Biology For Dummies
Part II Cell Reproduction and Genetics: Let’s Talk about Sex, Baby
Chapter 8
Reading the Book of Life: DNA and Proteins
In This Chapter
Grasping why DNA and proteins are so important
Producing proteins
Surveying the various DNA mutations that can occur
Controlling your genes
Without deoxyribonucleic acid, or DNA for short, your cells — and all the cells of every other living thing on Earth — wouldn’t exist. That’s because DNA controls the structure and function of organisms, largely because it’s essential to the production of the proteins that determine your traits. When changes occur in the DNA of one or more cells, the effects on the organism made up of those cells can be disastrous.
We show you just how important DNA and proteins are to your everyday life in this chapter. Get ready to discover how DNA and RNA (ribonucleic acid) work together to produce proteins, the types of DNA mutations that occur and how they affect you, and more.
Proteins Make Traits Happen, and DNA Makes the Proteins
You probably know that DNA is your genetic blueprint and that it carries the instructions for your traits. But what you may not know is exactly how your DNA causes you to look and function a certain way. DNA contains the instructions for the construction of the molecules that carry out the functions of your cells. These functional molecules are mostly proteins, and the instructions for creating these proteins are found in your genes, sections of DNA that lie along your chromosomes.
Think of your cells as little factories that have to carry out certain functions. Each function relies upon the actions of the robot workers in the factory. DNA is the instructions for the construction of each type of robot. If the robots are built correctly according to their instructions, they work the way they’re supposed to. Based on how the robots work, the factory accomplishes specific tasks. Your cells don’t have little robots running around, but they do have worker molecules that carry out the functions of the cell. If one of these molecules isn’t working correctly, your cells will work differently than they’re supposed to, which could affect your traits.
 One gene equals one blueprint for a functional molecule. Because many of the functional molecules in your cells are proteins, genes often contain the instructions for building the polypeptide chains that make up proteins (for more on protein structure, head to Chapter 3). So, one gene often equals one polypeptide chain.
One gene equals one blueprint for a functional molecule. Because many of the functional molecules in your cells are proteins, genes often contain the instructions for building the polypeptide chains that make up proteins (for more on protein structure, head to Chapter 3). So, one gene often equals one polypeptide chain.
Sometimes it’s hard to imagine how one little protein can affect you in an important way. After all, humans have about 25,000 genes, so you make a lot of different proteins. How can just one faulty protein make a difference? Well, if your skin cells couldn’t make the protein collagen, your skin would fall off at the slightest touch. Also, if the cells in your pancreas couldn’t make the protein insulin, you’d have diabetes. So you see, all the functions of your body that you probably take for granted — the way it’s built, the way it looks, and the way it functions — are controlled by the actions of proteins.
Moving from DNA to RNA to Protein: The Central Dogma of Molecular Biology
The instructions in DNA determine the structure and function of all living things, which makes DNA pretty darn important. Every time a cell reproduces, it must make a copy of these instructions for the new cell. When cells need to build a functional molecule (usually a protein), they copy the information in the genes into an RNA molecule instead of using the DNA blueprint directly (see Chapter 3 for more about RNA molecules). Here’s an outline of the process:
Cells use transcription to copy the information in DNA into RNA molecules.
The information to build proteins is copied into a special type of RNA called messenger RNA (mRNA), which carries the blueprint for the protein from the nucleus to the cytoplasm where it can be used to build the protein.
Cells use translation to build proteins from the information carried in mRNA molecules.
 The idea that information is stored in DNA, copied into RNA, and then used to build proteins is considered the central dogma of molecular biology.
The idea that information is stored in DNA, copied into RNA, and then used to build proteins is considered the central dogma of molecular biology.
 Transcription and translation are two pretty similar sounding words for two very different processes in cells. One way to remember which process is which is to think about the English meanings of these words. When you transcribe something, you copy it. Transcription in cells takes the information in DNA and uses it to make RNA. DNA and RNA are similar molecules, so it’s not like you’re really changing anything — you’re just copying the information down. When you translate something, on the other hand, you change it from one language to another. Translation in cells takes the information in mRNA and uses it to build a protein, which is a different type of molecule. So, translation changes the language of molecules from RNA to protein.
