levels do not normally cause an increase in transcription (C). Tryptophan does not bind to the operator (D).
36. B The level of tryptophan changes, which is meant to change the levels of transcription so tryptophan and the repressor don’t need to be in equal parts (A). There also is not ongoing tryptophan synthesis (C). RNA polymerase is not described as having more than one promoter (D). The repressor must always be expressed though, so it is there if tryptophan is or isn’t there. trpR must always be expressed.
37. A Without tryptophan, the repressor is bound to tryptophan at the operator, preventing transcription.
38. A The A-E genes are for synthesis of TRP, so when TRP is high they should be low and when TRP is low they should be high. This eliminates (B) and (C). The repressor should always be high because it is always expressed (A).
39. D The virus was capable of making some bacteria resistant to ampicillin. This is likely because it received ampicillin resistance when it previously infected bacteria that were resistant to ampicillin. However, the virus has never infected bacteria that are resistant to tetracycline, so the bacteria would die when exposed to it.
40. C Regulated secretion is when the proteins are made but are held before released. This would give the ability to respond in a more limited and specific manner (C), but it would not affect how they pass through the membrane or how big they are. It would not allow them quicker release because they are waiting on a signal.
41. B When proteins are built, they are built into the membrane of the endosome because the transmembrane segments must be in the hydrophobic space. Choice (B) is the only one that puts the membrane protein in the membrane
42. A Lysosomal proteins should be able to withstand the enzymes in the lysosome (A). It does not matter how stable it is or which side of the membrane it is on.
43. D A pump moves things against the concentration gradient. This makes it active transport.
44. D Impulses always travel from dendrite to soma to axon. This eliminates choices (A) and (B). The inside of the cell becomes more positive as the sodium rushes in.
45. D A failing sodium potassium pump will mean that potassium is not pumped in and sodium is not pumped out, so (B) and (C) are true. Action potentials cannot occur without the resting membrane potential the pump creates, so (A) is true also. Choice (D) is not true. The inside would become less negative.
46. A Wobble pairing occurs because the anticodons on the tRNA contain nucleotides that don’t mind as much which base in the codon pairs to them. This means that each anticodon might have several possible codons (A). There are more amino acids than nucleotides (B), but this is not a cause of the wobble pairing. There are no codons that could not have several anticodons (D). The inosine base would be in the mRNA (C).
47. A The mRNA is read 5′ → 3′, so if the mRNA and tRNA are oriented antiparallel, then the 3′ of the mRNA is oriented with the 5′ side of the tRNA. The wobble pairing is with the 3rd position on the codon, which would be the 5′ end of the tRNA. Choice (D) is incorrect because even though tRNA folds up, it is still a linear chain.
48. C The results are based on the percentage of an unstimulated response, so the response to no stimulus (C) must be calculated.
49. C Receptor 4 is never above the unstimulated levels (100%), so it was not stimulated by chemical, temperature, movement, sound, or light. Receptor 4 could only be a receptor for pressure (C).
50. C Receptor 1 responds to movement, so it is likely found in the ear.
51. D At 2 hours, the blood osmolarity increases. This means the person must have drunk something hypertonic to his or her blood.
52. B The relationship between blood osmolarity and ADH shows that after blood osmolarity increases there is an increase in ADH (B). It is not the other way around because the ADH increased after the osmolarity (A). Neither clearly has a longer effect.
53. B Without ADH, the person would not be able to respond to the high osmolarity. Choices (C) and (D) can be eliminated. The osmolarity should not increase more than before because that was dependent on the beverage that they consumed.
54. D Ubiquinone is inside the membrane, so it is hydrophobic. Choices (B) and (C) can be eliminated. Cytochrome C is a peripheral membrane protein, so it is partly hydrophobic but mostly hydrophilic.
55. C Cytochrome C is in the intermembrane space, so that space would be dyed. If the dye has large hydrophilic molecules, then it would not leave the intermembrane space.
