MCAT Biology Review
Chapter 12: Genetics and Evolution
12.2 Changes in the Gene Pool
All of the alleles that exist within a species are known as the gene pool. When mutations or genetic leakage occur, new genes are introduced into the gene pool. Genetic variability is essential for the survival of a species because it allows it to evolve to adapt to changing environmental stresses. Certain traits may be more desirable than others and confer a selective advantage—an advantage that allows for the individual to produce more viable, fertile offspring. In this section, we will consider genetic diversity and mutations, leakage, and genetic drift, which cause changes to the alleles present in the gene pool.
MUTATIONS
A mutation is a change in DNA sequence. New mutations may be introduced in a variety of ways. Ionizing radiation, such as ultraviolet rays from the sun, and chemical exposures can damage DNA; substances that can cause mutations are called mutagens. DNA polymerase is subject to making mistakes during DNA replication, albeit at a very low rate; proofreading mechanisms also help prevent mutations from occurring through this mechanism. Elements known as transposons can insert and remove themselves from the genome. If a transposon inserts in the middle of a coding sequence, the mutation will disrupt the gene.
Flawed proteins can arise in other ways without an underlying change in DNA sequence, as well. Incorrect pairing of nucleotides during transcription or translation, or a tRNA molecule charged with the incorrect amino acid for its anticodon, can result in derangements of the normal amino acid sequence.
The major types of nucleotide-level mutations are discussed in great detail in Chapter 7 of MCAT Biochemistry Review, so we offer just a brief overview here of each type.
Nucleotide-Level Mutations
Many mutations occur at the level of a single nucleotide (or a very small number of nucleotides). These mutations are shown in Figure 12.3 and are summarized below.
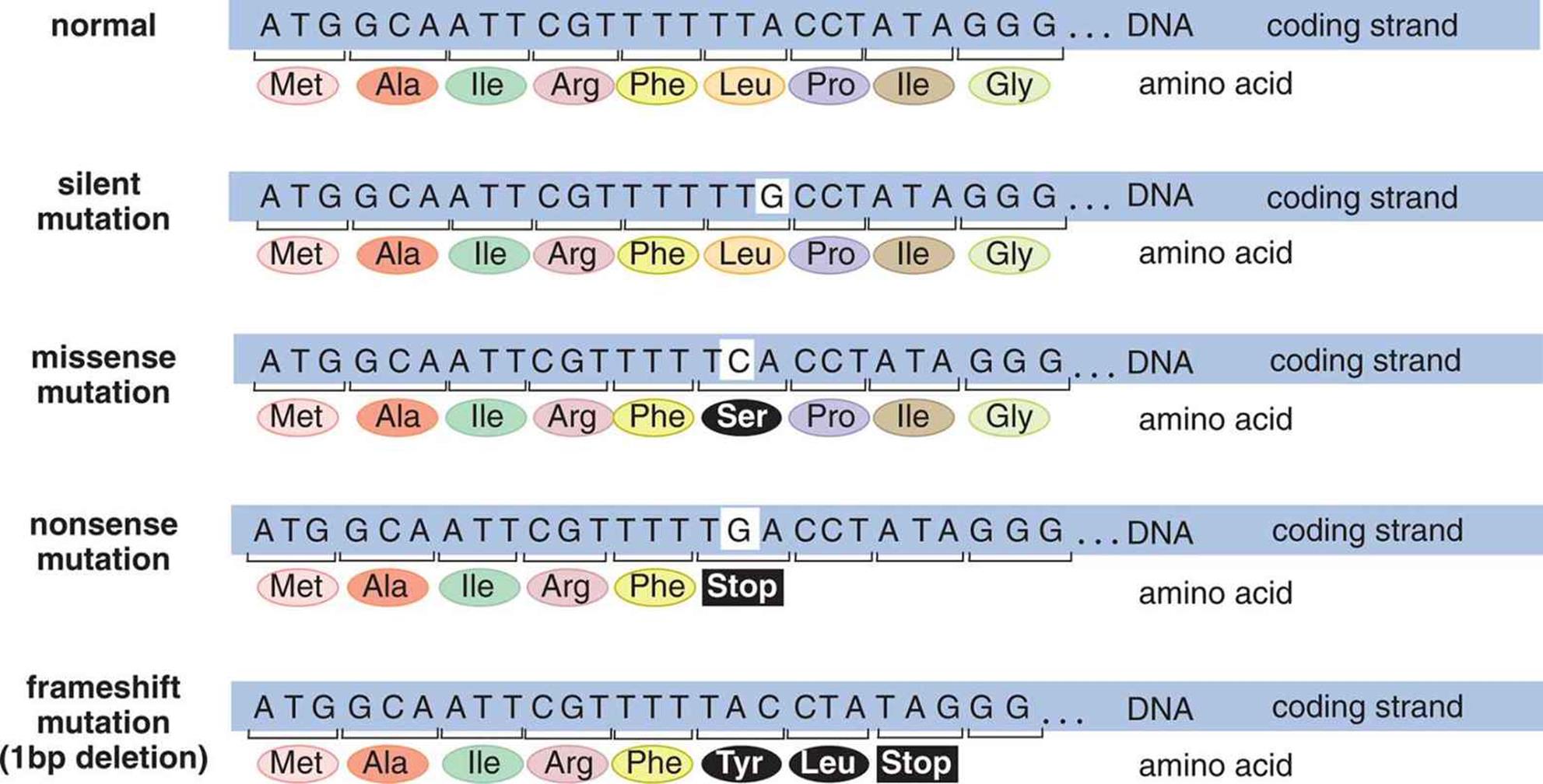 Figure 12.3. Common Nucleotide-Level Mutations
Figure 12.3. Common Nucleotide-Level Mutations
Point mutations occur when one nucleotide in DNA (A, C, T, or G) is swapped for another. These can be subcategorized as silent, missense, or nonsense mutations:
· Silent mutations occur when the change in nucleotide has no effect on the final protein synthesized from the gene. This most commonly occurs when the changed nucleotide is transcribed to be the third nucleotide in a codon because there is degeneracy (wobble) in the genetic code.