Transcription and translation are two pretty similar sounding words for two very different processes in cells. One way to remember which process is which is to think about the English meanings of these words. When you transcribe something, you copy it. Transcription in cells takes the information in DNA and uses it to make RNA. DNA and RNA are similar molecules, so it’s not like you’re really changing anything — you’re just copying the information down. When you translate something, on the other hand, you change it from one language to another. Translation in cells takes the information in mRNA and uses it to build a protein, which is a different type of molecule. So, translation changes the language of molecules from RNA to protein.
The sections that follow give you an in-depth look at transcription, RNA processing, and translation.
Rewriting DNA’s message: Transcription
DNA molecules are long chains made from four different building blocks called nucleotides that biologists represent with the letters A, T, C, and G (flip to Chapter 3 for the scoop on DNA structure). These chemical units are joined together in different combinations that form the instructions for cells’ functional molecules, which are mostly proteins.
When your cells need to build a particular protein, the enzyme RNA polymerase locates the gene for that protein and makes an RNA copy of it. (RNA polymerase is shown in Step 2 of Figure 8-1.) Because RNA and DNA are similar molecules, they can attach to each other just like the two strands of the DNA double helix do. RNA polymerase slides along the gene, matching RNA nucleotides to the DNA nucleotides in the gene.
 The base-pairing rules for matching RNA and DNA nucleotides are almost the same as those for matching DNA with DNA (see Chapter 3). The exception is that RNA contains nucleotides with uracil (U) rather than thymine (T). During transcription, RNA polymerase pairs C with G, G with C, A with T, and U with A. (Figure 8-1 gives you the visual of this. Note that the new RNA strand CUAG pairs up with the DNA sequence GATC in the gene.)
The base-pairing rules for matching RNA and DNA nucleotides are almost the same as those for matching DNA with DNA (see Chapter 3). The exception is that RNA contains nucleotides with uracil (U) rather than thymine (T). During transcription, RNA polymerase pairs C with G, G with C, A with T, and U with A. (Figure 8-1 gives you the visual of this. Note that the new RNA strand CUAG pairs up with the DNA sequence GATC in the gene.)
Figure 8-1:Transcribing DNA and processing mRNA within the nucleus of a eukaryotic cell.
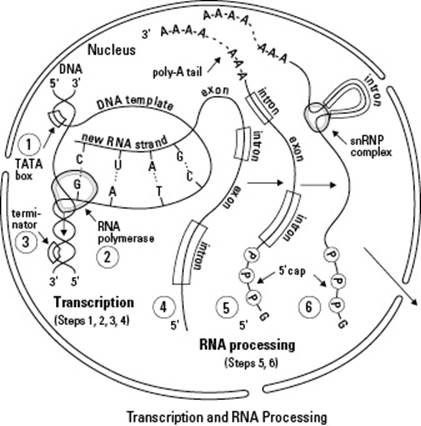
It may seem strange that your cells copy the information in your genes into a sort of mirror image made of RNA, but it actually makes a lot of sense. Your genes are extremely important and need to be protected, so they’re kept safe in the nucleus at all times. Your cells make copies of any information they need so the original DNA doesn’t get damaged.
 You can think of your chromosomes like drawers in a file cabinet. When your cells need information from the files, they open a drawer, take out a file (the gene), and make a copy of the information (the RNA molecule) that can travel out into the world (the cytoplasm). The original blueprint (the DNA) is kept safely locked away in the file cabinet.
You can think of your chromosomes like drawers in a file cabinet. When your cells need information from the files, they open a drawer, take out a file (the gene), and make a copy of the information (the RNA molecule) that can travel out into the world (the cytoplasm). The original blueprint (the DNA) is kept safely locked away in the file cabinet.
Of course, RNA polymerase and DNA aren’t the only things involved in transcription. The following sections introduce you to the other players and walk you through the process of transcription step by step.
