Microreactors in Organic Chemistry and Catalysis, Second Edition (2013)
10. Bioorganic and Biocatalytic Reactions
10.3. Biocatalysis by Enzymatic Microreactors
10.3.1 Classification of Enzymatic Microreactors Based on Application
Enzymatic microreactors offer various possibilities for performing chemical and biochemical assays such as enzyme assays and immunoassays, especially in the analysis of proteins and nucleic acids, and other biomolecules. In addition to biochemical analysis, they also have their applications in biocatalysis. Many of these microdevices were developed not only as model systems but also as answers to specific problems. Several developments have focused on the usage of microreactors in enzymatic diagnostics. These systems are characterized by chemical reactions faster than the traditional assays with minute amounts of sample and reagents, repeatability and reproducibility, which is attributed to replacing iteration of procedures in batch reactions, and to discrete sample treatment by flow injection systems. Such features make microreactors ideal for integration of several processes into automated and high-throughput analytical systems for precise environmental control, hence eliminating the possibility of human errors.
In the following discussion, applications of enzymatic microreactors will be presented. Comprehensive and up-to-date references are included that will give readers access to detailed information on the arising trends in the development of enzymatic microreactors for bioorganic and biocatalytic reactions and their current applications for applied analytical chemistry and biochemical studies.
10.3.1.1 Applications of Microreactors for Enzymatic Diagnostics and Genetic Analysis
Recent developments in microfluidic technology have focused on microreactors application in enzymatic diagnostics. These systems enable chemical reactions at scales many-fold higher than the conventional assays with minimum requirement for sample and reagent volumes. These advantages make microreactors ideally suited for high-throughput portable devices for precise control of microenvironment. These devices miniaturize and integrate several processes in a single chip, with the added advantage of automation. The most notable application of microtechnology has been setting up systems with microreactors for polymerase chain reaction, enabling automation of DNA amplification. Nucleic acid analysis chips have been developed that rely on technique such as PCR. Microfluidic technologies have focused on the usage of microreactors in enzymatic diagnostics, which have allowed construction of high-throughput systems for fast analysis of genetic material. The detailed discussion of development of PCR chip, however, is beyond the scope of this chapter and we refer readers to some excellent reviews [37–43]. We focus on the recently developed microreactors and their applications on diagnostics and genetic material analysis.
DNA analysis on-chip can be used in disease diagnosis with a genetic basis. PCR-based amplification and analysis can provide information on patient's gene expression profiles. The standard method of PCR takes several days but there is an immense need for miniaturized, low cost, disposable, robust, and portable diagnostic devices capable of rapid detection and accurate identification of a wide range of pathogens. The emerging field of microfluidics, in combination with microelectromechanical systems (MEMS) technology, to create LOC or μTAS, promises exciting solutions to realize PCR-based diagnostics. Microfluidics involves the precise control and manipulation of fluids in the microscale environment. Biological systems based on microfluidics allow miniaturization and integration of processes that were normally done at larger scale in separate operations with enhanced speed and efficiency [44]. The strengths of microfluidics can be widely exploited in miniaturization of PCR. Rapid thermal cycling can be achieved through heat transfer in microfluidic-based PCR due to the small reaction mass and the high surface to volume ratio of the microreactor. PCR requires temperature cycles during which a DNA strand is first melted at a higher temperature followed by annealing at a low temperature. The small thermal mass due to microscale enables significant reduction in cycle times from 1 to 2 h to only 5–10 min, which can be a major breakthrough in diagnostics given the long time needed for the conventional tests. The small length scale can also lead to a more uniform temperature distribution and enhance the yield and integrity of PCR. Microfluidic approach also features the advantages such as shorter assay time, lower reagent consumption, rapid heating/cooling, portability, as well as the possibility of integrating multiple pre- and post-PCR processing modules in a self-contained system with full automation. Furthermore, the closed system of microreactors decreases the risk of infection in the diagnostics of patients' samples. Typically, the substrates for PCR reaction and polymerase enzyme are injected into the reaction zone and a thermal cycling program is applied to enable amplification of the DNA. On-chip PCR is typically coupled with reverse transcription and capillary electrophoresis. Recent progress has been made on applying these techniques for specific diseases. For example, reverse transcription, PCR, and gel electrophoresis have been integrated in a microfluidic chip to detect influenza virus called the influenza virus genotyper [45]. The device converts RNA from throat swabs into billions of copies of complementary DNA and integrated the system with gel electrophoresis for DNA detection. The most recent on-chip PCR for detection of influenza virus was reported by Cao et al. [50]. In this study, the authors developed a single-use microfluidic chip that integrates solid phase extraction (SPE) and molecular amplification via a reverse transcription polymerase chain reaction (RT-PCR) to amplify influenza type A RNA and the PCR products are read using a commercial capillary electrophoresis chip. The recent trend is to integrate pre-PCR sample preparation with post-PCR analysis into a streamlined and automated system for point-of-care applications. Integration of pre-PCR sample preparation includes platforms that utilize filter separation [46–50], embedded SPE [51–53], and immunomagnetic beads [51–53]. We discuss below, some examples of the studies that represent the key advancements in PCR gene expression measurements in a single chip purposely developed for clinical diagnostics.
A fully integrated LOC was reported by Sauer-Budge and others [54] for detecting bacteria. The system enables bacterial lysis, nucleic acid isolation and concentration, PCR, and end-point fluorescent detection. They utilized a novel porous polymer monolith (PPM) embedded with silica that has been shown to lyse bacteria and isolate the nucleic acids [55]. The chip made of Zeonex, a thermoplastic with high melting temperature allowed PCR, good UV transmissibility for UV curing of the PPM, and low autofluorescence detection of the amplicon. Fluid control without the use of valves or pumps on the chip was achieved by utilizing a remote switching technique. Fluid flow rate changes on the valveless chip were made possible by incorporating speed changing fluid reservoirs. The microfluidic chip was integrated with PCR thermal cycling, which was achieved by ceramic heater and air cooling and an optical spectrometer for end-point fluorescence detection. A Taqman assay was employed to detect the isolated Bacillus subtilis DNA. This integrated system can replace the traditional clinical nucleic acid analysis, with the applicability not only in modern laboratories but also in resource-limited settings.
Current developments are not only aimed at integration but also at parallel genetic analysis as well. Parallel genetic analysis allows detection of different DNA samples simultaneously. A microfluidic device integrated with multichamber PCR and multichannel CE for parallel genetic analysis was developed (Figure 10.4) [56]. The microdevice consists of three functional units: temperature control, multiple PCR (four chambers PCR), and multiple channel separation (four separation channels, each channel connected to a PCR chamber). A separation channel is directly connected to the PCR chamber and PCR products were introduced into parallel separation channels for subsequent separation and automated detection by applying an electric field. This device was successfully used for simultaneous detection of hepatitis virus (HBV), Mycobacterium tuberculosis (MTB), and genotyping of human leukocyte antigen (HLA)-B-27. Detection of pathogens, HBV, and MTB, and genotyping of HLA-B27, expressed in patients with ankylosing spondylitis were realized on this microdevice, indicating an important step toward a completely integrated microfluidic system for parallel genetic analysis.
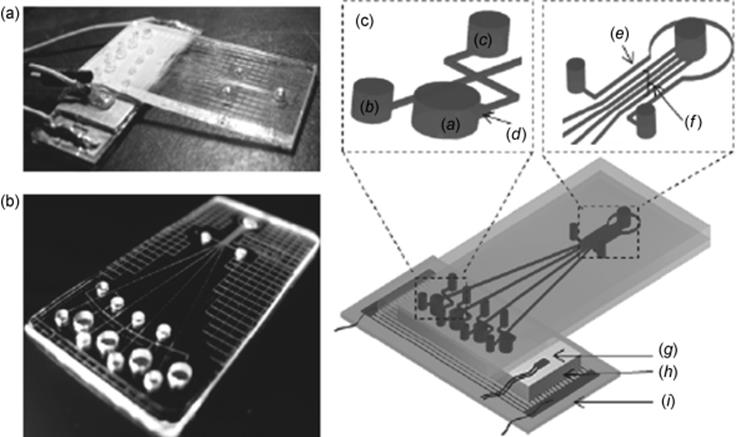
Figure 10.4 Schematics and photographs of the integrated PCR-CE microdevice. (a) Pictures of the integrated microfluidic device. (b) Pictures of glass chip (multiple PCR unit and multiple channel separation unit). (c) Schematics of the integrated microfluidic device; (a) PCR chamber, (b) cathode reservoir, (c) sample waste reservoir, (d) expanded zone, (e) reference channel, (f) separation channels, (g) platinum-chip temperature sensor, (h) Al block, and (i) glass wafer with Pt electrodes [56]. Source: Copyright 2010 Elsevier.
Further analysis of DNA also includes digestion with restrictases [57–59]. Such systems can improve automation in providing restriction maps for plasmids and can be incorporated into PCR as well as into product separation unit such as capillary electrophoresis. Xie and colleagues developed a microfluidic device for integrated restriction digestion reaction and resulting DNA fragment analysis [59]. A temperature-controlled reactor, analyte delivery, chip electrophoresis separation, and fluorescence detection was developed, in which the heaters, resistance temperature detectors, enzymatic reactors, CE channels, and pneumatic valves/pumps were integrated onto a single glass–PDMA chip. The digestion reaction was carried out using restriction endonuclease BamHI and FokI in surface passivated PDMS plate sandwiched between glass wafers.
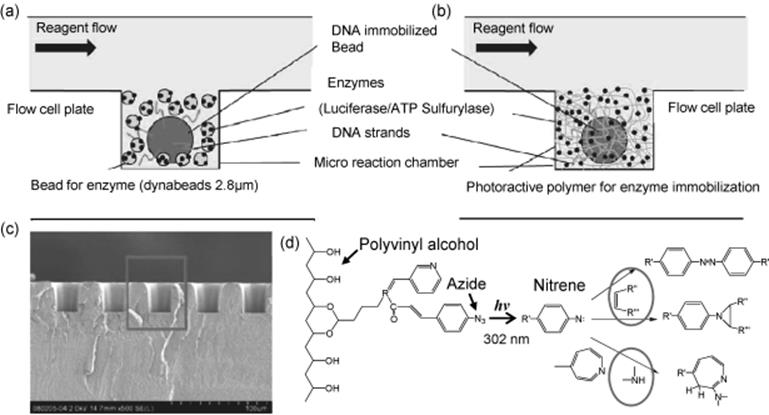
Another application of microreactors is DNA sequencing. Pyrosequencing in microreactors were found to lead to a faster sequencing with smaller quantities of DNA. Pyrosequencing is a variant sequencing method, in which individual native nucleotides are added sequentially. Pyrosequencing follows DNA polymerase progression along a DNA strand by allowing only a single dNTP to be available for incorporation at any time, and then take advantage of the chemical reaction that occurs when dNTP is incorporated by the polymerase. This reaction is followed by inducing a bioluminescence cascade and detecting the emitted light. A highly sensitive pyrosequencing system using polymer-supported enzymes illustrated in Scheme 10.5 was applied to high-throughput DNA analysis by Shirai and others [60]. A gel matrix was used to immobilize enzymes at high density in microreaction chambers (Figure 10.5). Reducing the size of microreaction chambers in a DNA analyzer is important to achieve a high throughput utilizing a commercially available detection device or camera. The use of small beads immobilizing DNA has a disadvantage in detecting luminescence because only small amounts of DNA can be immobilized on the bead surfaces for sequencing therefore, they enhanced luminescence by increasing the luciferase density in the chambers through immobilization in a gel matrix at a high concentration in the microreaction chambers. Luminescence one order of magnitude higher could be observed with the new method compared to the conventional method.
Figure 10.5 (a and b) Schematics of the intersectional view of microreaction chambers. Beads immobilizing enzymes were used as enzyme supporter in (a), and photoreactive polymer was used to immobilize enzymes in (b). (c) SEM image of the intersection of the reaction plate. The rectangle corresponds to areas in (a) and (b). (d) Reaction diagram for gelling and immobilization of enzymes [60]. Source: Copyright 2011 American Chemical Society.
Another example of DNA sequencing in microreactor by the pyrosequencing method is described by Sims and others [61], a multiplex sequencing-by-synthesis method combining terminal phosphate-labeled fluorogenic nucleotides (TPLFNs) and resealable PDMS microreactors. In the presence of phosphatase, primer extension by DNA polymerase using nonfluorescent TPLFNs generates fluorophores, which are confined in the microreactors and detected. The primed DNA templates were immobilized in the microreactors, then one of the four identically labeled TPLFNs was sequentially introduced, the microreactor sealed, and the fluorescence image after template-directed primer extension was recorded. A 30 base read with ~99% raw accuracy as demonstrated by this approach and a rapid turnaround, one-color detection, and generation of native DNA were realized.
Enzyme microreactors have also received wide application not only in the analysis but also in the synthesis of DNA. A nanoscale platform for real-time single-molecule sequencing by recording the DNA synthesis process by an immobilized DNA polymerase has been developed and has been covered by US patent application number 7,329,492 [62]. The process relies on fluorescence resonance energy transfer (FRET) between the fluorescence receptor moiety on the dNTPs and the fluorescent dye attached to the donor DNA that emits an enhanced signal upon nucleotide incorporation. Immobilization of polymerase enables increased sensitivity detection as the DNA strand extends unattached.
10.3.1.2 Application of Microreactors for Enzyme-Linked Immunoassays
Quantitative immunological methods have long been essential to many clinical, medical, pharmaceutical, and basic scientific investigations. Routine assays perform immunological assays on a microtiter plate. In laboratory settings, the enzyme-linked immunosorbent assay (ELISA) is the gold standard. ELISA, for detection of an antigen (target molecule) is called sandwich ELISA, being carried out by transmitting an antigen-containing sample onto a surface functionalized with appropriate antibodies. The targets bind to the immobilized antibodies and the rest of the samples are washed away. The bound antigens are labeled with another target-specific antibody linked to an enzyme to form the sandwich antibody–antigen–antibody. The analyte is detected by monitoring the generation of colored enzymatic products. However, the conventional ELISA presents limitations such as lengthy steps and analysis time, complex handling, requirement for a spectrophotometer or a fluorometer for detection, and high consumption of samples and reagents, and exposure to infectious agents, thereby limiting their application for point-of-care diagnosis. ELISA systems include many steps but may be simplified in miniaturized and automated devices [63]. ELISAs can be readily transferred to a microfluidic format. Expansion in the use of DNA chips had also pushed further development of immunoassay chips. Microchips open new possibilities in immunoanalysis applications. Various microfluidic devices reported today in literature for antibody (Ab) or antigen (Ag) determination were based on either homogenous or heterogenous format. In heterogenous format, wherein Abs or Ags are immobilized on a solid substrate, Ab–Ag complex forms. Heterogenous immunoassays take advantage of the high surface area/volume ratio and result in good performance and sensitivity. In the homogenous format, on the other hand, conjugation takes place in the solution phase and the immunoassay combined with microchip CE was based on the separation of the free form and the complex of Ab and Ag.