· Missense mutations occur when the change in nucleotide results in substituting one amino acid for another in the final protein.
· Nonsense mutations occur when the change in nucleotide results in substituting a stop codon for an amino acid in the final protein.
Frameshift mutations occur when nucleotides are inserted into or deleted from the genome. Because mRNA transcribed from DNA is always read in three-letter sequences called codons, insertion or deletion of nucleotides can shift the reading frame, usually resulting in either changes in the amino acid sequence or premature truncation of the protein (due to the generation of a nonsense mutation). These can be subcategorized as insertion or deletion mutations.
Chromosomal Mutations
Chromosomal mutations are larger-scale mutations in which large segments of DNA are affected, as demonstrated in Figure 12.4 and summarized below.
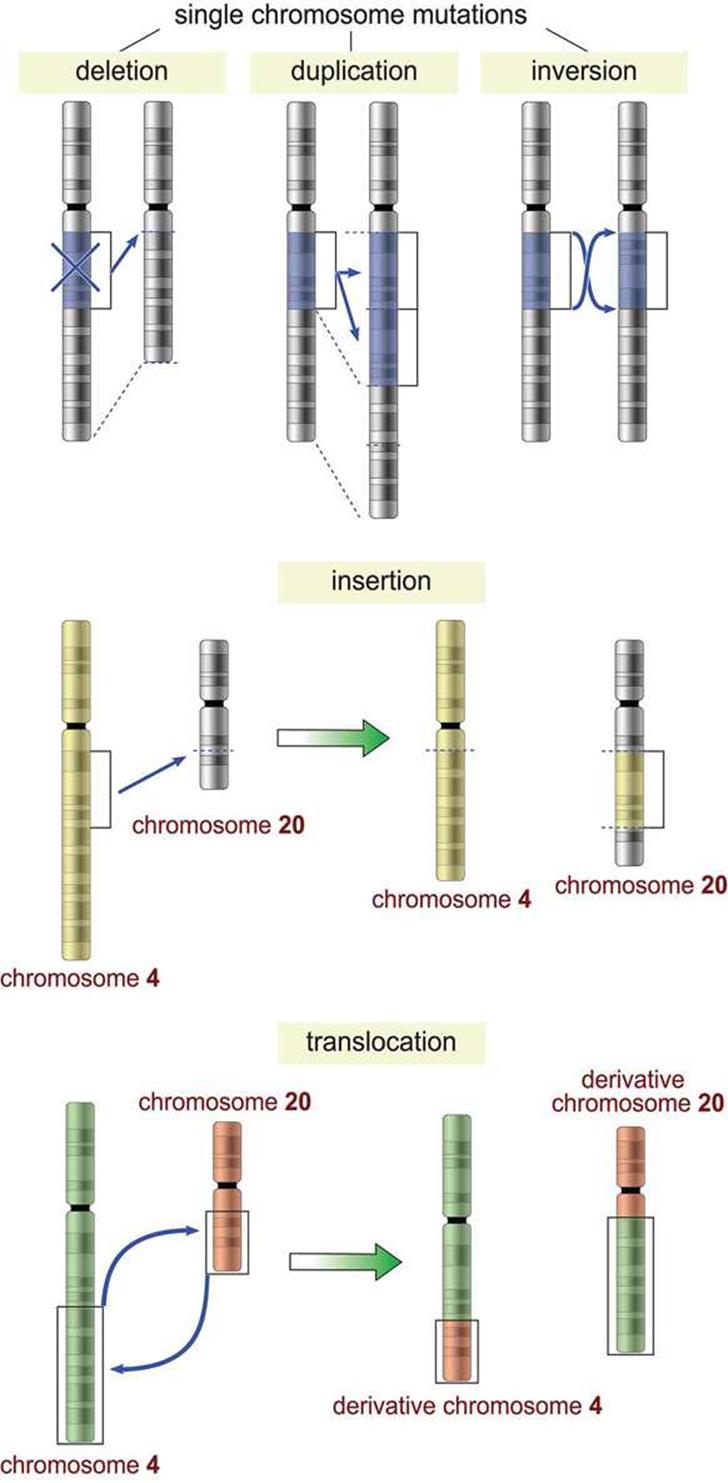 Figure 12.4. Common Chromosomal Mutations
Figure 12.4. Common Chromosomal Mutations
· Deletion mutations occur when a large segment of DNA is lost from a chromosome. Small deletion mutations are considered frameshift mutations, as described previously.