Finding out what else is involved
RNA polymerase locates the genes it needs to copy with the help of proteins called transcription factors. These proteins look for certain sequences in the DNA that mark the beginnings of genes; these sequences are called promoters.
Transcription factors find the genes for the proteins the cell needs to make and bind to the promoters so RNA polymerase can attach and copy the gene. Many promoters contain a particular sequence called the TATA box because it contains alternating T and A nucleotides. Transcription factors bind to the TATA box first, followed by RNA polymerase. (Step 1 in Figure 8-1 depicts a gene’s promoter, along with its TATA box.)
The ends of genes are marked by a special sequence called the transcription terminator. Transcription terminators can work in different ways, but they all stop transcription. (Step 3 of Figure 8-1 shows a transcription terminator.)
Walking through the process
 As you can see from the following, the process of transcription is pretty straightforward:
As you can see from the following, the process of transcription is pretty straightforward:
1. RNA polymerase binds to the promoter with the help of transcription factors.
By binding to the promoter, RNA polymerase gets set up on the DNA so it’s pointed in the right direction to copy the gene.
2. RNA polymerase separates the two strands of the DNA double helix in a small area.
By opening the DNA, RNA polymerase can use one of the DNA strands as its pattern for building the new RNA molecule. (The strand of DNA that’s being used as a pattern in Figure 8-1 is labeled as the DNA template. The new RNA strand is shown as it’s being built against the template strand.)
As RNA polymerase slides along the DNA, it opens a new area, and the DNA behind it closes back up.
3. RNA polymerase uses base-pairing rules to build an RNA strand that’s complementary to the DNA in the template strand.
Because the base-pairing rules are specific, the new RNA molecule contains a mirror image of the code in the DNA.
4. RNA polymerase reaches the termination sequence and releases the DNA.
Some terminators have a sequence that causes the new RNA to fold up on itself at the end, making a little bump that causes RNA polymerase to get knocked off of the DNA.
 Your cells use transcription to make several types of RNA molecules. Some of these RNA molecules are worker molecules for the cell, others are part of cellular structures, and one type — mRNA — carries the code for proteins to the cytoplasm.
Your cells use transcription to make several types of RNA molecules. Some of these RNA molecules are worker molecules for the cell, others are part of cellular structures, and one type — mRNA — carries the code for proteins to the cytoplasm.
Putting on the finishing touches: RNA processing
After RNA polymerase transcribes one of your genes and produces a molecule of mRNA, the mRNA isn’t quite ready to be translated into a protein. In fact, when the mRNA is hot off the presses, it’s called a pre-mRNA or primary transcript because it’s not quite finished.
 Before the pre-mRNA can be translated, it has to undergo a few finishing touches via RNA processing (see Steps 5 and 6 in Figure 8-1):
Before the pre-mRNA can be translated, it has to undergo a few finishing touches via RNA processing (see Steps 5 and 6 in Figure 8-1):
The 5' cap, a protective cap, is added to the beginning of the mRNA. The 5' cap (shown in Step 5 of Figure 8-1) tells the cell it should translate this piece of RNA.
The poly-A tail, an extra bit of sequence, is added to the end of the mRNA. Like its name suggests, the poly-A tail (see Step 5 of Figure 8-1) is a chain of repeating nucleotides that contain adenine (A). It protects the finished mRNA from being broken down by the cell.
The pre-mRNA is spliced to remove introns (noncoding sequences). One kind of weird thing about your genes (and ours too) is that the code for proteins is interrupted by sequences called introns. Your cells remove the introns before shipping the mRNA out to the cytoplasm (see Step 6 in Figure 8-1). The sections of the pre-mRNA that wind up getting translated are called exons. When your cells cut the introns out of pre-mRNA, the exons all come together to form the blueprint for the protein.
 If you get confused about what introns and exons do, just remember that introns interrupt and exons get to exit the nucleus.
If you get confused about what introns and exons do, just remember that introns interrupt and exons get to exit the nucleus.
Converting the code to the right language: Translation
After a mature mRNA leaves the nucleus of a cell, it heads for a ribosome in the cell’s cytoplasm where the code it contains can be translated to produce a protein (for more on ribosomes, see Chapter 4). As the strand of mRNA slides through the ribosome, the code is read three nucleotides at a time.