56. A The mosquito levels and eggshell thickness decrease when the DDT increases. It is unlikely the mosquito predators disappeared (C). It does not seem like condors adapted because their eggshells became brittle (D). The mosquitoes do not seem to be stronger because their population dwindled (B). The DDT is harming the eggshells (A).
57. D More malaria means more mosquitoes, but a large year in mosquitoes could still be combated by DDT. The likely cause is DDT resistant mosquitoes leading to high mosquito numbers. An additional pesticide would lead to fewer mosquitoes (C). The bird nesting would not affect the malaria cases.
58. B Eggshell thickness is at approximately 90% at the beginning of the graph. To decrease it by 20% would bring it to 72% (90 × 0.8 = 72%). The graph shows that just after 1995 thickness was at 72%. The DDT in use at that time was around 1 ppm or a little higher.
59. D To disprove the theory, the schooling behavior would have to be proved to not conserve energy. If schooling fish take longer to fatigue, survive longer swimming away from a predator, or experience less water resistance, schooling would be helping them conserve energy. If they had elevated heart rate while schooling, that would show that they are working harder and are not conserving energy (D).
60. A Schooling requires the group to travel in the same exact directions. Choices (B) and (C) would not be moving in the same direction. Those are more like shoals. The couple dancing is moving the same direction, but they are only two people. The traffic is more like a school of people.
61. D In its high-energy conformation, myosin is bound to ADP. ATP is hydrolyzed as it enters the high-energy state, so it should only have ADP bound once it gets there. It will also bind actin.
62. C Without a fresh ATP to bind, myosin will be bound to actin in the low-energy conformation and be unable to release it.
63. D Without calcium, myosin cannot bind to actin. With more EDTA, there should be more myosin alone, as shown in (D). The states bound to actin will disappear.
64. 7.5 The formula page gives the Ci Vi = Cf Vf formula for diluting a stock solution. The stock solution is 20 mM, so that is Ci. The new volume should be 10 mL and the new concentration should be 15 mM, corresponding to Vf and Cf, respectively.
20 mM × Vi = 15 mM × 10 mL
Vi = 7.5 mL
65. 25.00 The volume of mL held in the cylinder from bottom to the top line can be determined with the formula for volume of a cylinder, which is given on the formula sheet as V = πr2h. Then, that volume should be divided by 5 to determine what volume will fill to the bottom line. By plugging into the equation, we get V = 3.14 (1)2 (39.81) = 125.0034 mL. When divided by 5, each division on the cylinder holds 25.00068 mL
66. 6 Iguanodon had one common ancestor with Hypsilophodon before Hypsilophodon diverged. Iguanodon had two with Tenontosaurus because one come from where Tentontosaurus diverged and they both share the common ancestor with Hypsilophodon. By that logic, Iguanodon has three common ancestors with Zalmoxes, four with Dryosaurus, and five with Camptosaurus. It will also share those five common ancestors with the other species on the tree, because everything above Camptosaurus still shares those original five. The higher levels also share the ancestor that existed when Iguanodon diverged. Either way, six species share at least four common ancestors with Iguanodon. Remember, don’t count Iguanodon itself.