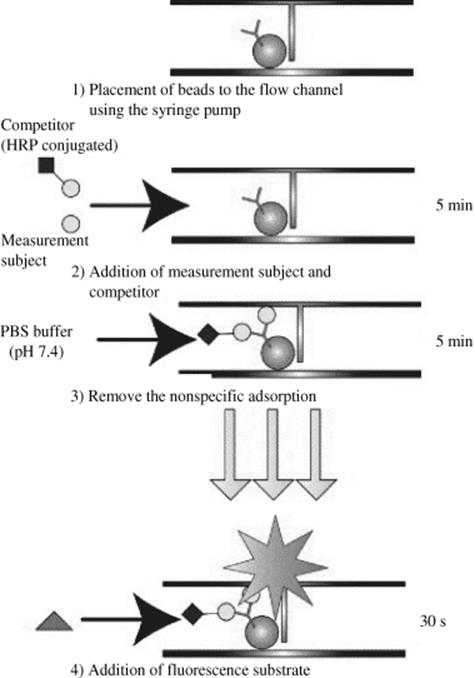
Generally, the detection mode used for a specific immunoassay technique is a function of its label, whether the antigen/antibody has an optical or enzyme label, which may also facilitate signal enhancement. The enzyme-labeled antibody manifests its signal through fluorescent product turnover from the enzyme reaction. Alkaline phosphatase (ALP) and horseradish peroxidase (HRP) are two of the most common enzyme labels. For instance, Endo and colleagues developed an immunosensor chip using the competitive immunoassay (Figure 10.6) for pollutant detection wherein polystyrene beads act as solid support for immobilization of coplanar polychlorinated biphenyls (Co-PCB) antibody [64]. The antibody-immobilized beads were introduced into the flow channel by simple control of the syringe pump. The antigen–antibody competitive reactions were performed on the surfaces of beads in the flow channel. The sample solution containing the HRP conjugated antigen, and non-HRP conjugated antigen were reacted in the microchannel yielding a linear range of 0.1 pg/ml to 1.0 μg/ml coplanar polychlorinated biphenyls. The use of enzyme-labeled antigen or antibody is by itself a signal enhancement technique.
Figure 10.6 Scheme for the competitive immunoassay on the polystyrene bead captured in the flow channel of the chip [64]. Source: Copyright 2005 Elsevier.
In another study, a microchip-based electrochemical enzyme immunoassay was developed by Chatrathi and colleagues [65] and its performance is demonstrated for the determination of monoclonal mouse IgG as a model analyte. The direct homogeneous immunoassay requires the integration of electrokinetic mixing of ALP-labeled anti-mouse IgG antibody (Ab-E) with the mouse IgG antigen (Ag) analyte in a precolumn reaction chamber, injection of immunochemical products into the separation channel, followed by rapid electrophoretic separation of enzyme-labeled free antibody and enzyme-labeled antibody–antigen complex. The separation is followed by a postcolumn reaction of enzyme tracer with p-aminophenyl phosphate (p-APP) substrate (S) and downstream amperometric detection of p-aminophenol (p-AP) product.
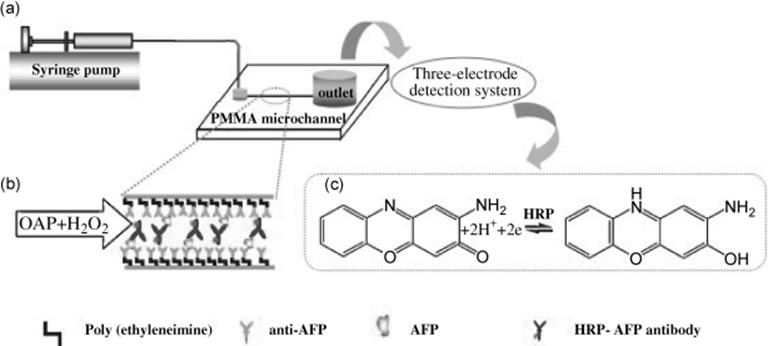
A microchip-based ELISA strategy for the detection of low-level disease biomarker in serum was reported by Liu et al. [66]. The on-chip immunoassay platform was used to determine a trace level of α-fetoprotein (AFP), hepatocellular carcinoma biomarker, using PMMA chip coupled with electrochemical detection system (Figure 10.7). The AFP monoclonal antibody was immobilized in PMMA microchannels modified with poly(ethyleneimine) (PEI) containing abundant amino groups for covalent immobilization. Then, the antigen AFP and HRP-conjugated antibody against AFP sequentially bind through antigen–antibody specific interaction. A detection limit of AFP down to 1 pg/ml using electrochemical detection system was obtained, and a detectable linear concentration range of 1–500 pg/ml by differential pulse voltammetry. The AFP existing in human serum was detected to demonstrate the utility of the immunochip.
Figure 10.7 (a) Schematic of the experimental setup including three parts: syringe pump, PMMA microfluidic chip, and three-electrode detection system, (b) schematic representation of the enzyme-linked immunosorbent assay in the PMMA microchannel, and (c) the electrochemical catalytic reaction occurred on the electrode [66]. Source: Copyright 2009 Elsevier.
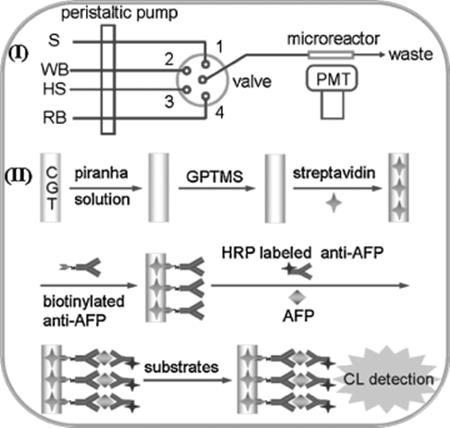
Recently, a streptavidin-functionalized capillary immune microreactor (Figure 10.8) was developed for highly efficient chemiluminescent immunoassay [67]. The functionalized capillary was used as both a support for highly efficient immobilization of antibody and a flow cell for flow-through immunoassay. For the immunoassay protocol, a mixture of AFP and HRP-labeled anti-AFP antibody was delivered into the immuno-microreactor and incubated under stop flow at room temperature for 20 min. This immunoassay system showed a linear range of three orders of magnitude from 0.5 to 200 ng/ml and a low detection limit of 0.1 ng/ml, thus making up the shortcoming of conventional capillary immunoreactors and providing a promising alternative for highly efficient flow-injection immunoassay.
Figure 10.8 Scheme of the automated flow-through CL immunoassay system: (I) flowthrough analysis system. S, incubation mixture; WB, wash buffer, HS, CL substrate for HRP; RB, regeneration buffer; (II) biofunctionalization of inside wall of glass capillary and one-step sandwich immunoassay procedure for AFP [67]. Source: Copyright 2011 Elsevier.
10.3.1.3 Applications of Microfluidic Enzymatic Microreactors in Proteomics
The field of proteomics can be divided into profiling, functional, and structural subtypes. The primary applications of microfluidic enzymatic microreactors in proteomics have been in profiling and functional proteomics while structural proteomics has not yet been studied on the microscale. Profiling proteomics is used in the identification of all proteins in a sample or in comparison of changes in protein expression among several samples. Because proteomics involves a great deal of complexity, accurate analysis demands intensive sample processing, including digestion by proteolysis, multidimensional separations, and mass spectroscopy. Another major application of enzymatic microreactors that benefit from the advantages of microfluidic microreaction technology is functional proteomics, which comprises a variety of assays to identify functional groups in proteins and determine the significance of binding events in order to elucidate the functional properties of proteins. Such advantages include favorable reaction kinetics, ability to integrate multiple processes, reduction of reagent consumption, and faster analysis time. The utility of functional proteomics is demonstrated in enzyme assays and immunoassays to characterize the behavior of a protein within a biological system.
The principal field of application of enzymatic microreactors refers to protein and peptide mapping, an essential process in the identification and sequencing of proteins. Different developments and advances in the analysis of proteins by enzymatic microreactors are highlighted in this chapter.
The identification of unknown proteins requires digestion into a group of constitutive peptides, which are interrogated with mass spectrometry. Prior to analysis of proteins, proteolytic digestion by endoproteases, typically trypsin is required for protein-expression profiling. Trypsin is the most frequently used enzyme that catalyzes the process of protein digestion through hydrolysis of peptide bonds at basic amino acid residues. Other enzymes commonly used are chymotrypsin, endoproteinases, thermolysin, and pepsin.
10.3.1.3.1 Microreactors for the Analysis of Proteins: Protein Digestion and Protein Mapping
One of the most important domains of enzymatic microreactors is protein-expression profiling, which encompasses protein and peptide mapping. This section discusses microdevices and applications stemming from microfluidic technology in the field of proteomics. Proteomics is the large-scale study of all proteins translated from the genes coded in a genome, particularly their structures and functions. One of its significant tasks is to develop approaches for efficient and rapid identification of proteins. However, the major challenges in proteomics include the relatively large number of proteins, presence of individual proteins at widely dynamic abundance levels, enormous complexity, for example, structural similarities but functional differences (protein isoforms) that complicate absolute identification, and the continued change with time, all of which points to the need for improvement in terms of efficiency in separation, sensitivity in characterization, and high throughput analysis. These challenges are the driving force for the development of new and efficient proteomic tools. Peptide mapping is a known strategy in protein identification. An interesting niche of enzymatic microreactor application is peptide mapping, a commonly used strategy in the identification and characterization of proteins, by means of enzymatic cleavage (protein digestion) followed by peptide identification by mass spectrometry (MS). The application of enzymatic microreactors in proteomics holds the promise of faster and efficient digestions, recyclability, resistance to autolysis, ease of automation, ability to process minimal sample, minimal cost, and high digestion reproducibility.
Peptide mapping is typically performed by enzymatic cleavage of the protein and identification of peptide fragments in the resulting mixture using electrospray ionization-mass spectrometry (ESI-MS) or matrix-assisted laser desorption/ionization-mass spectrometry (MALDI-MS). Enzymatic cleavage is performed using proteolytic enzymes, for example, trypsin, chymotrypsin, endoproteinase Lys-C, endoproteinase Asp-N, endoproteinase Glu-C, endoproteinase Arg-C (clostripain), thermolysin, and pepsin [68]. Prior to mass spectrometric analysis, separation of peptide fragments in the mixture either by micro-high performance liquid chromatography or capillary electrophoresis minimizes the ionization suppression and improves the sequence coverage [69].
The following discussions present recent developments that may contribute to the development of new proteomic tools through microfluidic microreaction technology.
The conventional in-solution method of protein digestion is the traditional proteolytic digestion performed by addition of free enzyme in protein solution but this method leads to long incubation time, inefficient digestion for low-abundance proteins and diluted protein samples, in addition to the possibility of generation of interfering fragments due to autolysis of protease. These unwanted fragments may lead to the ionization suppression in MS analysis and complicate interpretation of data. Enzymes are also not stable for extended time in solution and activity gradually decreases during their storage. There is, however, a requirement for rapid sample processing, targeted cleavage, and efficient database interrogation. To fulfill such requirements, attempts have been made in the past decade by researchers working on peptide mapping and proteomics through development of immobilized microfluidic enzymatic reactors. Microfluidic enzymatic microreactors are an alternative to in-solution method employing immobilization of proteases on microchannels of chip-based reactors or surfaces of capillaries. The microreactors that enable proteolytic digestion by enzymes immobilized on solid supports are also referred to as immobilized enzyme reactors, IMERs. The great potential of IMERS for proteomic applications comprise rapid and enhance digestion, automation, repeatability, efficient protein identification, and improved proteomic coverage. IMERs can be readily coupled to separation and identification systems that enable fast, efficient, high-throughput, and automated proteome analysis. Many studies on further coupling of these microreactors with MS to perform the efficient digestion of low-level proteins and sensitive peptide identification were reported [70–76]. The enzymes immobilized in microchannels provide a narrow space, hence high enzyme-to-substrate ratio can be achieved providing better molecular-level interactions between the immobilized enzymes and the flowing substrates and thus, substantially improving digestion capacity within a shorter digestion time in comparison to the free enzymes in solutions. Moreover, immobilizing the enzyme on solid supports eliminates unwanted enzyme autodigestion and interfering fragments, and an extremely high local concentration of proteolytic enzyme provides rapid catalytic turnover. These features were achieved even for low-abundance proteins and minute proteomic samples. As mentioned earlier, such microreactors could be easily coupled with separation and detection systems, leading to high-throughput analysis of proteomes. Immobilized enzymes in microchannels could be categorized as those packed in fabricated microchips and those in capillaries. We discuss enzymatic microreactors in reference to these two categories.
10.3.1.3.2 Enzymatic Microreactors for Profiling Proteomics
Prior to identification of proteins, digestion into a group constitutive peptides is required followed by interrogation by mass spectrometry. Conventional method of protein digestion entails exposure to proteolytic enzymes such as trypsin or other lytic agents such as cyanogen bromide, a tedious process that demands a multiprocess that is time consuming. The application of microfluidic enzymatic reactors holds the promise of faster and efficient protein digestion. Several device configurations have been used for proteomic processing in microfluidic devices, including open channel devices, immobilized beads, monoliths, and use of hydrogel microstructures. Many papers have been published and an excellent list of examples of IMERs on microchip and capillary microreactors for protein proteolysis and identifications are presented in Tables 10.1 and 10.2, respectively. Open-channel microdevices are capable of achieving complete tryptic digestion in a very short timeframe (several minutes to <5 s). Another alternative method for increasing the surface-to-volume ratio in microfluidic devices is the use of trypsin modified monoliths, or from membrane sandwiched between channels. Sol–gel entrapment method is also employed to immobilize enzyme on the wall of the channel or capillary. A variety of methods are available for enzyme immobilization such as covalent linking, and physical methods such as adsorption, and entrapment.
Table 10.1 Immobilized-enzyme reactors (IMERs) in microchips and their applications in proteolysis.
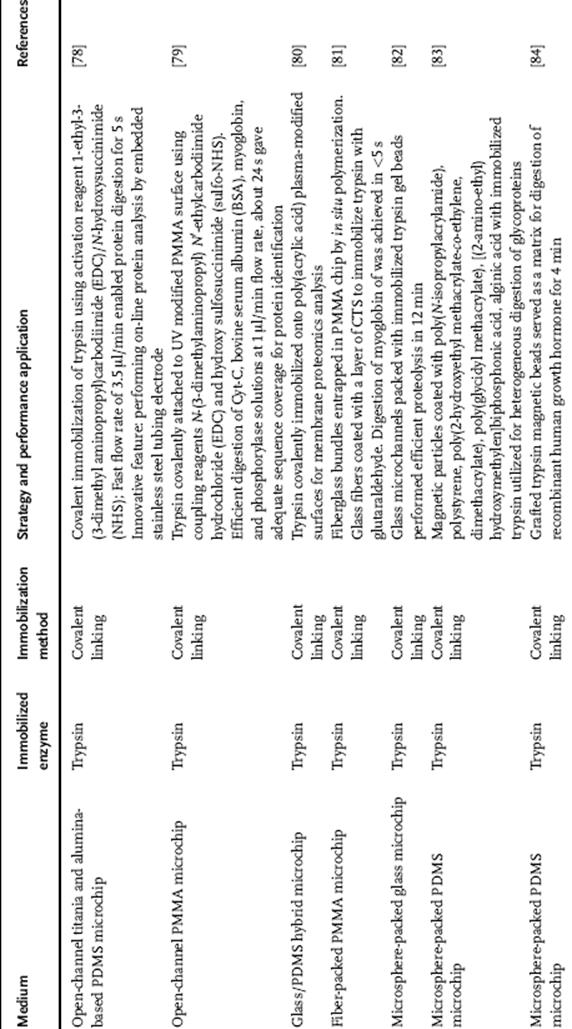



Table 10.2 Immobilized enzyme reactors (IMERs) in capillary and their applications in proteolysis.