 A group of three nucleotides in mRNA is called a codon. If you take the four kinds of nucleotides in RNA — A, G, C, and U — and make all the possible three-letter combinations you can, you’d come up with 64 possible codons. Each codon specifies one amino acid in the polypeptide chain of a protein.
A group of three nucleotides in mRNA is called a codon. If you take the four kinds of nucleotides in RNA — A, G, C, and U — and make all the possible three-letter combinations you can, you’d come up with 64 possible codons. Each codon specifies one amino acid in the polypeptide chain of a protein.
To figure out the amino acid a singular codon represents, follow the labels on the edges of the table in Figure 8-2. So to find out what the codon CGU represents:
1. Look first to the left side of the table and find the row marked by the first letter in the codon.
The letter C is the second letter down, so the amino acid represented by the C portion of the codon CGU is found in the second row of the table.
2. Then look to the top of the table and find the column marked by the second letter in the codon.
The letter G is the last letter in the row, so the amino acid represented by the G part of the codon CGU is found at the intersection of the second row (which is marked by C) and the last column under the Second Letter heading.
3. Look to the right side of the table and find the row marked by the third letter in the codon.
The letter U is listed first, so the amino acid represented by the U portion of the codon CGU is the first one listed in the intersection of the second row and the last column under the Second Letter heading. Put it all together and you find that the amino acid represented by the codon CGU is arginine.
 To translate a molecule of mRNA, begin at the start codon closest to the 5’ cap of the mRNA, divide the message up into codons, and look the codons up in a table of the genetic code that shows the names of the 20 different amino acids found in the proteins of living things. For example, 5’CCGCAUGCGAAAAUGA3’ translates into methionine-arginine-lysine.
To translate a molecule of mRNA, begin at the start codon closest to the 5’ cap of the mRNA, divide the message up into codons, and look the codons up in a table of the genetic code that shows the names of the 20 different amino acids found in the proteins of living things. For example, 5’CCGCAUGCGAAAAUGA3’ translates into methionine-arginine-lysine.
Figure 8-2: The genetic code.
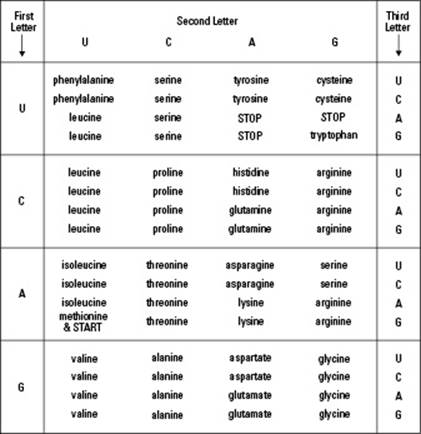
The following sections get you acquainted with specialized codons and the anticodons all codons need to pair up with for translation to occur. They also help you understand the overall process of translation.
Making sense of codons and anticodons
The genetic code is amazingly similar for all the organisms on Earth, from you to E. coli. To read it, you need to know about the unique features of some of the codons:
The codon AUG is called the start codon. AUG is called the start codon because translation begins here. When one of your cells starts translating mRNA into a polypeptide, the first AUG closest to the 5' cap of the mRNA is the first codon to be read. AUG also represents the amino acid methionine, so methionine is the first amino acid added to the polypeptide chain.
The codons UAA, UAG, and UGA are called stop codons. Translation ends when a stop codon is read in the mRNA. A stop codon only indicates when translation should end; it doesn’t represent an amino acid. When stop codons are read in the mRNA, translation stops without adding any new amino acids to the polypeptide chain.
Some amino acids are represented by more than one codon. For instance, arginine is represented by more than one codon; CGU, CGC, CGA, and CGG all represent arginine. Because of this situation, biologists say that the genetic code is redundant (more than one codon represents some amino acids).
In order for your cells to decode mRNA, they need the help of an important worker: transfer RNA (tRNA). Transfer RNA decodes the message in the mRNA. Like all RNA molecules, tRNA is made of nucleotides that can pair up with other nucleotides according to base-pairing rules.