67. 0.5 The crosses are shown below for each of the genes. The percentage of children with A blood is 50%
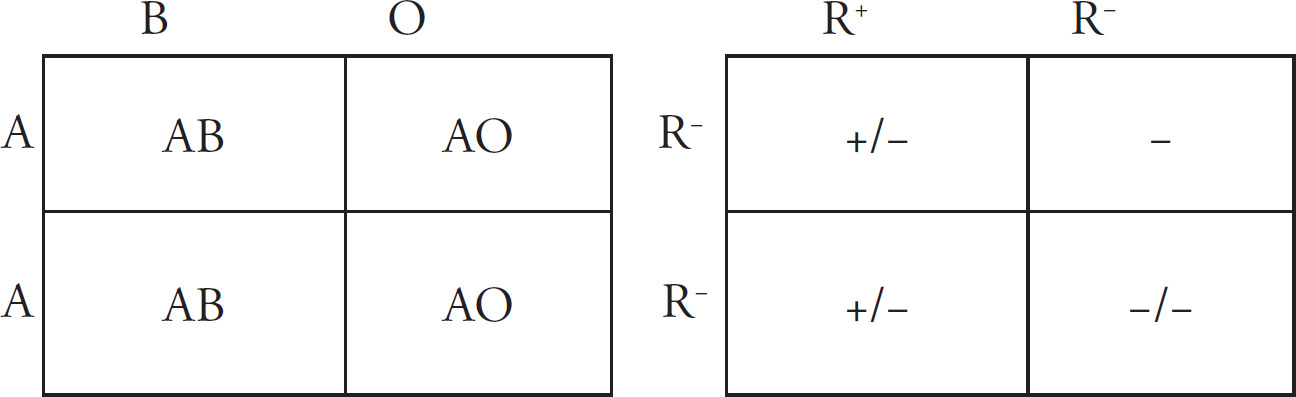
68. 0.25 Only blood types AA or AO and -/- will be able to donate to the mother. The formula sheet shows that for probability of A and B occurring, you must multiply probability of A and the probability of B. The probability of AA or AO blood is 0.5, and the probability of -/- is 0.5.
0.5 × 0.5 = 0.25 = 25%
69. 0.44 There are 40/120 homozygous recessives = the frequency of homozygous recessive = q2, and then q = 0.333333 and if p + q= 1, then p = 0.666667. The frequency of heterozygotes is 2pq = 2 (0.333333) (0.666667) = 0.444442.
Section II: Free Response
On the following pages you’ll find two types of aids for grading your essays: checklists and sample responses. Checklists like these are used to grade your essays on the actual test. They’re actually quite simple to use.
For each item you mentioned, give yourself the appropriate number of points (1 point, 2 points, and so on). Remember that you can only get a maximum of 10 points for each essay. As you evaluate your work, don’t be kind to yourself just because you like your own essay. If the checklist mentions an explanation of structure and function and you failed to give both, then do not give yourself a point. The testing board is very particular about this. You need to mention precisely the things they’ve listed, in the way they’ve listed them, in order to gain points.
What if something you’ve mentioned doesn’t appear on the list? Provided you know it’s valid, give yourself a point. If it was something you pulled out of your hat at the last minute, odds are it’s not directly applicable to the question. However, because you might come up with details even more specific than those contained in the checklist, it’s not unlikely that the example or structure you’ve cited is perfectly valid. If so, go ahead and give yourself the point. Remember that the testing board hires college professors and high school teachers to read your essays, so they’ll undoubtedly recognize any legitimate information you slip into the essay.
The second part of the answer key involves short paragraphs. These are not templates; the testing board does not expect you to write this way. They are simply additional tools to help illustrate the things you need to squeeze into your essay in order to rack up the points. They explain in some detail how the various parts of the checklist relate to one another and may give you an idea about how best to integrate them into your own essays come test time.
If you find that you didn’t do too well on the essay portion, be sure to read carefully the chapter on the free-response questions (Chapter 14). You can use the chapters themselves as checklists. If you find it too difficult to grade your own essays, see if your teacher or a classmate will help you out. Good luck!
Long-Form Free-Response Checklist 1
(10 points total)
I. Identify 4 categories and give an example of each (2 points)
II. Describe proteins (2 points)
III. Describe nucleic acids (2 points)
IV. Describe carbohydrates (2 points)
V. Describe lipids (2 points)
Example of a 10-Point Response
The four macromolecules are proteins, carbohydrates, nucleic acids, and lipids. Hemoglobin is a protein. Glucose is a carbohydrate. DNA is a nucleic acid. A phospholipid is a lipid.
Proteins are built from amino acids that are connected by peptide bonds. There are 20 amino acids. Amino acids each have an amino end and a carboxyl end and a different functional group. They have different characteristics, and when long chains of amino acids come together, they form proteins that can fold up, making unique shapes. These proteins have unique characteristics depending on the amino acids within them. The sequence of amino acids determines the structure of the protein. Many proteins are enzymes with a special job that requires a particular shape.