10.3.1.3.3 Enzyme Immobilization in Microfluidic Chips for Efficient Proteolysis
Basically, all the advantages of immobilized enzymes in conventional reactors also apply to network of microchannels and capillaries [9]. Considering the large surface-to-volume ratio in microsystems, interfacial interactions play a significant role [8]. Owing to the predominant laminar flow and much stronger diffusion in these microsystems, kinetics are expected to be different from the large reactors.
There are several methods available on immobilization of enzymes. An open tubular enzyme microreactor has the protease immobilized on the channel surfaces of the microchip by adsorption (Figure 10.9), covalent linking (Figure 10.10), cross-linking immobilization (Figure 10.11), entrapment (Figure 10.12a and b), or encapsulation (Figure 10.13). A common immobilization technique is covalent linking characterized by strong binding and minimal enzyme leakage from the support. Aside from immobilization on the inner surfaces of the channel, proteases are also immobilized in monoliths in channels, or on the in-channel microspheres. Table 10.1 presents reports on IMERs in microchips employed for efficient proteolysis.
Figure 10.9 Immobilization of enzyme using the adsorption technique.

Figure 10.10 Immobilization of enzyme using the covalent technique.

Figure 10.11 Immobilization of enzyme by the cross-linking technique.

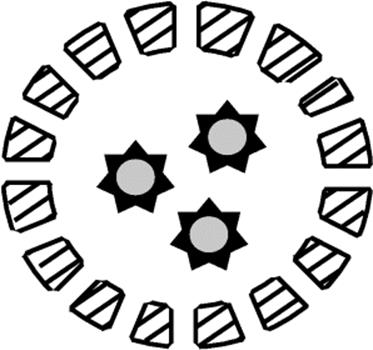
Figure 10.12 (a) Immobilization of enzyme using the entrapment technique in a matrix and (b) in a fiber.
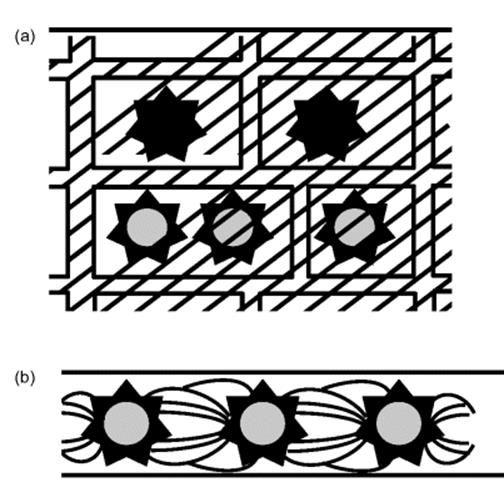
Figure 10.13 Immobilization of enzyme by the encapsulation technique.
10.3.1.3.4 Immobilization Approaches in Chip-based Microfluidic Systems
There are a number of strategies for enzyme immobilization on channels or microchip such as covalent attachment to the activated support, physical adsorption of the protein onto a solid matrix, and copolymerization of the protein with polymers [77]. These immobilization approaches applied to microfluidic microchips are discussed below.
Covalent Immobilization of Enzymes in Microchips
Covalent bonding of enzymes on a functionalized solid support is the most intensely used immobilization technique, as evident from Table 10.1. This technique offers the great advantage of preventing desorption phenomenon given the presence of substrates and solutions at high ionic strength, reducing spontaneous enzyme autodigestion. Moreover, the use of covalent bonding involves a higher enzymatic thermal stability since strong interaction of enzyme to support causes rigidity of the protein structure and consequently limits the thermal movement of the protein at high temperature. Owing to the difficulty of enzyme to unfold, inactivation is not easily observed and high enzymatic turnover is achieved with fewer diffusional restrictions. Usually immobilization is carried out in the presence of carbodiimide, cross-linking by glutaraldehyde or cyanogen bromide activation of the support material. Covalent binding to the activated support gives the advantage of minimization of the leakage of the immobilized enzyme and improving the lifetime of enzyme-immobilized microreactors. The functional groups of the proteins suitable for covalent bonding under mild conditions are (i) the α-amino group of the backbone, and the ε-amino group of lysine, and the guanidine group of arginine, (ii) the α-carboxyl group of the chain end and the β- and γ-carboxyl groups of aspartic and glutamic acids, (iii) the phenol ring of tyrosine, (iv) the thiol group of cysteine, (v) the hydroxyl groups of serine and threonine, (vi) the imidazole group of histidine, and (vii) the indole group of tryptophan. It is important that the catalytic functional groups of the enzymes are not involved in the covalent linkage to the support in order to prevent modification of the enzymatic activity or complete inactivation of the immobilized protein.
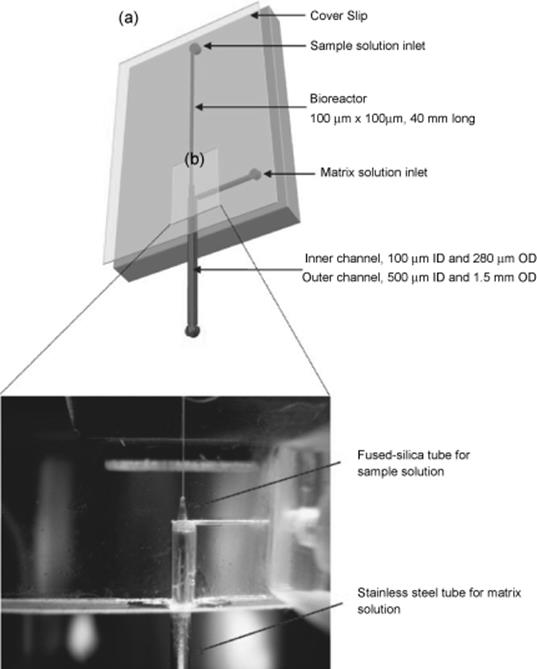
An example of an enzyme-immobilized microchip by covalent linking is based on polydimethylsiloxane developed by Wu and colleagues [78]. Polymerization of acrylic acid on the channel surface in order to introduce carboxyl groups was achieved by UV exposure. The covalent immobilization of trypsin was achieved using the activation reagents; (1-ethyl-3-(3-dimethyl aminopropyl)carbodiimide (EDC) and N-hydroxysuccinimide (NHS) were used and a fast flow rate of 3.5 μl/min enabled protein digestion for 5 s and the digests were identified by matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) and electrospray ionization (ESI)-MS. The innovative feature of this report is performing on-line protein analysis by embedded stainless steel tubing electrodes. Another study on immobilization by covalent linking of enzyme on surface of microchannel in a microreactor was reported by Lee and others [79]. An automated proteolytic digestion bioreactor and droplet deposition system was constructed with a plastic microfluidic device for off-line interfacing to MALDI-TOF-MS (Figure 10.14). Trypsin was covalently attached to UV exposed PMMA surfaces. PMMA chip surfaces can be derivatized to yield carboxylic acid termini by exposure to a 254 nm UV lamp at 15 mW/cm2 for 20 min. The stainless steel and silica tubes were then inserted into guide channels on the PMMA substrate and the UV exposed PMMA cover slip was thermally annealed to the PMMA substrate at 98 °C for 20 min. The UV modified channels were chemically treated with coupling reagents such as N-(3-dimethylaminopropyl)-EDC and water-soluble sulfo-NHS. Cytochrome C (Cyt-c), bovine serum albumin (BSA), myoglobin (MYO), and phosphorylase solutions that were efficiently digested at a flow rate of 1 μl/min for 24 s gave adequate sequence coverage for protein identification. A study by Pereira-Medrano and others [80] demonstrated a glass–PDMA hybrid microIMER with enzymes covalently immobilized onto poly(acrylic) plasma-modified surfaces for the purpose of rapidly (as low as 30 s) generating peptides suitable for MS analysis. The microdevice also allows, for the first time, rapid digestion of insoluble proteins. Membrane protein identification through this method was achieved after only 4 min digestion time, up to ninefold faster than either dual-stage in-solution digestion approaches or other commonly used bacterial membrane proteomic workflows.
Figure 10.14 Schematic of the PMMA microfluidic chip. (a) The bioreactor measured 40 mm × 100 μm × 100 μm and the solution of sample and matrix are loaded by syringe pumps. On the end of the bioreactor, coaxial tubes were sealed to mix digests with a matrix solution and to deposit onto a MALDI target plate. (b) Assembled tryptic digest microfluidic chip: components including PMMA chip and cover slip, inlet and outlet connectors, capillary and stainless steel tube [79]. Source: Copyright 2008 American Society for Mass Spectrometry.
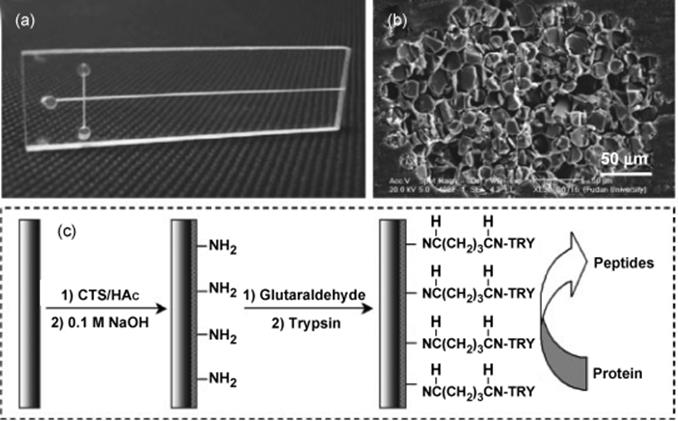
An interesting enzyme-immobilized microchip was reported by Fan and others (Figure 10.15) [81]. Fiberglass bundles were packed in a PMMA chip by in situ polymerization. The feasibility and performance of the unique microchip were demonstrated by analyzing the tryptic digestion of MYO and BSA. The on-chip digestion was carried out at a flow rate of 2.0 μl/min and the digestion time was significantly reduced to <5 s. The digests were identified by MALDI-TOF-MS with sequence coverages of 66% (MYO) and 40% (BSA) that were comparable to those obtained by the conventional in-solution tryptic digestion. This fiber-based microchip bioreactor provides a promising platform for the high-throughput protein identification.
Figure 10.15 (a) Photograph of a typical fiber-based microchip, (b) SEM image of the cross-section of a fiberglass-packed microchannel in the PMMA substrate, and (c) immobilization process of trypsin on the surface of glass fibers in the channel. Magnification of SEM 6400 [81]. Source: Copyright 2007 Wiley-VCH Verlag GmbH, Weinheim.
Electroosmotic flow (EOF) in the proteolytic system packed with immobilized trypsin gel beads was developed for digestions of proteins for 12 min [82]. The tryptic peptides were analyzed independently using CE and MALDI-TOF-MS. MALDI-TOF-MS was used to identify the parent protein and the tryptic peptides using MS-Fit database searching. The potential utility of the EOF-driven proteolytic system was demonstrated by direct electro-elution of proteins from an acrylamide gel into the proteolytic system, with elution and tryptic digestion achieved in a single step. The EOF-driven proteolytic system thus, provides a simple way to integrate protein digestion into an electrophoretic micro total analysis system for protein analysis and characterization.
The preparation of an easily replaceable protease microreactor for microchip application was described by Bilkova and colleagues [83]. The most important advantage of microsphere-packed microchip bioreactors is that the enzyme-immobilized microspheres in the channel can be replaced. Magnetic particles coated with poly(N-isopropylacrylamide), polystyrene, poly(2-hydroxyethyl methacrylate-co-ethylene dimethacrylate), poly(glycidyl methacrylate), [(2-amino-ethyl)hydroxymethylen]biphosphonic acid, or alginic acid with immobilized trypsin were utilized for heterogeneous digestion. The properties were optimized, with the constraint of allowing immobilization in a microchannel by a magnetic field gradient. To obtain the highest digestion efficiency, submicrometer spheres were organized by an inhomogeneous external magnetic field perpendicularly to the direction of the channel. Kinetic parameters of the enzyme reactor immobilized in microchip capillary (microchip immobilized magnetic enzyme reactor (IMER)) were determined. The capability of the IMER was demonstrated by five model (glyco)proteins.
The use of grafted trypsin magnetic beads in a microchip for performing protein digestion was described by Slovakova and others [84]. The PDMS device used strong magnets to create a magnetic field parallel to the flow with a strong gradient pointing through the center of the chip channel. A plug of low hydrodynamic resistance magnetic beads loaded with trypsin that serves as a matrix for protein digestion was formed. This device represents an inexpensive way of fabricating a multi open-tubular-like column with an appropriate pore size for proteins. Kinetics studies of the hydrolysis of a model peptide showed a 100-fold increase in digestion speed obtained by the microsystem when compared to a batch wise system. High performance and reproducibility for digesting recombinant human growth hormone are confirmed by analyzing the digest products in both CE and MALDI-TOF-MS. Similar sequence coverage (of about 44%) is obtained from MS analysis of products after 10 min on-chip and 4 h with soluble trypsin in bulk.
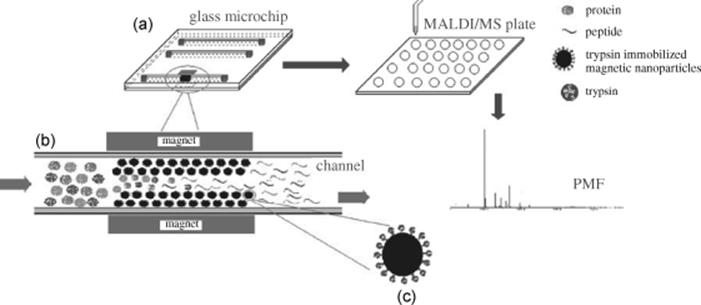
An easily replaceable microchip enzymatic microreactor has been fabricated based on the glass microchip with trypsin-immobilized superparamagnetic nanoparticles (Figure 10.16) [85]. Magnetic nanoparticles (50 nm in diameter) were synthesized. Amine-functionalized magnetic nanoparticles with high magnetic responsivity and excellent dispersibility were prepared through a facile one-pot strategy, followed by functionalization with numerous aldehyde groups by treating the amine-functionalized magnetic nanoparticles with glutaraldehyde. Finally, immobilization of trypsin onto the aldehyde-functionalized magnetic nanoparticles was achieved through reaction of the aldehyde groups with amino groups of trypsin. The prepared magnetic nanoparticles were packed onto the glass microchip by the application of a strong magnetic field using a magnet to form an on-chip magnetic nanoparticles packing bed. This proteolytic microreactor was used for complete digestion of Cyt-c, BSA, and MYO, which was achieved in 10 s under the flow rate of 5 μl/min. The digestion products were characterized using MALDI-TOF-MS with sequence coverage of 83%, 43%, and 79% observed, respectively.
Figure 10.16 (a) Schematic representation of on-chip protocol for protein digestion and identification coupled with MALDI-TOF/TOF-MS. (b) Amplified on-chip microreactor within the microchannel. (c) Amplified magnetic nanoparticles for trypsin immobilization [85]. Source: Copyright 2007 Elsevier.
A microreactor for proteinase-K (PK) digestion was prepared by immobilizing PK-grafted magnetic beads inside a PDMA microchannel using a longitudinal magnetic field parallel to the flow direction and a magnetic field gradient, thereby forming a matrix for enzymatic digestion [86]. The microreactor's capability was first evaluated using a model substrate, succinyl-ala-ala-ala-paranitroanilide (SAAAP). Reaction kinetics were typically accelerated a 100-fold as compared to conventional batch reactions. This microsystem was then applied to the digestion of prion protein from brain tissues. Controlled proteolysis could be obtained by varying the on-chip flow rate, while a complete proteolysis of normal protein was achieved in only 3 min.