· Duplication mutations occur when a segment of DNA is copied multiple times in the genome.
· Inversion mutations occur when a segment of DNA is reversed within the chromosome.
· Insertion mutations occur when a segment of DNA is moved from one chromosome to another. Small insertion mutations (including those where the inserted DNA is not from another chromosome) are considered frameshift mutations, as described previously.
· Translocation mutations occur when a segment of DNA from one chromosome is swapped with a segment of DNA from another chromosome.
Consequences of Mutations
Mutations can have many different consequences. Some mutations can be advantageous, conferring a positive selective advantage that may allow the organism to produce more offspring. For example, sickle cell disease is a single nucleotide mutation that causes sickled hemoglobin. While the disease itself is detrimental to life, heterozygotes for sickle cell disease usually have minor symptoms, if any, and have natural resistance to malaria because their red blood cells have a slightly shorter lifespan—just short enough that the parasitic Plasmodium species that cause malaria cannot reproduce. Thus, heterozygotes for sickle cell disease actually have a selective advantage because they are less likely to die from malaria.
On the other hand, some mutations can be detrimental or deleterious. For example, xeroderma pigmentosum (XP) is an inherited defect in the nucleotide excision repair mechanism. In patients with XP, DNA that has been damaged by ultraviolet radiation cannot be repaired appropriately. Ultraviolet radiation can introduce cancer-causing mutations; without a repair mechanism, patients with XP are frequently diagnosed with malignancies, especially of the skin.
One important class of deleterious mutations is known as inborn errors of metabolism. These are defects in genes required for metabolism. Children born with these defects often require very early intervention in order to prevent permanent damage from the buildup of metabolites in various pathways. For example, in phenylketonuria (PKU), the enzyme phenylalanine hydrolase, which completes the metabolism of the amino acid phenylalanine, is defective. In the absence of this enzyme, toxic metabolites of phenylalanine accumulate, causing seizures, impairment of cerebral function, and learning disabilities, as well as a musty odor to bodily secretions. However, if the disease is discovered shortly after birth, then dietary phenylalanine can be eliminated and treatments can be administered to aid in metabolizing any additional phenylalanine.
LEAKAGE
Genetic leakage is a flow of genes between species. In some cases, individuals from different (but closely related) species can mate to produce hybrid offspring. Many hybrid offspring, such as the mule (hybrid of a male horse and a female donkey), are not able to reproduce because they have odd numbers of chromosomes—horses have 64 chromosomes and donkeys have 62, so mules, with 63 chromosomes, cannot undergo normal homologous pairing in meiosis and cannot form gametes. In some cases, however, a hybrid can reproduce with members of one species or the other, such as the beefalo (a cross between cattle and American bison). The hybrid carries genes from both parent species, so this results in a net flow of genes from one species to the other.
GENETIC DRIFT
Genetic drift refers to changes in the composition of the gene pool due to chance. Genetic drift tends to be more pronounced in small populations. The founder effect is a more extreme case of genetic drift in which a small population of a species finds itself in reproductive isolation from other populations as a result of natural barriers, catastrophic events, or other bottlenecks that drastically and suddenly reduce the size of the population available for breeding. Because the breeding group is small, inbreeding, or mating between two genetically related individuals, may occur in later generations. Inbreeding encourages homozygosity, which increases the prevalence of both homozygous dominant and recessive genotypes. Ultimately, genetic drift, the founder effect, and inbreeding cause a reduction in genetic diversity, which is often the reason why a small population may have increased prevalence of certain traits and diseases. For example, branched-chain ketoacid dehydrogenase deficiency (also called maple syrup urine disease) is especially common in Mennonite communities; this implies a common origin of the mutation, which may be a very small original population.
This loss of genetic variation may cause reduced fitness of the population, a condition known as inbreeding depression. On the opposite end of the spectrum, outbreeding or outcrossing, is the introduction of unrelated individuals into a breeding group. Theoretically, this could result in increased variation within a gene pool and increased fitness of the population.
MCAT Concept Check 12.2:
Before you move on, assess your understanding of the material with these questions.
1. What are the three main types of point mutations? What change occurs in each?
·
·
·
2. What are the two main types of frameshift mutations?
·
·
3. What are the three main types of chromosomal mutations that do NOT share their name with a type of frameshift mutation? What change occurs in each?
·
·
·
4. Why would genetic leakage in animals be rare prior to the last century?
5. Why is genetic drift more common in small populations? What relationship does this have to the founder effect?