The main nucleic acids are DNA and RNA. They are formed from chains of nucleotides that connect by phosphodiester bonds. A nucleotide has a nitrogenous base (either A, C, G, T, or U), and a sugar (either ribose or deoxyribose), and a phosphate. DNA stores the genetic information (the genome). It is always in a double helix form. The order of the nucleotides is a recipe for making all the things that the organism needs to make. RNA can take many shapes. mRNA is a temporary molecule that carries a message transcribed from the DNA to the ribosome for translation.
Carbohydrates are sugars. Polysaccharides are built from individual sugar units called monosaccharides. They are made of carbon, hydrogen, and oxygen. Glucose is a monosaccharide. The body uses it for energy storage. Glucose is used in glycolysis at the start of cellular respiration. Carbs have a lot of energy within their chemical bonds.
Lipids are hydrophobic. Many types of lipids have long hydrophobic chains made of carbon and hydrogen. The body uses phospholipids to make membranes because they are amphipathic, which means they are partially hydrophilic and partially hydrophobic. A lipid bilayer is made from two rows of phospholipids making a hydrophobic sandwich. Another lipid is cholesterol, which is used in some types of hormone.
Long-Form Free-Response Checklist 2
(10 points total)
I. Describe how you test if they are the same species (1 point)
II. Discuss why red plants are only in the middle and white plants are at the ends (1 point)
III. Create three hypotheses (3 points)
IV. Describe experiment(s) to test them (3 points)
V. Show mock data to prove one of the hypotheses (2 points)
Example of a 10-Point Response
To test if the flowers were from the same species, you would have to try to breed them together by pollination. If they are the same species, then you should be able to breed them. If you cannot breed them, then they might be different species.
Since the trail is a loop, there must be some feature near the beginning and the end of the trail that is different from the middle. Maybe the trail goes higher on the mountain in the middle, where it is colder or there is more shade. Maybe the soil is a different blend in the two spots. Maybe one part of the trail has more water or a different pH of soil. Maybe one part was burned by fire. Something must be different in the two areas.
After looking at the environment more closely, the beginning of the trail has more sun, warmer temperature, and moist soil. Three hypotheses are:
1—The white flowers prefer warmer temperatures than the red flowers.
2—The white flowers prefer more sun than the red flowers.
3—The white flowers prefer a moister soil than the red flowers.
To test each of these experiments, the different environments would need to be set up and then both colors of flowers should be grown and measured to see where they grow better. Everything should be the same in the two groups except for the one variable being tested. Multiple plants should be included in each group. The height of the plants should be measured throughout the experiment to see which condition causes each plant to grow better. Each group should have multiple white plants and multiple red plants for replicates. The results below show that the differing factor is the level of moisture. Red plants prefer dry, and white plants prefer wet.
| Condition | Average Red Growth | Average White Growth |
| Warm | 20 | 21 |
| Cool | 21 | 23 |
| Sun | 35 | 34 |
| Shade | 36 | 35 |
| Wet | 10 | 40 |
| Dry | 40 | 10 |
Short-Form Free-Response Checklist 3
(3 points total)
I. Describe three properties of water and how they contribute to life (3 points)
Example of a 3-Point Response
Three properties of water are cohesion, low density as a solid, and high heat capacity.
Cohesion is the property by which water connects to other water molecules. This is important because it allows water to move in chains. An example of this is in transpiration, where long chains of water molecules travel up xylem in plants and evaporate out the stomata to pull water out of the roots.
Water is less dense in its frozen state than it is in a liquid state. This means that ice floats to the top of water. Because life began in the oceans and continues in many lakes, it is important that when the water freezes the ice doesn’t sink down and crush the life in the water.
High heat capacity allows water to accept a lot of heat without changing temperature. This makes it important in the body for reactions where it can act as a heat buffer. It also helps humans to remove heat by sweating because the water retains heat.