Enzyme Immobilization by Cross-linking on Microchip Channel Surface
Immobilization of enzymes could be achieved by intermolecular cross-linking of the protein, either to other protein molecules or to functional groups on an insoluble support matrix. Cross-linking an enzyme to itself is both expensive and insufficient, as some of the protein material will inevitably be acting mainly as a support, resulting in relatively low enzymatic activity. Generally, cross-linking is best used in conjunction with one of the other methods. Since the enzyme is covalently linked to the support matrix, very little desorption is likely using this method.
An example of introduction of a functional group on the channel surface for covalent cross-linking was described by Ekstrom and group [87]. The immobilization of proteolytic enzymes onto the mirochip IMER was made according to standard procedures for enzyme coupling to silica matrixes. Trypsin was immobilized by surface modification of silicon by using 3-aminopropylsilane and glutaraldehyde. Protein solutions each equivalent to 1 μl volume of lysozyme, MYO, ribonuclease A, and Cyt-c were enzymatically digested within 1–3 min and identification with high sequence average of 50–100% was obtained.
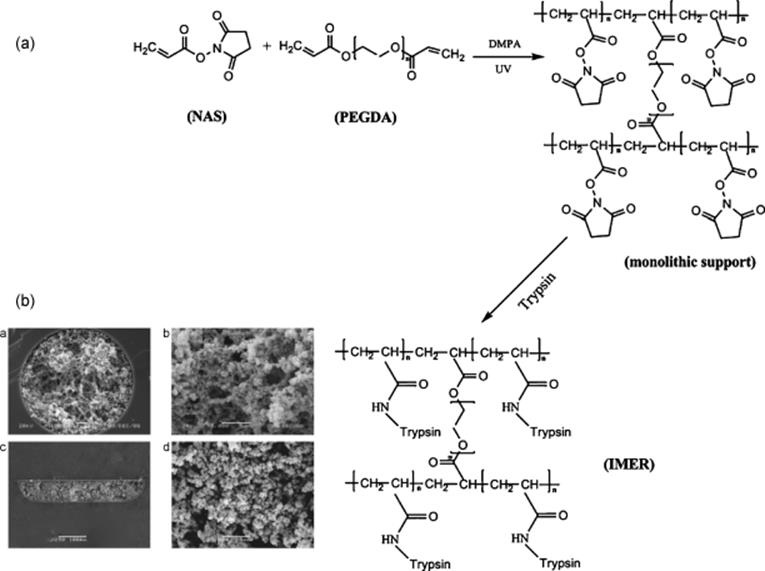
Recently, an integrated microchip-based IMER was described by Liang and others [88]. A novel kind of hydrophilic monolithic support was prepared on microchips by photopolymerization using poly(ethylene glycol)diacrylate (PEGDA) as hydrophilic cross-linker and N-acryloxysuccinimide (NAS) as active monomer, and then trypsin was immobilized on the monolithic support (Figure 10.17a and b). This microchip-based IMER was integrated with nanoRPLC–ESI-MS/MS via a packed electrospray emitter tip. Proteins were on-line digested by the microchip-based IMER and the digests were trapped by the electrospray emitter tip, and then separated by nanoRPLC–ESI-MS/MS. A mixture of BSA, MYO, and Cyt-c (200 ng for each protein) was analyzed. The sequence coverages for the three proteins were 51%, 61%, and 60%, respectively. To evaluate the applicability of the integrated platform for complex analysis, a randomly selected RPLC fraction of E. coli extract was analyzed, and for three consecutive runs, 48, 43, and 47 proteins were identified, respectively. The results from the analysis of standard proteins and complex samples are proofs that such a platform offers great potential for high-throughput proteome profiling.
Figure 10.17 (a) Procedures for the preparation of monolithic supports and trypsin immobilization. (b) SEM images of poly(NAS-co-PEGDA) monoliths in capillaries (a × 1000, b × 5000) and microchannels (c × 250, d × 5000 [90]. Source: Copyright 2011 Elsevier.
Enzyme Immobilization by Adsorption on Microchip Channel Surface
Physical adsorption of an enzyme onto a solid is probably the simplest way of preparing immobilized enzymes. The method relies on nonspecific physical interaction between the enzyme protein and the surface of the matrix, brought about by mixing a concentrated solution of enzyme with the solid. Binding forces involve ionic interactions, hydrogen bonds, van der Waals forces, hydrophobic interactions, and others. No reactive species are involved, hence, deleterious modifications and conformation changes, which may result in the change of biological activity will not pose as a problem. A major advantage of adsorption as a general method of insolubilizing enzymes is that it requires no reagents and involves minimum activation steps. Therefore, adsorption is easily carried out, and tends to be less disruptive to the enzymatic protein than chemical means of attachment, the binding being mainly by the forces mentioned above. This gives the advantage of keeping the structure and the activity of the enzyme. In this respect, the method bears the greatest similarity to the situation found in biological membranes in vivo and has been used to model such systems. However, because of the weak bonds involved, desorption of the protein resulting from changes in temperature, pH, ionic strength, or even the mere presence of substrate, is often observed. Another disadvantage in using immobilized enzymes by adsorption is the nonspecific further adsorption of other proteins or other substances. This may alter the properties of the immobilized enzyme or, if the substance adsorbed is a substrate for the enzyme, may probably decrease the rate of reaction depending on the surface mobility of enzyme and substrate. Adsorption of the enzyme may be necessary to facilitate the covalent reactions discussed in the earlier section. Stabilization of enzymes temporarily adsorbed onto a matrix has been achieved by cross-linking the protein in a chemical reaction subsequent to its physical adsorption.
An example of enzyme immobilized by adsorption was described by Gao and others, who developed a PDMS microreactor for proteolytic digestion with online ESI-MS identification [89]. Trypsin was adsorbed in a poly(vinylidine fluoride) porous membrane. Peptide identification for Cyt-c was reported using as little as 0.04 pmol.
A report by Liu and coworkers described layer-by-layer (LBL) coatings that have been assembled on the inner surfaces of the microchip [90]. Natural polysaccharides, positively charged chitosan (CS), and negatively charged hyaluronic acid (HA) were multilayer-assembled onto the surface of a poly(ethylene terephthalate) (PET) microfluidic chip to form a microstructured and biocompatible network for enzyme immobilization. Trypsin was adsorbed in the multilayer membrane composed of CS/HA assembled multilayers. The resulting peptide analysis has been carried out by MALDI-TOF-MS. The maximum proteolytic velocity of the adsorbed trypsin was ~600 mM/min/μg, thousands of times faster than that in solution. BSA, MYO, and Cyt-c were used as model substrates for the tryptic digestion. The standard proteins were identified at a low femtomole per analysis at a concentration of 0.5 ng/μl with the digestion time <5 s. This simple technique may offer a potential solution for low-level protein analysis.
A study by Liu and others demonstrated a PET microreactor coated with gold nanoparticle network in which trypsin was entrapped [91]. The nanostructured surface coating was assembled via a LBL electrostatic binding of poly(diallyldimethylammonium chloride) and gold nanoparticles. Enzyme within the two-bilayer assembly showed a maximum reaction rate of 2.0 mm/s, that is, the value of Vmax per unit of trypsin is about 400 mm/(min μg). Trace amounts of proteins down to femtomole per analysis were digested using the microchip reactor. The resulting tryptic products were identified by MALDI-TOF-MS/MS. The protein mixtures extracted from the mouse macrophages were efficiently identified by on-line digestion and LC–ESI-MS/MS analysis.
An enzymatic microreactor has been fabricated based on the PMMA microchip surface-modified with zeolite nanoparticles [92]. By introducing the silanol functional groups, the surface of PMMA microchannel has been successfully modified with silicalite-1 (S-1, all silica MFI-type zeolite nanoparticles) and trypsin was immobilized in the microchannel to form a bioreactor using silica sol–gel matrix. Trypsin immobilization was realized with a stable gel network through a Si—O—Si bridge via tethering to the silanol groups. The maximum proteolytic rate constant of the immobilized trypsin was measured ~6.6 mM/s. Using MALDI-TOF-MS, the proposed microreactor provides an efficient digestion of Cyt c and BSA at a fast flow rate of 4.0 μl/min, which affords a very short reaction time of <5 s.
A microfluidic reactor for rapid, low-pressure proteolysis using PMMA chip with pepsin-agarose immobilized by straightforward adhesion-based approach has been demonstrated by Liuni and others [93]. The device presented a shallow (10 μm) reactor “well” functionalized with pepsin-agarose. Electron spray ionization was carried out directly from a short metal capillary integrated into the chip outlet. Digestion efficiency was tested using horse heart myoglobin, bovine ubiquitin, and bovine BSA. Complete digestion was achieved in <4 s with 9% sequence coverage for myoglobin, <4 s with 64% sequence coverage for ubiquitin, and 15 s with 34% sequence coverage for BSA. Stress testing revealed little loss of performance over ~2 h continued use at high flow rates of 30 μl/min. The device provides a convenient platform for a range of applications in proteomics.
Immobilization of Enzymes on Microchip Channel Surface by Entrapment
Another approach for immobilization is entrapment method, which consists of the physical trapping of the active components (e.g., protease) into a film, gel, fiber coating, or microencapsulation [94]. The method involves a physical step, that is, confining enzymes within the lattices of polymerized gels. This allows the free diffusion of low-molecular weight substrates and reaction products. The reaction involves hydrolysis and polycondensation of alkoxysilane monomers. During the process, proteins are incorporated into a gel matrix. The usual method is to polymerize the hydrophilic matrix in an aqueous solution of the enzyme and break up the polymeric mass to the desired particle size. Although there is no chemical bond formation between the enzyme and the polymer matrix, the proteins can be chemically modified without loss of their activity by conjugation to one of the monomers before polymerization leading to covalent bond to the matrix [95]. The advantages of this immobilization method are the extremely large surface area between the substrate and the enzyme, within a relatively small volume, and the real possibility of simultaneous immobilization [96].
Wu and group reported a titania-based and alumina-based (PDMS) microfluidic enzymatic reactor along with their analytical features in coupling with MALDI-TOF and ESI-MS [97]. Microreactors with microchannel and stainless steel tubing (SST) were fabricated using PDMS casting and O2-plasma techniques, and were used for the preparation of an enzymatic reactor. Plasma oxidation of the PDMS microfluidic system enabled the microchannel wall to generate a layer of silanol groups. These silanol groups act as anchors onto the microchannel wall linked covalently with the hydroxyl groups of trypsin-encapsulated sol matrix. As a result, the trypsin-encapsulated gel matrix was anchored onto the wall of the microchannel, and the leakage of the gel matrix from the microchannel was effectively prevented. These microdevices provided an excellent extent of digestion even at a fast flow rate of 7.0 μl/min, which afforded a very short residence time of about 2 s. The encapsulated trypsin exhibited an increased stability even after continuous use. Such features are required for high-throughput protein identification.
Another report of immobilization of enzymes on microchannel surface demonstrated sol–gel structures on PMMA [98]. The PMMA surface was modified with a copolymer of butyl methacrylate/γ-(methylacryloxy) propyltrimethoxysilane and silica-sol–gel to immobilize trypsin. Trypsin was immobilized through the silicon–oxygen–silicon bridge formed by tethering to the silane functional groups attached on PMMA. Subsequent attachment of silica sol–gel with the silane-functionalized groups facilitated encapsulation of enzyme onto the hydrophobic plastic-based microchip. Using this microreactor, Cyt-c and BSA were digested at a fast flow rate of 4.0 μl/min, which afforded a residence time of <5 s. The digestion products were characterized using MALDI-TOF-MS with sequence coverage of 75% and 31% for Cyt-c and BSA, respectively.
Enhancement of proteolysis through the silica-gel-derived microfluidic reactor was reported using a PET microchip fabricated by laser ablation [99]. The PET surface was subjected to hydrolysis reaction and protease was encapsulated using silica sol–gel. The carboxyl and hydroxyl groups extending from the hydrolyzed surfaces contributed to the stable construction of silica sol–gel network on the polymeric surface. The capability of the microreactor was tested using protein solutions of BSA, Cyt-c, MYO, and hepatitis A virus vaccine injected through the gel-modified PET microchannel at a flow rate of 120 μl/h. High proteolysis efficiency was observed which could be attributed to the high surface area to volume ratio and the microscopic confinement of the gel-derived microreactor. For BSA and Cyt-c digests, 30% and 69% amino acid sequence were identified, respectively. The reactor can be used for low-level protein digestion at 16 fmol per analysis.
Sakai-Kato and group presented trypsin encapsulation in sol–gel, which is derived from alkoxysilane [100]. The trypsin encapsulated in silica-sol–gel was fabricated in a sample reservoir of the PMMA chip. Fluorescent-labeled arginine ethyl ester (ArgOEt) and bradykinin were digested within the sample reservoir. This microchip is applicable to digest proteins with multiple cleavage sites and separate digested fragments. The encapsulated enzyme exhibited increased stability even after continuous use, compared with the free enzyme in solution.
An on-chip microreactor was proposed toward the acceleration of protein digestion through the construction of a nanozeolite-assembled network [101]. The nanozeolite microstructure was assembled using a LBL technique based on poly(diallyldimethylammonium chloride) and zeolite nanocrystals. Trypsin was entrapped in nanozeolite network. It was found that the controlled trypsin-containing nanozeolite networks assembled within a microchannel could act as a stationary phase with a large surface-to-volume ratio for the highly efficient proteolysis of both proteins at low levels and with complex extracts. The maximum proteolytic rate of the adsorbed trypsin was measured to be 350 mM/min/μg, much faster than that in solution. Moreover, due the large surface-to-volume ratio and biocompatible microenvironment provided by the nanozeolite-assembled films as well as the microfluidic confinement effect, the low-level proteins down to 16 fmol per analysis were confidently identified using the microreactor in <5 s. The capability of the microreactor was further demonstrated in the identification of the complex extracts from mouse macrophages integrated with two-dimensional liquid chromatography–electrospray ionization-tandem mass spectrometry.
Other Techniques for Immobilization of Enzyme in Microreactor
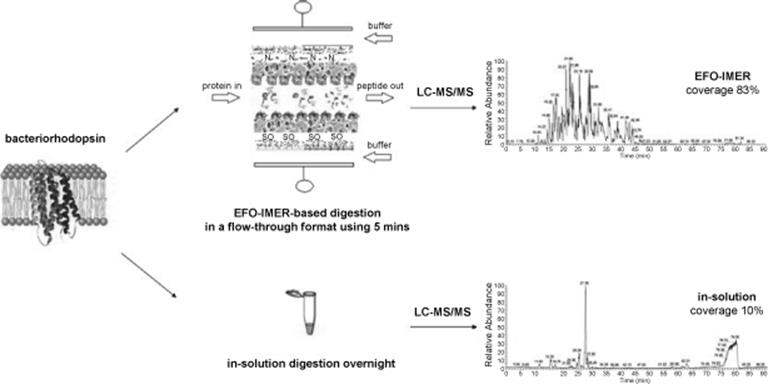
Rapid and enhanced proteolytic digestion using an electric-field-oriented enzyme reactor was recently described by Zhou and others [102]. This research group created a novel enzyme reactor using electric field-mediated orientation and immobilization of proteolytic enzymes (trypsin/chymotrypsin) on biocompatible PVDF membranes in a continuous flow-through chamber (EFO-IMER) (Figure 10.18) [102]. They were able to fix the enzyme dipoles at the surface of PVDF by employing a controllable electric field under 50 V in a continuous flow-through chamber within only 15 min. In <5 min, this reactor in various enzyme combinations can produce enhanced rapid digestion at an optimized flow rate of 2 μl/min or less for standardized prototypic proteins, hydrophilic proteins, and hydrophobic transmembrane proteins when compared to in-solution techniques. With improved digestive efficiency, EFO-IMER improved the overall functional analysis of lipid raft proteomes by identifying more closely functionally linked proteins and elucidated a richer set of biological processes and pathways linked to the proteins than traditional in-solution methods.