During translation, tRNA molecules pair up with the codons in mRNA to figure out which amino acid should be added to the chain. Each tRNA has a special group of three nucleotides, called an anticodon, that pairs up with the codons in mRNA. Each tRNA also carries a specific amino acid. So, the tRNA that has the right anticodon to pair with a specific codon adds its amino acid to the growing polypeptide chain.
 Because the pairing of anticodon to codon is specific, only one tRNA can pair up with each codon. The specific relationship between tRNA anticodons and mRNA codons ensures that each codon always specifies a particular amino acid.
Because the pairing of anticodon to codon is specific, only one tRNA can pair up with each codon. The specific relationship between tRNA anticodons and mRNA codons ensures that each codon always specifies a particular amino acid.
Breaking down the translation process
Although the process of translation is fairly complicated, it’s pretty easy to understand when you break it down into three main steps: the beginning (initiation), the middle (elongation), and the end (termination). Follow along in Figure 8-3 as we present these three steps:
1. During initiation, the ribosome and the first tRNA attach to the mRNA (see #1 in Figure 8-3).
The small subunit of the ribosome binds to the mRNA. Then the first tRNA, which carries the amino acid methionine, attaches to the start codon. The start codon is AUG, so the first tRNA has the anticodon UAC (see #2 in Figure 8-3). After the first tRNA is bound to the mRNA, the large subunit of the ribosome attaches to form a complete ribosome.
2. During elongation, tRNAs enter the ribosome and donate their amino acids to the growing polypeptide chain.
Each tRNA enters a pocket in the ribosome called the A site (see #3 in Figure 8-3). An adjacent pocket, called the P site, holds a tRNA with the growing polypeptide chain (see #4 in Figure 8-3). When a tRNA is parked in the A site and the P site, the ribosome catalyzes the formation of a peptide bond between the growing polypeptide chain and the new amino acid. In Figure 8-3, a bond is forming between the amino acids cysteine (cys) and proline (pro) because they’re next to each other in the ribosome.
After the new amino acid is added to the growing chain, the ribosome slides down the mRNA, moving a new codon into the A site. After a new codon is in the A site, another tRNA can enter the ribosome, and the process of elongation can continue.
3. During termination, a stop codon in the A site causes translation to end.
The ribosome slides down the mRNA until a stop codon enters the A site. When a stop codon is in the A site, an enzyme called a release factor enters the ribosome and cuts the polypeptide chain free. Translation stops, and the ribosome and mRNA separate from each other.
Following translation, polypeptide chains may be modified a bit before they fold up and become functional proteins. Often, more than one polypeptide chain combines with another chain to form the complete protein.
Growing wild: Cancer and aging
Cancer, a disease caused by uncontrolled cell division, occurs much more frequently as people age, and the reality is that the longer you live, the more likely you are to develop cancer. Why? Because throughout your life, tiny genetic changes may occur that you don’t even realize are happening — a tiny mutation caused by an X-ray here or another tiny mutation caused by breathing in polluted air there. Some mutations are repaired; others just exist in your body’s cells without much notice. Each individual mutation may be harmless, but after genetic changes accumulate over time, the chance that some mutation will occur to change the proteins that control cell division grows greater.
If your proteins don’t do their jobs correctly, your cells may be thrown into an uncontrolled cycle of dividing and growing, causing a mass of cells called a tumor to form. As even greater numbers of mutations accumulate in the cells over time, their characteristics change, and the tumor may become malignant (more likely to spread and cause death). The spread of cancer cells around the body occurs through metastasis, when malignant cells leave their tissue of origin and enter the circulatory or lymphatic systems.
People living in the United States have a 1 in 3 chance of developing cancer during their lifetimes. That statistic is alarming, but you can do your part to reduce your risk of cancer by making good lifestyle choices — avoiding saturated fat, cigarettes, and obesity will all lower your cancer risk.
Figure 8-3:Translating mRNA into protein.