Short-Form Free-Response Checklist 4
(4 points total)
I. Explain where a chemical synapse would be found (1 point)
II. Explain how it functions and how info is sent/received (3 points)
Example of a 4-Point Response
A chemical synapse would be found between neurons in the body. The cell before the synapse is the pre-synaptic cell, and the part of the neuron that would be there is the axon. The cell after the synapse is the postsynaptic cell, and the part of the neuron that would be there is the dendrite.
Chemical synapses work when the axon of one cell receives an action potential that causes vesicles of neurotransmitter to be released into the synapse. The neurotransmitter diffuses across and binds to receptors on the dendrite of the other cell. Depending on the receptors and the message, the other cell might fire an action potential or it might be inhibited and not fire an action potential.
Short-Form Free-Response Checklist 5
(3 points total)
I. Describe evolutionary fitness (1 point)
II. Explain how adaptations can increase fitness but also increase predation (2 points)
Example of a 3-Point Response
Evolutionary fitness is the ability to reproduce better than others of your species due to heritable traits. This is sometimes due to traits that help survival and thus increase the ability to reproduce. Other times this may be because of sexual selection that directly improves reproduction ability.
If an adaptation increases predation and increases fitness, then it must not be one that simply increases survival. It is likely something that directly increases reproduction, such as the tail on a peacock. The long tails on male peacocks make them a target for predation. However, the female peacocks just love them. Only males with a nice, long tail will get mated with, so the long tail increases evolutionary fitness even though it increases predation.
Short-Form Free-Response Checklist 6
(3 points total)
I. Calculate the surface area-to-volume for each cube (2 points)
II. Explain which cube would be better at gas transport (1 point)
Example of a 3-Point Response
The cube with the larger surface area-to-volume ratio would be the better at gas exchange because it would have a lot of surface area.
To calculate the surface area of each cube, use the formula A = 6s2 The volume of a cube is V = s3.
The small cube has a surface area of 6(6)2 = 216, and its volume is (6)3 = 216. So the surface area-to-volume ratio is 216/216 = 1.
The large cube has a surface area of 6(10)2 = 600, and its volume is (10)3 = 1000. So the surface area-to-volume ratio is 600/1000 = 0.6.
The small cube has a larger surface area-to-volume ratio and would be better at gas exchange because it has more surface area per inside volume space.
Short-Form Free-Response Checklist 7
(4 points total)
I. Identify producers and consumers (1 point)
II. Describe the impact of the invasive species (1 point)
III. Create a mock graph (2 points)
Example of a 4-Point Response
The phytoplankton are producers because they produce energy through photosynthesis. The benthic invertebrates are primary consumers because they eat the phytoplankton. Sea ducks that eat the benthic invertebrates are secondary consumers. Tertiary consumers would be things like eagles that eat the sea ducks.
The phytoplankton and zooplankton would be harmed if the invasive species were competing for resources without being eaten by predators. Everything that relies on the plankton would be affected too. The benthic invertebrates would have no food. The small planktivorous fish would have no food. The wading birds wouldn’t have small planktivorous fish to eat. The entire system would go down because of the invasive species disrupting the balance.