Figure 10.18 Graphical representation of an experiment on the development of an enzyme reactor using electric field-mediated orientation and immobilization of proteolytic enzymes (trypsin/chymotrypsin) on PVDF membranes in a continuous flow-through chamber. LC–MS/MS spectra show enhanced rapid digestion efficiency in EFO-IMER when compared to in-solution techniques [102]. Source: Copyright 2011 Elsevier.
Immobilization of Enzyme in Capillary Microreactor
IMER systems in capillary microreactors are now routinely being used in protein digestion for peptide mapping and proteomics. Similar to microfluidic channels on chips, enzyme immobilization in capillary can be classified into two categories based on the immobilization location. First, a wall or assembly of membrane channels called open channel or open tubular capillary and secondly, a solid support inside channels such as monoliths [103–105], sol–gel [106–110], controlled pore glass, silica, and polystyrene beads [111–115]. The representative IMERs in Table 10.2 presents the various developed IMERs in capillary and the different immobilization techniques employed.
Work has been geared toward incorporation of further peptide analysis by MALDI-TOF and ESI-TOF. Fused silica capillary was also employed as a support for immobilizing proteases to fabricated microfluidic reactors for efficient proteolysis. The cost of capillaries are much cheaper than microfluidic chips, hence they have been widely used in fabrication of microreactors.
Enzyme Immobilization by Covalent Linking in Capillary Microreactor
This technique of immobilization involves formations of a covalent bond between the enzyme and support material. Covalent bonds usually provide the strongest linkages between enzyme and carrier. Leakage of enzyme is often minimized with covalently bound immobilized enzymes. Using the inner surface of a fused silica capillary as the direct carrier for enzyme immobilization, Licklider and Kuhr immobilized trypsin to the inner wall of a capillary via carboxyl-modified beads [116]. Digestion of globular proteins Cyt-C and lactalbumin was enhanced by applying mild vibration driven by a sinusoid waveform (~0.75 W), which was delivered by the audio oscillator at a frequency of 200 Hz or at another frequency characteristic of the piezoelectric element (i.e., 750 Hz, 2.3 kHz, 3.3 kHz, and 7.1 kHz).
Bossi and his colleagues [117] achieved enzyme immobilization on a portion of silica capillary by photo-initiated polymerization. The capillary was then filled with N-(4-azidobenzoyloxy)succinimide (ABS), which was cross-linked to the capillary surface upon UV light exposure. The enzyme was immobilized via the succinimide group of ABS. The enzyme was immobilized with a surface density of 15.8 μg/cm2, the stability was high (80% after 38 days) and the rate of conversion was 0.2 ng/s. On-line protein mapping was tested with proteins of different dimensions, showing competitiveness in terms of time (completed map within 15 min) and exhaustive maps of small proteins. The performance of the CE-microreactor in peptide mapping was studied with different protein substrates. Carbonic anhydrase, Cyt-c, and hemoglobin were used as substrates. The protein substrates were injected in the CE-microreactor at a concentration of 1-1.1 mg/ml by pressure-injection into the capillary at 3 psi/s and a voltage of 2 or 10 kV was applied.
Zhao's group also integrated an immobilized enzyme microreactor in fused silica capillaries with a nanoelectrospray mass spectrometry for on-line digestion and fast peptide mass mapping [118]. Trypsin was immobilized onto the surface of the inner wall of the fused silica capillary tubing as illustrated in Scheme 10.11. The procedure was demonstrated using dilute solutions of angiotensin II, Cyt-c, hemoglobin, and β-casein. Because the inner walls of the capillaries are modified by covalent chemical bonds, the adsorption of peptides and proteins to the inner walls of the capillaries is suppressed. This procedure was performed with solutions as dilute as 1 fmol/μl (1 nM) Cyt-c. This method shows generation of tryptic peptides with sequence coverage up to 90% within minutes, without detection of any trypsin autolysis products. In addition, the immobilized enzyme can be cleaned easily, enabling the microreactor to be reused for nanoelectrospray.
Scheme 10.11 Immobilization of trypsin onto the inner wall surface of the capillary [118]. Source: Copyright 2006 Elsevier.
Krenkova and others [75] studied open tubular capillary enzyme reactors for rapid protein digestion and possible on-line integration into a CE-ESI-MS system. The need to minimize the time of the analyte molecules to diffuse toward the surface immobilized enzyme and to maximize the surface-to-volume ratio of the open tubular reactors dictated the use of very narrow bore capillaries. Extremely small protein amounts (atto-femtomoles loaded) could be digested with enzymes immobilized directly on the inside wall of a 10 μm inner diameter capillary. The capillary was modified with (3-aminopropyl)trimethoxysilane and l-1-tosylamido-2-phenylethyl chloromethyl ketone. The enzymatic activity was characterized in the flow-through mode with on-line coupling to electrospray ionization-time of flight-mass spectrometer (ESI-TOF-MS) under a range of protein concentrations, buffer pHs, temperatures, and reaction times. The optimized reactors were tested as the nanospray needles for fast identification of proteins using CE-ESI-TOF-MS.
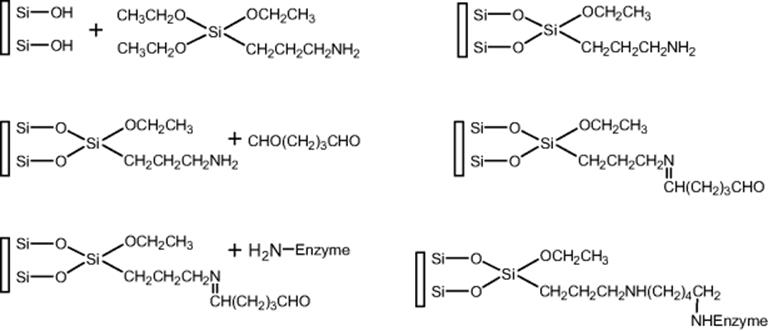
Covalent linking of enzyme on fused silica capillaries was also demonstrated by Stigter [119]. Pepsin was covalently immobilized on dextran-modified capillaries as illustrated in Scheme 10.12. The applicability of pepsin-immobilized microreactor was tested with a number of proteins varying in molecular weight, isoelectric point, and sample composition. This open tubular microreactor demonstrated a flow-dependent digestion as represented by native hemoglobin. Complete digestion of human hemoglobin, horse myoglobin, and BSA was observed at a flow rate of 1 μl/min and residence time of 3 min with on-line micro-HPLC with UV detection. The sequence coverage for hemoglobin β- and α-chain were 55% and 97%, respectively, while the sequence coverage for various casein variants were 35–55%. The protein sequence of these and other proteins could largely be retrieved with a database search on the MS/MS data, which make the microreactors an interesting tool for the characterization of unknown proteins.
Scheme 10.12 Scheme of the surface modification and enzyme immobilization in fused silica capillaries [119]. Source: Copyright 2008 Elsevier.
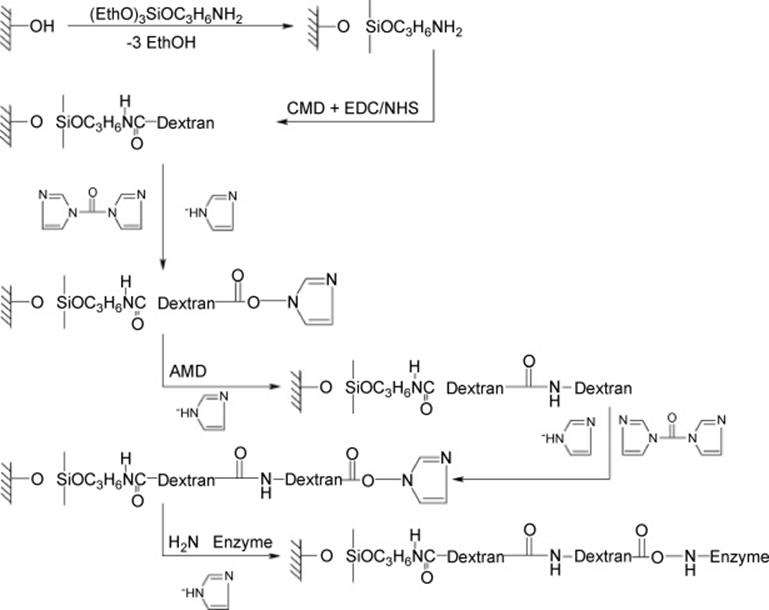
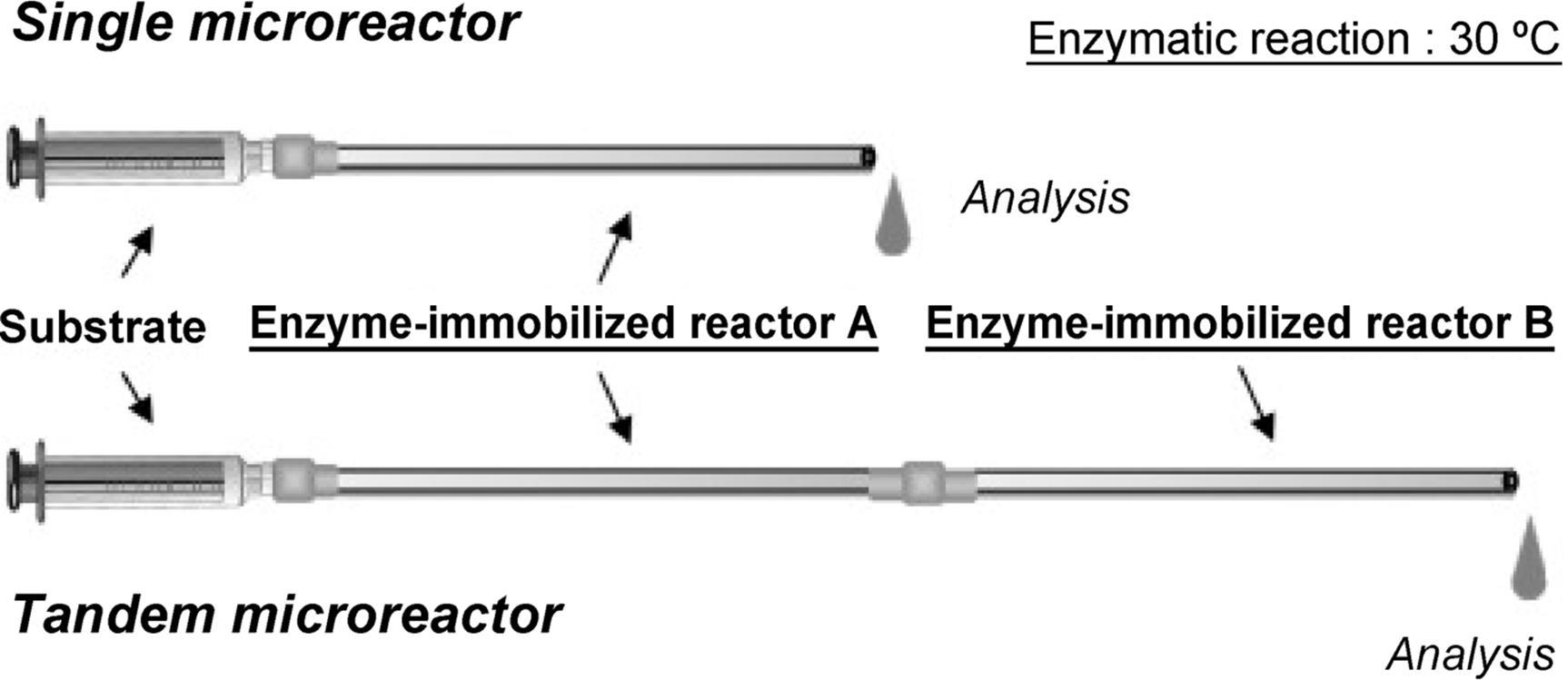
The Miyazaki group developed a preparation method for nanostructure on the microchannel surface formed by sol–gel like simple treatment with 3-aminopropyltriethoxysilane [120]. In addition, a novel and facile method for the preparation of an enzyme-immobilized microreactor was proposed (Figure 10.19) [121]. Enzymes are immobilized as an enzyme-polymer membrane formed on the inner wall of the microchannel by a cross-linking polymerization using bifunctional cross-linker agents, paraformaldehyde and glutaraldehyde. The resulting microreactor showed excellent reaction performance and stability against denaturating agents. Furthermore, this novel method was applied to more enzymes using a supporting material, lysine-polymer (Figure 10.20) [122]. Utilizing this method, Yamaguchi and colleagues [123] developed tandem-enzymatic microreactors for protein digestion (Figure 10.21). Single trypsin microreactor (TY) digested Cyt-c at a flow rate of 2.5 μl/min for 10.4 min and yielded 93% sequence coverage. Chymotrypsin (CT) reactor digested Cyt-c with 38% sequence coverage. TY-CT tandem microreactor digested Cyt-c for 20.8 min with 88% sequence coverage. Tandem microreactors digested β-casein with higher sequence coverage (70%) than in-solution digestion. Proteolysis was achieved within a short period of time (~10 min) at 30 °C. The approach using a tandem enzyme-immobilized microreactor coupled with MS, and using poly-lys as a booster can be a useful approach for rapid analysis of protein sequence and for other postranslational modification analyses.
Figure 10.19 Preparation of enzyme-membrane on the inner wall of a PTFE tube. (a) Enzyme and aldehyde solutions were each charged into a 1 ml syringe, the solutions were supplied to a PTFE tube using a syringe pump. (b) Cylindrical enzyme-membrane (dry state) exposed from PTFE tube, which forms on the inner wall of the tube. (c) Possible mechanism of polymerization process of enzyme and cross-linker reagent in a microchannel [121]. Source: Copyright 2005 The Royal Society of Chemistry.