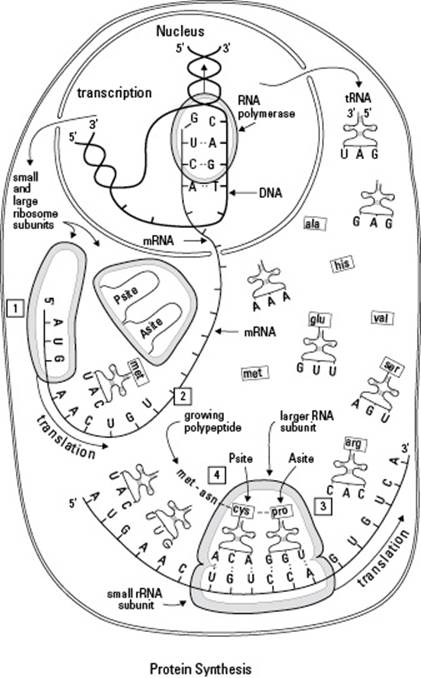
Mistakes Happen: The Consequences of Mutation
If a mistake in a strand of DNA goes undetected or unrepaired, the mistake becomes a mutation. A mutation is a change from the original DNA strand — in other words, the nucleotides aren’t in the order that they should be.
 Mutations in DNA lead to changes in RNA, which can lead to changes in proteins. When proteins change, the functions of cells, and the traits of organisms, can also change.
Mutations in DNA lead to changes in RNA, which can lead to changes in proteins. When proteins change, the functions of cells, and the traits of organisms, can also change.
Mutations usually happen as the DNA is being copied during DNA replication (see Chapter 6 for a description of DNA replication). Two main types of mutations occur:
Spontaneous mutations: These result from uncorrected mistakes by DNA polymerase, the enzyme that copies DNA. DNA polymerase is a very accurate enzyme, but it’s not perfect. In general, DNA polymerase makes one mistake for every billion base pairs of DNA it copies. One in a billion isn’t bad . . . unless you’re talking about your DNA. Then, any change can eventually cause problems. Cancer, for example, usually occurs as people age because they’ve lived long enough to accumulate mutations in certain genes that control cell division.
Induced mutations: These result from environmental agents that increase the error rate of DNA polymerase. Anything that increases the error rate of DNA polymerase is a mutagen. The most common mutagens are certain chemicals (such as formaldehyde and compounds in cigarette smoke) and radiation (like ultraviolet light and X-rays).
When mutations occur during DNA replication, some daughter cells formed by mitosis or meiosis inherit the genetic change (we explain how cells divide in Chapter 6). The types of mutations these cells inherit can be divided into three major categories:
Base substitutions: When the wrong nucleotides are paired together in the parent DNA, a base substitution occurs. If the parent DNA molecule has a nucleotide containing thymine (T), then DNA polymerase should bring in a nucleotide containing adenine (A) for the new strand of DNA. However, if DNA polymerase makes a mistake and brings in a nucleotide with guanine (G) by mistake, that’s a base substitution. Because just one nucleotide was changed, the mutation is called a point mutation. The effect of point mutations ranges from nothing to severe.
• Silent mutations have no effect on the protein or organism. Because the genetic code is redundant, changes in DNA may lead to changes in mRNA that don’t cause changes in the protein (see the earlier section “Making sense of codons and anticodons” for more on the redundancy of the genetic code).
• Missense mutations change the amino acids in the protein. Changes in DNA can change the codons in mRNA, leading to the addition of different amino acids into a polypeptide chain. The severity of missense mutations depends on how different the original amino acid is from the new amino acid and where in the protein the change occurs.
• Nonsense mutations introduce a stop codon into the mRNA, preventing the protein from being made. If the DNA changes so that a codon in the mRNA becomes a stop codon, then the polypeptide chain gets cut short. Nonsense mutations usually have severe effects and are the cause of many genetic diseases, including certain forms of cystic fibrosis, Duchenne’s muscular dystrophy, and thalassemia (an inherited form of anemia).