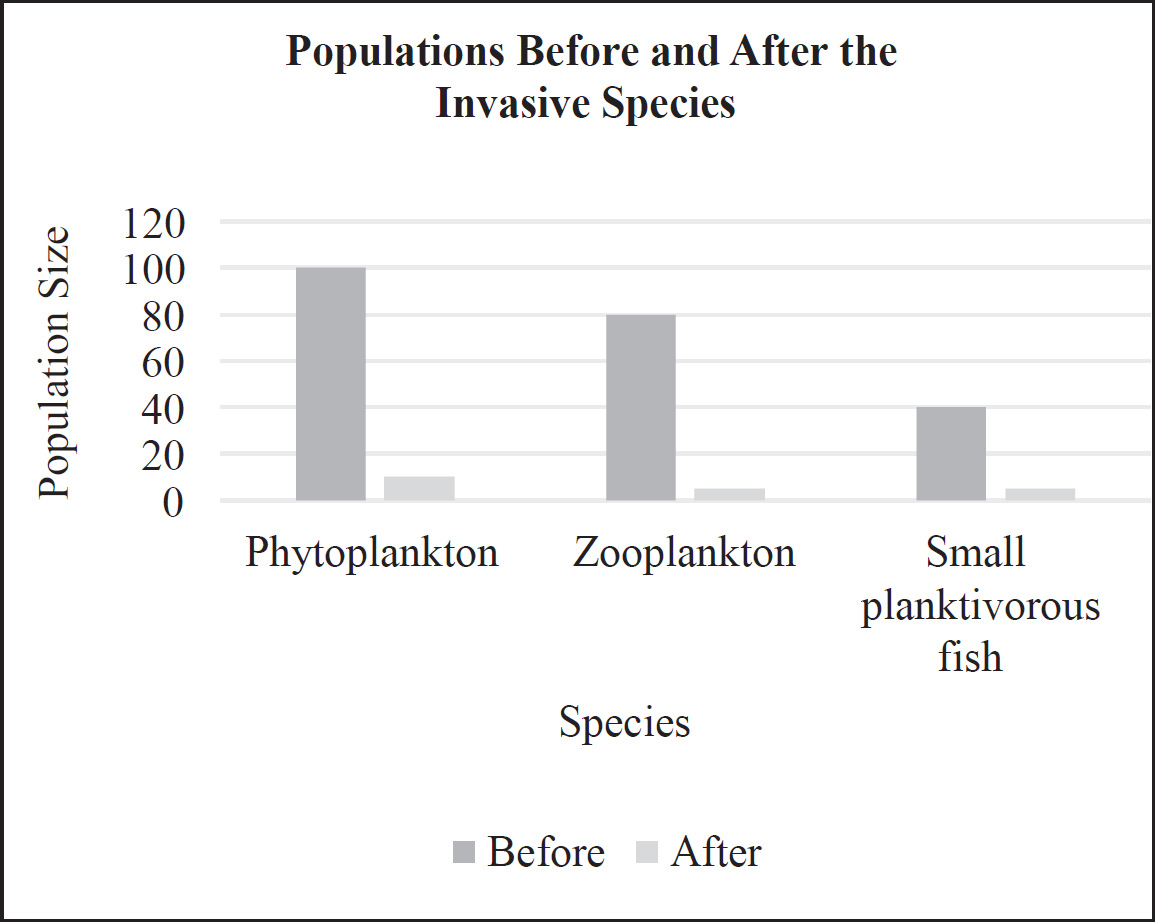
Short-Form Free-Response Checklist 8
(4 points total)
I. Describe the difference between germline and somatic cells (2 points)
II. Explain a germline mutation and a somatic cell mutation (2 points)
Example of a 4-Point Response
Somatic cells are the regular cells found everywhere in the body that make up your tissues and body parts. Your skin cells are somatic cells. The germline cells are special cells that will make gametes. In males, gametes are spermatogonia and the sperm that come from them. In females, these are the oogonia and the eggs that come from them. Gametes are the cells that will be passed on to future generations.
If a mutation occurs in your somatic cells, this will affect you, but it will not be passed on to your children. The mutation will only affect that cell and the cells that that cell divides into. The mutation will not be in all of your other cells. If the mutation is deadly, then the cell that is happened in will just die. One cell dying is no big deal.
On the other hand, if you have a mutation in your germline cells, the mutation will be passed on to your children. They will have the mutation in all of their cells because they will grow from the one fertilized egg that they begin with. Inherited disease occur because at some point there was a mutation in a germline cell that resulted in an entire person being built with that mutation in every single one of their cell. Of course, that affected person’s germline cells will all have the mutation too. All their future offspring that get that mutated gene will have in it every cell as well.

What’s next on
your reading list?
Discover your next
great read!
Get personalized book picks and up-to-date news about this author.
Sign up now.