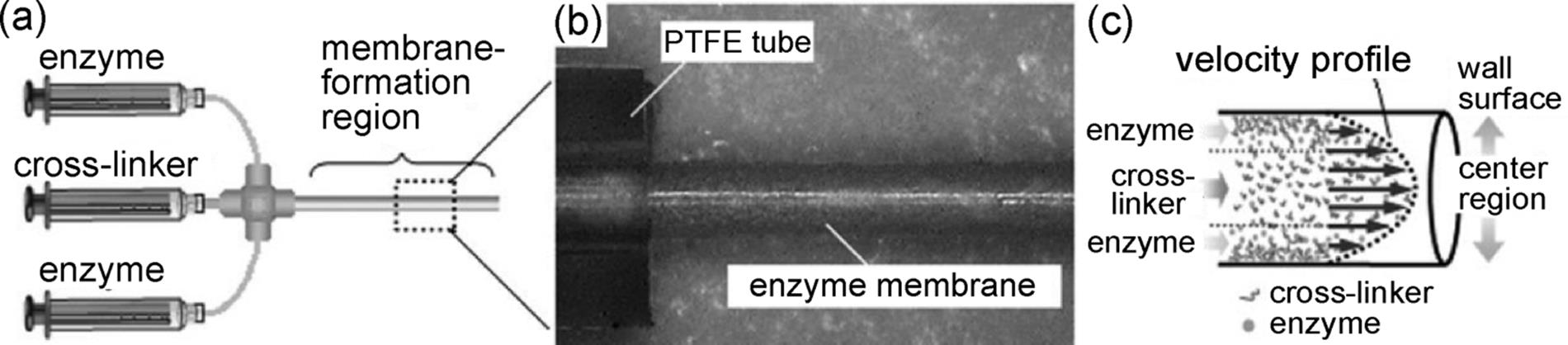

Figure 10.20 (a) Schematic illustration of the procedure used to prepare an acylase-CEM. The cross-linking polymerization was performed in a concentric laminar flow. A silica capillary was fitted to the outer diameter of the T-shape connector by attaching to a PTFE tube using heat-shrink tubing. The capillary was set in the connector located at the concentric position of the CEM tube. The cross-linker solution was supplied to the substrate PTFE tube through the silica capillary, corresponding to a central stream in the concentric laminar flow. A solution of acylase-poly(Lys) mixture was poured from the other inlet of the T-shape connector, and formed an outer stream of the laminar flow. CCD images of (b) cylindrical enzyme-membrane (dry state) exposed from the PTFE tube, which forms on the inner wall of the tube and (c) a sectional view of the obtained CEM [122]. Source: Copyright 2006 Wiley-VCH Verlag GmbH, Weinheim.
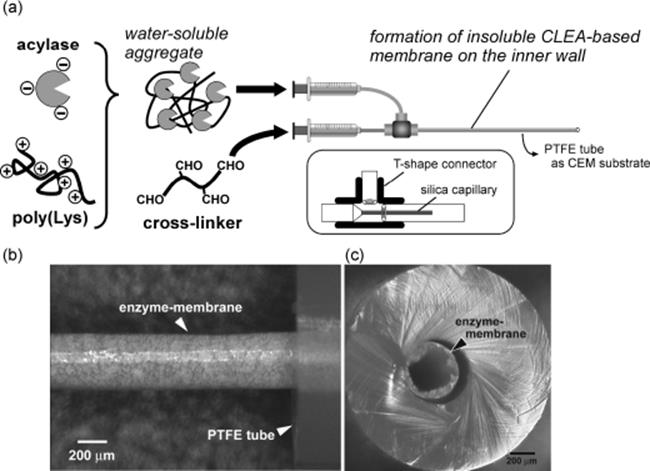
Figure 10.21 Schematic representation of enzymatic reaction using enzyme-immobilized microreactors. The substrate was pumped through the enzyme-immobilized microreactors using a syringe pump. The reaction was carried out at 30 °C. The digests were collected in a sample tube and then analyzed by reverse-phase HPLC and ESI-TOF-MS [123]. Source: Copyright 2010 Elsevier.
Although the vast majority of microfluidic reactors feature open channel architectures, the channels exhibit small surface-to-volume ratios and since only channel walls can be used to provide the desired interactions, the microreactor can handle only minute amount of compounds. This lead to the attempts to extend the techniques to the microfluidic format that enabled packing the chip with immobilized enzyme beads. However, the difficulty of packing a bed of beads in the channel prompted researchers to turn to monolith materials. Monolith materials have also been used as supports for immobilization of enzymes. Monolithic enzyme reactor have the unique properties that include better accessibility of the active site for substrates, very low back pressure stability in most solvents, and versatility of the active functional group available on pore surface of the monoliths. The major advantage of monolithic supports is the outstanding mass transfer due to the convective flow through pores. Peterson and coworkers prepared enzymatic capillary microreactor by immobilizing trypsin on porous monoliths consisting of 2-vinyl-4,4-dimethyl azlactone, ethylene dimethacrylate, and acrylamide or 2-hydroxyethyl methacrylate. Immobilization of trypsin was achieved by covalent bonding of the amine and thiol groups of the enzyme with the azlactone functionalities. The performance of the monolithic microreactors has been demonstrated with BAEE and myoglobin. The proteolytic activity of myoglobin was evaluated at 0.5 μl/min flow rate with residence time of 11.7 s. The digests were characterized off-line using MALDI-TOF-MS. The sequence coverage obtained was 67%. These results have opened new avenues for the design and construction of rapid, high-throughput, protein mapping systems.
Ye and others reported a nanoliter enzyme microreactor that enables on-line protein digestion and peptide mapping by capillary electrophoresis with post-column labeling for laser-induced fluorescence detection [125]. The enzyme microreactor was formed by immobilizing trypsin onto a monolithic capillary column, which was prepared by in situ polymerization of glycidyl methacrylate and ethylene dimethacrylate in a capillary. Digestion efficiency was evaluated with substrate N-α-benzoyl-dl-arginine-ρ-nitroanilide (BAPNA) and α-lactalbumin. The microreactor has a 30 nl volume and is coupled with a separation capillary via a fluid joint for on-line digestion. It took only 16 min for overall analysis, including digestion and separation. Column efficiencies of >300 000 plates per meter were obtained for most peaks in the electropherogram of an on-line mapping experiment of denatured α-lactalbumin under optimal conditions.
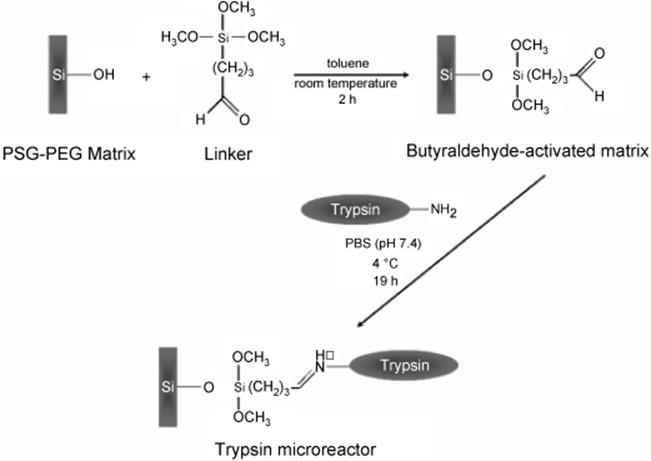
Trypsin covalently linked to a photopolymerized sol–gel monolith modified by incorporating poly(ethylene glycol) (PSG-PEG) (Scheme 10.13) was described by Dulay and others [126] for on-column digestion of N-α-benzoyl-l-arginine ethyl ester (BAEE) and peptides, neurotensin, and insulin chain B. They prepared a novel IMER by preparation of a reactive hydrophilic macroporous PEG-modified photo-polymerized sol–gel (PSG-PEG) monolith, followed by the functionalization with an alkoxysilane reagent containing aldehyde group via Schiff chemistry and the covalent attachment of trypsin to butyraldehyde-activated PSG-PEG monolith was achieved by the reaction of the aldehyde group of the bound linker with the terminal primary amine of trypsin. Proteolytic acitivity of a trypsin-PSG-PEG monolith for the digestion of BAEE was evaluated by the hydrolytic cleavage of the ester linkage in BAEE to form Nα- benzoyl-l-arginine (BA). BA product was formed even when substrate was exposed for only 10 s. The BA was separated from BAEE by capillary electrophoresis and detected downstream by absorption (254 nm). Digestion of neurotensin was demonstrated by the respective fragment peaks. The residence time of the neurotensin sample plug in the monolith determined how much of the neurotensin was digested. For insulin chain B, digestion was observed within 15 min as suggested by the resulting 48% decrease of the insulin peak area. The enzyme microreactor formed by covalent attachment of the enzyme trypsin was able to facilitate protein digestion at short digestion times under low temperature in contrast to the longer reaction times at higher temperatures in tryptic digests in free solution. The enhanced proteolytic activity of the immobilized trypsin could be ascribed to the elimination of trypsin autolysis, stabilization of the trypsin structure, resulting in higher accessibility of the substrate to the active site of the enzyme. Aside from the covalent linking, the hydrophilic PSG-PEG monolith contributed to the enhancement of the structural stability of immobilized trypsin by providing a favorable microenvironment.
Scheme 10.13 Strategies for functionalization of the PSG-PEG monolith surface with trimethoxysilylbutyraldehyde linker and attachment of trypsin to a butyraldehyde-activated PSG-PEG monolith [126]. Source: Copyright 2005 American Chemical Society.
Duan and group developed a monolithic enzymatic microreactor in a fused silica capillary by in situ polymerization of acrylamide, glycidyl methacrylate, and ethylene dimethacrylate in the presence of a binary pyrogenic mixture of dodecanol and cyclohexanol, followed by ammonia solution treatment, glutaraldehyde activation, and trypsin modification [127]. Improvement of the efficiency of trypsin modification was achieved by employing acrylamide as comonomer was found useful thus, increasing the enzyme activity. The optimized microreactor offered very low back pressure enabling the fast digestion of proteins flowing through the reactor. The performance of the monolithic microreactor was demonstrated with the digestion of Cyt-c at high flow rate. The digests were then characterized by CE and HPLC–MS/MS with the sequence coverage of 57.7%. The digestion efficiency was found over 230 times higher compared with the conventional method.
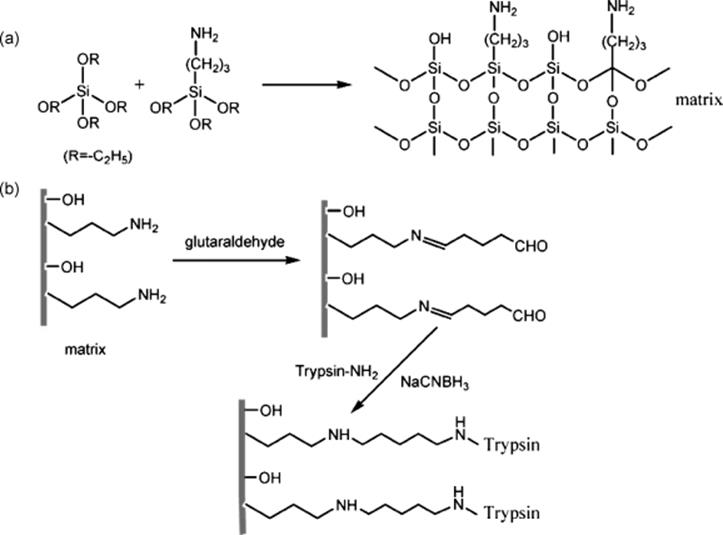
Another novel immobilized trypsin reactor based on organic–inorganic hybrid silica monoliths has been developed by Ma and others [128]. With the presence of cetyltrimethyl ammonium bromide in the polymerization mixture, the hybrid silica monolithic support was prepared in a 100 μm inner diameter capillary by the sol–gel method with tetraethoxysilane (TEOS) and 3-aminopropyltriethoxysilane (APTES) as precursors. The electrostatic interaction played an important role in the formation of a network. The resulting monolithic support bearing amine groups was activated using a common bifunctional reagent, glutaraldehyde, and trypsin was covalently immobilized (Scheme 10.14). Enzymatic activity of immobilized trypsin was investigated using a synthetic decapeptide, C-myc (EQKLISEEDL). Results showed that the digestion speed was about 6600-fold with the immobilized trypsin compared to that performed in free solution. The performance of such a microreactor was further demonstrated by digesting myoglobin, with the digested products analyzed by microflow reversed phase liquid chromatography coupled with tandem mass spectrometry (microRPLC–MS/MS). The yielding sequence coverage for on-column digestion was 92%, the same as that obtained by in-solution digestion, whereas the residence time of myoglobin in the immobilized enzyme reactor was mere 30 s about 1500 times shorter than 12 h required to achieve the same degree of digestion in solution. The applicability of the microreactors for real-life proteome analysis was also proven effective as demonstrated in the digestion proteins extracted from E. coli in a 10 cm long microreactor and LC/MS/MS. A total of 208 proteins were identified after 150 s digestion time in the microreactor compared to the 176 proteins recognized after 24 h long in-solution digestion.
Scheme 10.14 Scheme of the organic–inorganic hybrid silica monolithic matrix formation (a) and trypsin immobilization (b) [128]. Source: Copyright 2008 American Chemical Society.
Lin and Skinner fabricated polymer-based monolithic enzyme reactors in fused silica capillaries, and characterized their performance through protein digestion and identification by MALDI-MS and ESI-MS [129]. Microreactors were prepared by fabricating a porous methacrylate base monolith followed by photografting with glycidyl methacrylate, and immobilization of the enzymes with carbonyldiimidazole. Trypsin and Staphylococcus aureus V-8 protease (Glu-C) were used to produce three types of reactors: trypsin-based, Glu-C-based, and trypsin combined with Glu-C. Protein digestions, performed by perfusing (flushing) protein solutions through the reactor under pressure, were evaluated based on the peptide map generated when directly coupled to an ESI mass spectrometer. Efficient digestion was observed over flow rates from 0.2 to 1 μl/min, which corresponds to reactor residence times of 0.24–1.4 min. Chromatographic separation of model proteins followed by the digestion of specific fractions using these proteolytic enzyme reactors and ESI-MS was demonstrated to show the efficiency of the platform.
A few years ago, Sinz described a preparation of capillary trypsin IMER for a rapid and efficient digestion of proteins down to the femtomole level [130]. Trypsin was immobilized on a poly(glycidyl methacrylate-co-acrylamide-co-ethylene glycol dimethycrylate) monolith by glutaraldehyde technique. Digestion efficiencies were evaluated using mode protein, BAEE and mixture of proteins; Cyt-c, lysozyme, MYO, β-lactoglobulin, myoglobin, hPPARα, and BSA at 300 nl/min flow rate, with digestion time of 231 s. Digestion of cross-linked lysozyme at flow rate of 100 nl/min, digestion time of 11 min and 33 s allowed for identification of two cross-linking products in lysozyme. The greatest advantages of this trypsin IMER are short digestion times and processing of low-abundance proteins.
An integrated platform with the combination of protein and peptide separation was established online by Wang and group, by which proteins were first separated by capillary isoelectric focusing, online digested by a trypsin immobilized enzyme microreactor, trapped and desalted by two parallel columns, separated by nanoreversed phase and finally identified by MS (Figure 10.22) [131]. In this platform, two hollow fiber membrane interfaces were used; one to supply catholyte (the electrolyte adjacent to the cathode of an electrolytic cell) and electric contact, and another to supply adjustment buffer to improve the compatibility of protein separation and digestion. A hybrid silica monolith-based immobilized trypsin microreactor was used for online protein digestion. To evaluate the performance of the microreactor, a 0.29 μg of a four-protein mixture of BSA, myoglobin, β-globulin, and ribonuclease A was digested, and sequence coverages of 8, 26, 10, and 54%, respectively, were obtained. Protein extracts from E. coli were also digested and 101 proteins were identified, suggesting that the integrated platform holds promise for low abundance protein digestion and identification.
Figure 10.22 Schematic diagrams of the hollow fiber membrane interface (a) and the CIEF-IMER-nanoRPLCMS platform (b) [132]. Source: Copyright 2011 Wiley-VCH Verlag GmbH, Weinheim.