Deletions: When DNA polymerase fails to copy all the DNA in the parent strand, that’s a deletion. If nucleotides in the parent DNA are read but the complementary bases aren’t inserted, the new strand of DNA is missing nucleotides. If one or two nucleotides are deleted, then the codons in the mRNA will be skewed from their proper threes, and the resulting polypeptide chain will be very altered. Mutations that change the way in which the codons are read are called frameshift mutations. Deletions of three nucleotides result in the deletion of one amino acid. Serious diseases such as cystic fibrosis and Duchenne muscular dystrophy result from deletions.
Insertions: When DNA polymerase slips and copies nucleotides in the parent DNA more than once, an insertion occurs. Just like deletions, insertions of one or two nucleotides can cause frameshift mutations that greatly alter the polypeptide chain. Huntington’s disease, an illness that causes the nervous system to degenerate starting when a person is in his 30s or 40s, is caused by insertions of the sequence CAG into a normal gene up to 100 times. Although the sequence is a multiple of three (so technically not a frameshift mutation), the abundance of these insertions messes up the reading of the normal genetic code, causing either abnormal protein production or a lack of protein production.
Fighting for breath
Cystic fibrosis (CF), a disease that clogs the lungs and smaller passageways of the body with mucus, is the most common genetic disease found among people of northern European ancestry (about 30,000 people are living with CF in the United States alone). The mucus in the lungs makes it difficult to breathe and provides a breeding ground for bacteria, causing repeated lung infections. People with CF don’t usually live past young adulthood because their lungs ultimately fail from all the damaging infections. CF patients and their families struggle against this disease daily, using therapies to break up the mucus and fight the infections, trying to hold back the effects of this dreadful disease.
All of this difficulty and pain is caused by a mutation in one protein: the CFTR protein. Normal CFTR protein is found in the plasma membranes of people’s cells, where it helps move chloride across the membrane. The movement of chloride affects the movement of water, which is why the mucus gets so thick on the outside of cells in people who have the disease. The most common mutation in CFTR is a deletion of three nucleotides, resulting in the loss of one amino acid. That doesn’t seem like a very big change, but the loss of that one amino acid alters the polypeptide chain so it doesn’t fold up properly. Other proteins, acting as quality-control proteins, see the abnormal folding and mark the mutated protein for destruction. So, in people with the most common form of CF, the CFTR protein never even makes it to the plasma membrane.
Scientists are currently searching for ways to correct this problem. One new and very exciting development is the discovery of a drug that helps mutant CFTR proteins make it to the plasma membrane. In lung cells grown in the laboratory, the mutant CFTR proteins were able to transport some chloride, suggesting that the drug may be able to reduce the severity of the disease. The researchers who discovered this effect are taking their discovery to the next level by working on appropriate doses of the drug and testing it on people who suffer from CF.
Giving Cells Some Control: Gene Regulation
Even though your DNA is in control of the proteins your body makes, and those proteins are in charge of determining your traits, your cells do have some say in life. Because each one of your cells has a complete set of your chromosomes, your cells are able to practice gene regulation,meaning they can choose which genes to use (or not use) and when.
 When a cell uses a gene to make a functional molecule, that gene is expressed in the cell. Gene regulation is the process cells use to choose which genes to express at any one time. (Scientists talk about gene regulation as cells turning genes “on” or “off.”)
When a cell uses a gene to make a functional molecule, that gene is expressed in the cell. Gene regulation is the process cells use to choose which genes to express at any one time. (Scientists talk about gene regulation as cells turning genes “on” or “off.”)
Genes are regulated by the action of proteins that bind to DNA and either help or block RNA polymerase from accessing the genes. In your cells, proteins that help RNA polymerase bind to your genes are called transcription factors. They bind to special sequences on the DNA near genes’ promoters and make it possible for RNA polymerase to bind the promoter. Transcription and translation occur, producing the protein in the cell.
Gene regulation allows your cells to do two things: adapt to environmental changes and make it so that each cell type has a distinct role in the body. We fill you in on both in the next sections.
Adapting to environmental changes
The world around you is always changing, which means you need to be able to respond to environmental signals in order to maintain your physiological balance. Gene regulation allows you to do just that. When your cells need to respond to environmental changes, they turn genes on or off to make the proteins needed for the response.