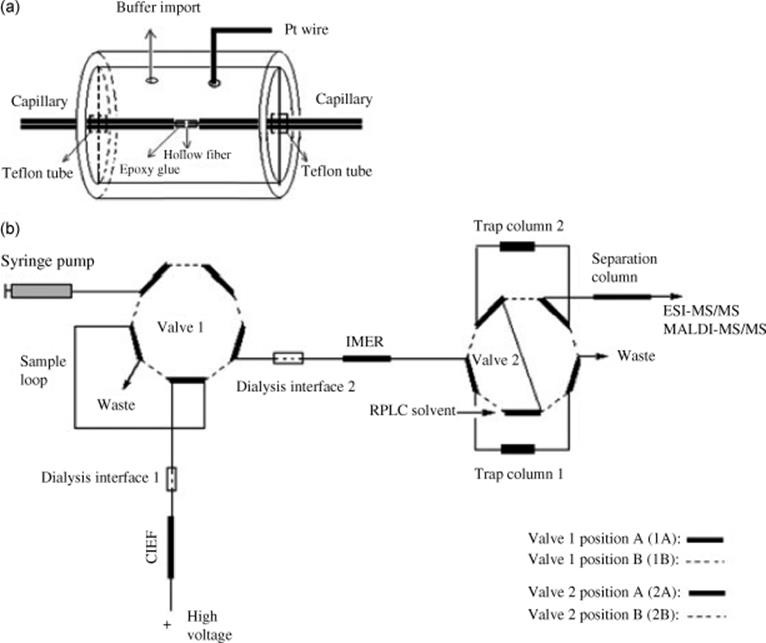
Recently, a replaceable microreactor for on-line protein digestion in a two-dimensional capillary electrophoresis system with tandem mass spectrometry detection was developed by Li and others [132]. Trypsin was immobilized onto the carboxylic acid-functionalized surface of the magnetic beads via EDC activation. The capillary reactor was coupled to CE-ESI-MS. The performance efficiency of the system was checked by digestion and separation of model proteins, insulin chain b oxidized, and β-casein. This platform delivers tryptic digests to the MS within a short window. The two-dimensional separation requires roughly 30 min to digest and separate 30 fractions.
All the publications discussed above are successful demonstrations that protein digestion through immobilized enzyme microreactors enables faster, efficient, and automated protein identification via peptide mapping while using very small amount of samples.
Immobilization of Enzyme by Adsorption in Capillary Microreactor
Adsorption leading to formation of physical links between the enzyme molecule and the surface of the capillary wall via hydrogen bonds, hydrophobic interactions, ionic bonds, and so on, is another enzyme immobilization technique applied in capillary microreactors. Immobilization by adsorption is the simplest method and involves reversible surface interactions between enzyme and support material as shown in Figure 10.9. The procedure of adsorption consists of mixing together the enzyme and a support with adsorption properties under suitable conditions of pH and ionic strength for a period of incubation. There are disadvantages of enzymes immobilized using adsorption technique such as leakage of enzymes from the carriers due to weak interaction between enzyme and carrier, which can be destroyed by desorption forces such as high ionic strength, pH, and so on steric hindrance by the support, nonspecific bonding, overloading on the support, and contamination of product. Therefore, a number of variations to counter these drawbacks have been developed such as adsorption-cross-linking, modification-adsorption, adsorption-coating, and selective adsorption-covalent attachment. A detailed discussion of these variations is presented in the book by Cao and Schmid [133]. Guo and others developed an immobilized metal-ion chelating capillary microreactor combined with MALDI-TOF-MS, which offers advantages such as ease in regeneration, less consumption of sample, and excellent reproducibility [134]. A regenerable capillary microreactor was prepared by synthesis of the chelator silane reagent and then reaction onto the capillary wall following the immobilization of the metal ion and enzyme. The metal ion of copper and subsequently enzyme was specifically adsorbed onto the surface of the capillary wall. The chelator of iminodiacetic acid (IDA) was first reacted with 3-glycidoxypropyltrimethoxysilane (3-GLYMO) to form GLYMO-IDA silane under optimum conditions of pH 11, then the GLYMO-IDA-silane was immobilized onto the inner capillary wall. Protein of Cyt-c (0.05 pmol) was digested using the microreactor for 15 min. As low as 20 fmol of Cyt-c could be digested.
Xu and his colleagues prepared an enzyme-immobilized capillary microreactor for rapid protein digestion and proteomics analysis [135]. The inner surface of the fused silica capillary was coated with poly(diallyldimethylammonium chloride) (PDDA)-entrapped silica sol–gel matrix, followed by assembly of trypsin onto the PDDA-modified surface via electrostatic adsorption. Protein samples including β-casein, myoglobin, and cytochrome c could be effectively digested and electrophoretically separated in such a modified capillary. Just 2.26 ng (corresponding to 0.10–0.14 pmol) of sample was sufficient for on-line capillary electrophoresis peptide mapping. The efficiency of the digestion was further demonstrated by digestion of a human liver cytoplasm sample and 253 proteins were identified in one unique run.
An enzymatic reactor was prepared by trypsin dynamically immobilized on HILIC (hydrophilic interaction chromatography) matrix packed in a polymer-sheathed fused silica tubing, Peeksil column [136]. Trypsin was immobilized in the HILIC matrix by dynamic immobilization method which allows enzyme to be linked with immobilized matrix by multiple interactions such as van der Waals force, hydrogen bonding interaction, ionic bonding interaction, and hydrophobic interaction. It could be found that the microreactor possess high hydrolytic activity according to the hydrolysis results of a substrate of trypsin, BAPNA. Moreover, high-enzymatic activity performance of the microreactor was demonstrated by standard proteins, BSA, Cyt-C, and MYO. The results showed that the enzymatic reactor can realize on-line enzymolysis of proteins, and the time to analyze and identify the proteins was greatly shortened.
Immobilization of Enzyme in Capillary Microreactor by Entrapment
Entrapment technique involves the entrapment of enzymes in gel matrix. The enzyme is mixed with gel formation ingredients and upon gel formation, the enzyme remains trapped in the matrix. Another form of entrapment is the formation of membrane around the droplet of enzyme, which is typically in solution. Immobilization by entrapment differs from adsorption and covalent bonding in that enzymes are free in solution but restricted in movement by the lattice structure of a gel. The membrane must be permeable to diffusion of substrate and product molecules but tight enough to prevent leakage of enzyme. Hence, properties such porosity, chemistry, and chemical strength of the support matrix are the key factors controlling the activity and the applicability of the microreactor. Entrapment can be achieved by mixing the enzyme with a polyionic polymer material, and cross-linking the polymer with multivalent cations.
Sakai-Kato's group developed a facile in situ encapsulation procedure to prepare IMER [110]. Trypsin was encapsulated in the hydrogel by mixing fully or partially hydrolyzed silane. In comparison to trypsin in free solution, the bound trypsin exhibited about 700 times higher enzymatic activity. This microreactor was only suitable for small molecules since large molecules could not pass through the nanopores of the hydrogel network. In 2004, they improved this technique by coating trypsin-containing gel onto the porous silica monolith (Figure 10.23) [139]. A miniaturized pepsin reactor inside a fused silica capillary (75 μm inner diameter) was prepared by coating a pepsin-containing gel on a photopolymerized porous silica monolith. The pepsin-encapsulated film was prepared by a sol–gel method. The sol–gel reaction was optimized so that the sol solution containing pepsin forms a thin film on the photopolymerized sol–gel (PSG) monolith that was initially fabricated at the inlet of the capillary. Pepsin was encapsulated into the gel matrix without losing its activity. The large surface area of the PSG monolith enabled the immobilized pepsin to achieve a high catalytic turnover rate, and the porous nature of the PSG promotes penetration of large molecular proteins into the column. The immobilized pepsin-digested peptides and proteins, and the resulting mixture of peptide fragments, could be directly separated in the portion of the capillary where no PSG monolith exists. The durability and repeatability of the fabricated pepsin-coated column was tested and found to be satisfactory. The on-line digestion of insulin chain-β and lysozyme provides identification of the proteolytic peptides. A 100% recovery was achieved for the insulin chain β-amino acid sequence and 73% for the lysozyme amino acid sequence.
Figure 10.23 Schematic illustration of an on-line enzyme reactor integrated into CE. (a) Substrates are introduced into the pepsincoated PSG column. (b) Substrates are catalyzed into products while they flow through the pepsin-coated PSG monolith by electrophoresis. (c) The products and unreacted substrates were separated in the separation section of the capillary by electrophoresis [139]. Source: Copyright 2004 American Chemical Society.
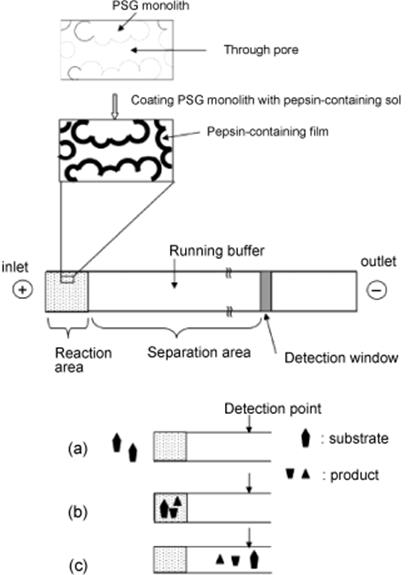
10.3.2 Enzymatic Microreactors for Biocatalysis
Biocatalysis is the use of natural catalysts, such as protein enzymes, to perform chemical transformations on organic compounds. Biocatalysis has become an enabling technology in the synthesis and production of application of biocatalysis to fine chemicals, especially for the pharmaceutical industry. Attractive features of biocatalysts and enzymes include versatility, substrate selectivity, high enantioselectivity, chemoselectivity, regioselectivity, catalysis at ambient temperatures and pressure, and the offer of “green chemistry,” that is, operation usually in an aqueous solvent. Bioprocesses are being explored as a way toward sustainable organic chemical synthesis. One of the criticisms of biocatalytic systems is that they are relatively slow to implement [137]. Considerable efforts which have been inspired by the rapid advances of microfluidic reactors for chemical synthesis and the development of microfluidic enzyme reactors as analytical tools for proteomic studies are now contributing much to speed up the development of bioconversion systems.
10.3.3 Advantages of Microreactors in Biocatalysis
Microscale processing techniques based on microfluidics are rapidly emerging as a means to increase the speed of bioprocess design and reduce material requirements. Many fundamental and practical advantages of microfluidic reactors include more efficient substrate processing owing to more effective thermal transfer due to larger surface area to volume ratios, more precise control over reaction conditions, reduce use of resources, real time monitoring of reaction efficiency, and increase throughput via parallelization [138]. Automation of these techniques can reduce labor intensity and enable a wider range of process variables to be examined. Advantages of microstructured reactors can be categorized according to whether benefits are realized in the development phase or during biocatalytic processing at the production scale. One of the practical benefits during development phase to be considered is the possibility of screening large libraries of biocatalysts in shorter periods of time. These microdevices can be regarded as screening devices that are best used during the stage of biocatalyst selection. They will constitute a useful scale-down system with which a range of realistic process conditions can be examined in a relatively short time. For example, a highly automated and flexible microreactor system can speed up research and development process as it supports high-throughput experimentation. Such system can contribute to optimization of reaction conditions and final selection of the candidate biocatalyst. Continuous processing in microstructured reactors adopted to fit the reaction kinetics of the enzymatic transformation applied can contribute to quality improvement. Design strategy based on microreactors can also offer control of selectivity. Besides the advantage of enabling screening of candidate enzymes from natural biodiversity or mutant libraries, they also provide an excellent system for biocatalytic process analysis and design. The rapid evaluation of different reaction conditions overcomes the time constraints typically associated with biocatalytic process development. Enhanced mass transfer provided by microreactor can be exploited for process intensification, particularly during multiphase biocatalytic processing where mass transfer across phase boundaries is often limiting.
However, for commercial production, the use of continuous microreactor technology should be justified by a clear cost advantage in comparison to presently applied technologies. The use of microfluidic bioreactors opens up possibilities for new production concepts, particularly continuous processing and flexible scale-up on demand via parallelization. There is, however, a need for a modular networks consisting of upstream steps (reaction) and downstream step (separation and purification), that is, the development of whole biocatalytic processes for real implementation in industrial setting.
10.3.4 Biocatalytic Transformations in Microfluidic Systems
10.3.4.1 Solution-phase Enzymatic Reactions
There are various configurations that have been used within the scope of microreactors but the simplest form is solution-phase enzyme reaction, wherein substrate and enzyme aqueous solution are continuously fed separately to the microreactor and the reaction takes place in the microchannel. Although the solution-phase reaction is not a favorable process because of the large volume of enzymes required, several significant achievements have been reported. Table 10.3 presents several examples of biocatalytic synthesis using microreaction technology with free enzymes. Kanno and others reported on on-chip synthesis of oligosaccharides using β-galactosidase in a continuous-flow microreactor as mentioned in Section 10.2.1 [25, 140]. Hydrolysis of p-nitrophenyl-β-d-galactopyranoside was performed in buffered media, whereas transgalactosylation of p-nitrophenyl-2-acetamide-2-deoxy-beta-d-glucopyranoside was conducted in buffer-acetonitrile solvent system. The authors did not use immobilized enzyme. The reaction was performed by separate loading of the enzyme in phosphate buffer solution and the substrate solution in acetonitrile into inlets, and was terminated by heating the recovered solution. Both reactions were enhanced in the microreactor channel compared with the conventional batch reaction [25, 140].
Table 10.3 Enzymatic transformations in microfluidic reactors in solution phase.



Maruyama's group also employed continuous flow in a microfluidic reactor for the enzymatic degradation of p-chlorophenol, which was carried out in a two-phase flow in a microchannel fabricated on a glass plate [141]. The surface of the microchannel was partially modified with octadecylsilane groups to be hydrophobic, thus allowing clear phase separation at the end-junction of the microchannel. The enzyme (laccase), which is surface active, was solubilized in a succinic aqueous buffer and the substrate (p-chlorophenol) was dissolved in isooctane. The degradation of p-chlorophenol occurred mainly at the aqueous-organic interface in the microchannel. A simple theoretical model for the degradation in the microchannel, assuming that diffusion of the substrate is the rate-limiting step in the enzymatic degradation of p-chlorophenol in the microchannel. The calculated degradation values agreed well with the experimental data.
The efficiency of solution-phase (two aqueous phase) enzymatic reaction in microreactor was demonstrated by laccase-catalyzed l-DOPA oxidation in an oxygen-saturated water solution, and analyzed in a Y-shaped microreactor at different residence times (Figure 10.24) [142]. Up to 87% conversions of l-DOPA were achieved at residence times below 2 min. A two-dimensional mathematical model composed of convection, diffusion, and enzyme reaction terms was developed. Enzyme kinetics was described with the double substrate Michaelis–Menten equation, where kinetic parameters from previously performed batch experiments were used. Model simulations, obtained by a nonequidistant finite differences numerical solution of a complex equation system, were proved and verified in a set of experiments performed in a microreactor. Based on the developed model, further microreactor design and process optimization are feasible.
Figure 10.24 Scheme of the microchannel (2W = 220 m, H = 50 m, and L = 332 mm) [142]. Source: Copyright 2009 Elsevier.
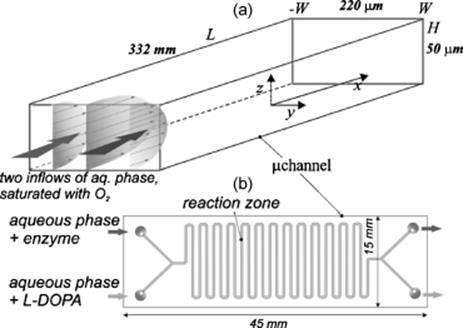
The efficiency of two-phase system was demonstrated by Swarts group who used lipase type B from Candida antarctica to catalyze the esterification of propionic acid and 1-butanol in a water/n-decane two-phase system [143]. The reaction was described by a Ping Pong Bi Bi mechanism with alcohol inhibition. No significant differences were found between the micro and bench scale. Furthermore, effects of temperature on activation and inactivation of the enzyme were found to be similar on micro and bench scale, which provides evidence that parameters found on either scale can be used for the other scale. Enzyme kinetic parameters can be determined on a microscale, with very low consumption of reagents and catalyst, and then be applied to bench scale thus, reducing the cost of optimizing enzyme processes by downscaling.
Another demonstration of a continuous flow operation is the psi-shaped microreactor that was used for lipase-catalyzed synthesis of isoamyl acetate in the 1-butyl-3-methylpyridinium dicyanamide/n-heptane two-phase system [144]. The chosen solvent system with dissolved Candida antarctica lipase B, which was attached to the ionic liquid/n-heptane interfacial area because of its amphiphilic properties, was shown to be highly efficient and enabled simultaneous esterification and product removal. The system allowed for simultaneous esterification and product recovery showed a threefold reaction rate increase when compared to the conventional batch. This was mainly a consequence of efficient reaction-diffusion dynamics in the microchannel system, where the developed flow pattern comprising intense emulsification provided a large interfacial area for the reaction and simultaneous product extraction. Another lipase-catalyzed isoamyl acetate synthesis in a continuously operated pressure-driven microreactor was reported by the same authors [145]. The esterification of isoamyl alcohol and acetic acid occurred at the interface between n-hexane and an aqueous phase with dissolved lipase B from Candida antarctica. Controlling flow rates of both phases reestablished a parallel laminar flow with liquid–liquid boundary in the middle of the microchannel and a separation of phases was achieved at the y-shaped exit of the microreactor (Figure 10.25). The microreactor approach demonstrated 35% conversion at residence time 36.5 s at 45 °C and at 0.5 M acetic acid and isoamyl alcohol inlet concentrations and has proven more effective and outperformed the batch operation, which could be attributed to the favorable mass and heat transfer characteristics.
Figure 10.25 Graphical presentation of the microfluidic device used. (a) Scheme of the main channel, (b) line drawing of the whole microreactor with indicated inlet and outlet constituents [145]. Source: Copyright 2009 Elsevier.
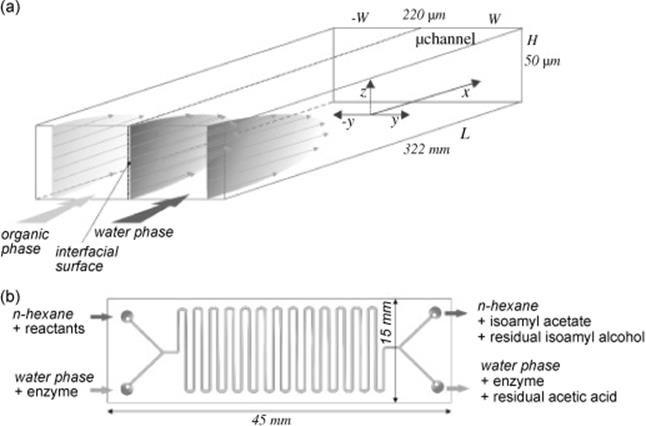
There are studies that take advantage of flow regimes in which two phases are segregated, often also called segmented flow conditions. Segmented or slug flow gives the advantage of higher interfacial area and enhanced mass interfacial transfer over parallel flow [146]. A microreactor based on a segmented flow biphasic system was employed to synthesize optically pure cyanohydrins using a cell lysate containing S-selective hydroxynitrite lyase [147, 148]. Asymmetric synthesis was achieved [148, 149]. Different aldehyde substrates were selected to be converted to their corresponding cyanohydrins (Scheme 10.15).
Scheme 10.15 Biocatalytic synthesis of (S)-cyanohydrins.
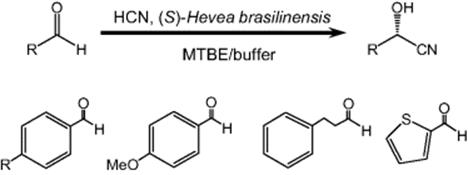
Continuous flow in microfluidic system was also used for the bioluminescent reaction between ATP and luciferin, which was promoted by the enzyme luciferase to ultimately yield oxyluciferin and AMP [149]. The microfluidic technique allowed for the determination of Michaelis–Menten rate constants with a single experiment.
The feasibility of in situ steroid biotransformation and product recovery in microchannels was investigated by Marques and others using a two-phase system comprising an organic phase as substrate and product pool, and an aqueous phase with dissolved enzyme (Scheme 10.15) [150]. The enzymatic oxidation of 4-cholesten-3-one in a Y-shape microreactor geometry (Figure 10.26) achieved roughly up to 70% conversion of cholesterol at residence times below 1 min. A mathematical model based on mass balances for cholesterol, 4-cholesten-3-one, and dissolved oxygen concentrations, comprising double-substrate Michaelis–Menten kinetics and the velocity profile of two immiscible fluids, was developed in order to describe and predict the process of cholesterol oxidation (Figure 10.27).
Figure 10.26 (a) Scheme of microchannel reactor (2W = 220 μm, H = 100 μm, and L = 332 μm). Parallel aqueous–organic two-phase flow (b) and velocity profile (c) at the inlet of the microchannels at combined flow rate of 14 μl/min (4.4 μl/min aqueous phase and 9.6 μl/min organic phase) [150]. Source: Copyright 2010 Elsevier.
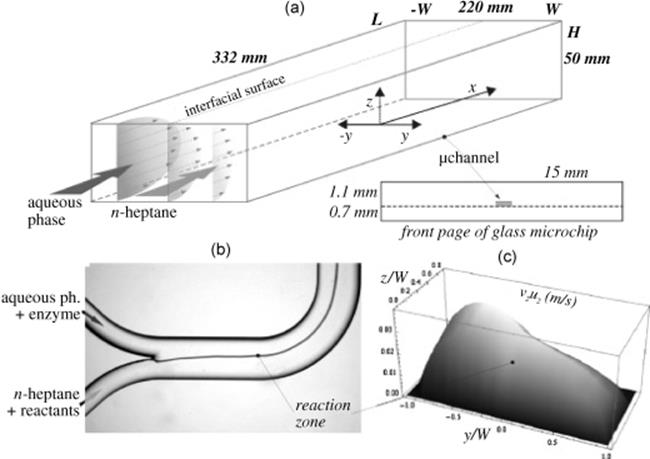
Figure 10.27 (a and b) Microscopic observation of a flow pattern of colored buffer and n-heptane entering through the separate inflow channels (total flow rate through the main microchannel 14 l/min). (c and d) Slug flow formation in the microchannels at equal inlet flow rates of both aqueous and organic phase (total flow rate through the main microchannel 14 l/min) [150]. Source: Copyright 2010 Elsevier.
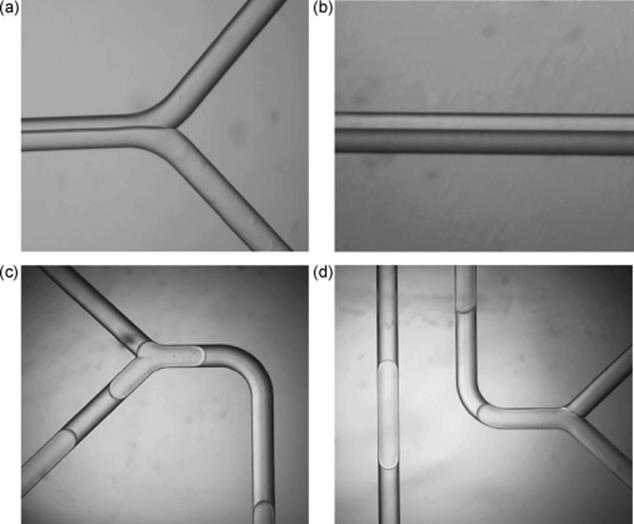

Mohr and his colleagues reported a continuous recirculating two-phase flow miniaturized bioreactor for biocatalytic transformations with the enzyme pentaerythritol tetranitrate reductase (PETN) using on-chip spectroscopic detection of the organic and aqueous phases [151]. A phase separation technique is described that uses electrostatic phase separation, wherein electrostatic attraction forced charged droplets to merge back into the aqueous phase and thus allowed the monitoring of both reaction phases during enzymatic turnover. An enhanced rate of enzyme catalyzed reduction of trans-2-(2-nitrovinyl)thiophene in comparison to batch reactor was reported.
A segmented flow capillary microreactor was used to perform conversion of 1-heptaldehyde to 1-heptanol in a two liquid–liquid phase system by Karande and others [152]. The enzyme used was the thermophilic alcohol dehydrogenase (TADH). Aqueous and organic liquids were pumped and segmented flow was attained through a T-piece connector and introduced into the PTFE tubing. The system showed high productivities and easy phase separation facilitating downstream processing.
Most recently, a modular microfluidic reactor and in-line filtration system for the rapid and small-scale evaluation of biocatalytic reactions have been demonstrated by O'Sullivan and others [153]. The system combined a substrate with a biocatalyst in free solution. The PMMA enzymatic microreactor worked by co-flowing the enzyme and substrate through a T-channel and mixing was achieved by staggered herringbone micromixer (SHM). The filtration unit composed of gaskets made of PDMS was integrated into the enzyme mixer. This platform was used for separation of biocatalyst for transketolase-catalyzed coupling of irreversible donor HPA to the acceptors glycoaldehyde and propanol. Complete or near complete conversions into (3S)-1,3-dihydroxypentan-2-one in microreactor comparable to batch reactors were obtained.
The reports mentioned above provide a systematic coverage of the nonimmobilized enzymatic reactors used in biocatalytic reactions under continuous flow operation. Results from microreactor experiments were comparatively higher than conventionally mixed batch reactors in terms of conversion rate and improvement of product yield as demonstrated for hydrolysis [140], dehalogenation [141], oxidation [142], esterification [143], synthesis of isoamyl acetate [144, 145], synthesis of cyanohydrins [147, 148], synthesis of chiral metabolites [153], reduction [151], and bioluminescent reaction [149]. The small volumes involved and the favorable mass transfer inherent to these devices make them particularly useful for the screening of biocatalysts and rapid characterization of bioconversion systems. The remarkable results of such studies revealed that the product yield could be enhanced significantly in comparison with the conventional batch runs.
10.3.4.2 Microfluidic Reactors with Immobilized Enzymes for Biocatalytic Transformations
Another category of enzymatic transformations in multiphase systems is enzymes immobilized on the reactor wall as presented in Table 10.4. Enzymes are advantageously used in immobilized form because this strategy allows for increased volumetric productivity and improves stability. Continuous mode of operation is employed in these systems. The approaches commonly used for immobilization in conventional multiphase biocatalysis can also be employed in microreactors such as covalent methods, cross-linked enzyme aggregates (CLEA), and adsorption methods. The experimental setups can either be chip-type reactors with activated channel surface walls where enzyme binds, or enzyme immobilized monolith reactors, where a support is packed inside a capillary tube.
Table 10.4 Enzymatic transformations in microfluidic reactors with immobilized enzymes.




Our group has developed a technique that forms an enzyme-immobilizing membrane on the microchannel surface [154]. This is a modification of CLEA formation, which is used in batchwise organic synthesis [155]. The microreactor has a cylindrical membrane composed of cross-linked polymerized acylase and polylysine matrix product on the internal surface of the microchannel (Figure 10.28). This microreactor, called CLEA-based enzyme microreactor (CEM), prevents high pressure better than carrier-filled microreactors because of its hollow structure [121]. This microreactor enabled a highly enantioselective reaction for a racemic amino acid derivative. The integration of enzyme microreactor (CLEA-CEM) and microextractor provided efficient continuous production of optically pure amino acids.
Figure 10.28 Continuous flow system for enantioselective separation of racemic amino acids. First, an L-body in the racemic substrate (4 mM) is enantioselectively converted into an amino acid through the tubing acylase-reactor. The pH of the resultant substrate solution is then lowered by addition of 0.2 M HCl that is supplied to the microextractor with ethyl acetate. Acetyl-d-Phe in the acidic aqueous phase is extracted selectively to the ethyl acetate phase flowing along the silicon-side in a microchannel [154]. Source: Copyright 2007 Royal Society of Chemistry.
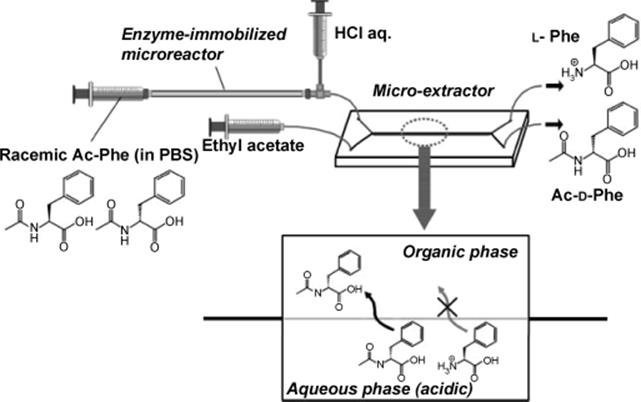
Enzyme immobilization by CLEA was also used by Hickey and others [156]. The aminoacylase enzyme was successfully immobilized in miniaturized flow reactors. Two approaches to enzyme immobilization have been examined, both involving enzyme cross-linking. The first reactor type has used monoliths, to which the enzyme was attached, and the second contained previously cross-linked enzyme trapped using frits, in the microfluidic channels. Two different microreactor designs were used in the investigation: microreactor chips for the monoliths and capillary flow reactors for the cross-linked enzyme. These systems allowed passage of the substrate and product through the system while retaining the aminoacylase enzyme performing the catalytic conversion. The enzyme has been successfully immobilized and used to produce stable biocatalytic microreactors that can be used repeatedly over a period of several months.
Applications of enzyme-immobilized microreactors for processing have also been presented including hydroxylation of macrolide in a microreactor [156]. PikC hydroxylase was immobilized on Ni-NTA agarose beads via in situattachment. The enzyme loading was ~6 μg/mg of beads resulting in a microchannel loading of 10.7 mg/ml. This high enzyme loading enabled the rapid hydroxylation of the macrolide YC-17 to methymycin and neomethymycin in about equal amounts with a conversion of >90% at a flow rate of 70 nl/min. This high reactivity allowed rapid hydroxylation reactions to be performed with short residence times, which is critical for complex enzymes with limited inherent stability. A similar immobilization technique was applied for the synthesis of (R)-benzoin, (R)-2-hydroxy-1-phenylpropan-1-one, and 6-O-acetyl-d-glucal in a flow-through mode. His-tagged protein was directly immobilized within the microstructured PASSflow reaction system through tyrosine-based Ni-nitrilotriacetic acid (Ni-NTA) system [158].
Biocatalytic reactions performed using immobilized enzyme microreactors under continuous flow mode have been found effective for hydrolysis reactions [121, 158–161], with the enzyme either trapped in the matrix [160], covalently linked to modified surface wall [160, 121], enzymes entrapped in hydrogels [162], or enzymes immobilized on monolith [179]. The experimental setup consists of either chip-type microreactors with activated channel walls where enzymes bind, enzymes that bind to beads, enzymes entrapped in the matrix, enzymes adsorbed in nanoporous materials, and most recently, nanosprings as supports for immobilized enzymes in chip-based reactors, or enzyme immobilized monolith reactors, where support is packed inside a capillary tube (Table 10.4).