Suppose you’re getting too much sunlight. To protect your skin, the cells on the tip of your nose need to darken a bit by making more of the skin pigment melanin. The extra sunlight on your skin triggers certain proteins to bind to the genes needed for melanin production and help RNA polymerase access the genes. RNA polymerase reads the genes, making mRNA that contains the blueprints for the necessary proteins. The mRNA is translated, and the proteins are made. The proteins do their jobs, and the skin on your nose turns a darker color. This example of how your skin gets darker illustrates how cells can access genes when they need them in order to respond to signals from the environment.
Becoming an expert through differentiation
You have more than 200 different types of cells in your body, including skin cells, muscle cells, and kidney cells. Each of these cells does a different job for your body, and like any good craftsman, each of these cell types requires the right tools for its job. To a cell, the right tool for the job is usually a specific protein. For instance, skin cells need lots of the protein keratin, muscle cells needs lots of contractile proteins, and kidney cells need water-transport proteins.
 Cell differentiation is the process that makes cells specialized for certain tasks. Your differentiated cells have all the blueprints for all the tools because they each have a full set of your chromosomes; what makes them different from one another is which blueprint they use.
Cell differentiation is the process that makes cells specialized for certain tasks. Your differentiated cells have all the blueprints for all the tools because they each have a full set of your chromosomes; what makes them different from one another is which blueprint they use.
Cells differentiate from one another due to gene regulation. For example, when a sperm met an egg to form the cell that would become the future you, that first cell had the ability to divide and form all the different cell types your body needed. As that cell and its descendents divided, however, signals caused different groups of cells to change their gene expression. Proteins in these cells bound to the DNA molecules, activating some genes and silencing others. As you grew and developed in your mother’s uterus, your cells became more and more distinct from each other. Some cells became part of your nervous tissue, whereas others formed your digestive tract. Each of these changes occurred as cells transcribed and translated the genes for the proteins that they needed to do their particular job.
 After a cell becomes differentiated, it usually can’t go back. It’s specialized for a certain task and can’t access the genes for proteins that aren’t in its job description.
After a cell becomes differentiated, it usually can’t go back. It’s specialized for a certain task and can’t access the genes for proteins that aren’t in its job description.
Stem cells: An introduction and a moral dilemma
Stem cells are cells that have the potential to become any type of cell in the body. When they divide, their descendants can either remain stem cells or become differentiated into a specialized cell type. Stem cells are also unique because they’re immortal — unlike most human cells, which stop dividing after about 40 or 50 cell divisions, stem cells can continue to produce new cells indefinitely. They’re therefore extremely important to the body because they can help replace cells and repair damage in almost every kind of tissue. They also have the potential to cure human diseases and save lives.
For many years the best source of stem cells for research was human embryos made by in vitro fertilization but no longer wanted by the people who donated the sperm and egg. Unwanted embryos are usually destroyed, but some people choose to donate their frozen embryos to scientific research. Human embryonic stem cells are special because they’re pluripotent, meaning they have the potential to become every type of human cell. Although adult stem cells can be taken from fully developed humans, these stem cells are usually only able to become a limited number of cell types based on their tissue of origin. For example, hematopoietic stem cells from the bone marrow have the ability to divide and produce many different types of blood cells, but they typically can’t produce nerve cells.
Because human embryonic stem cells are pluripotent, they’re more valuable for some types of research. However, some people feel that destroying human embryos is morally wrong, no matter how early in development they are (embryos frozen after in vitro fertilization typically have about five to eight cells). This belief even led to a ban on federal funding for the development of new stem cell lines derived from human embryos. Those who support stem cell research argue that the potential benefits of stem cells for treating disease and saving lives make it imperative that scientists continue their research with embryonic stem cells.
Will this moral dilemma ever be solved? One potential solution is to make adult stem cells behave as if they’re embryonic stem cells. A group of scientists has figured out a way to make this happen, but additional research is needed to determine whether these induced pluripotent stem cells are truly equivalent to embryonic stem cells.