THE LIVING WORLD
Unit Three. The Continuity of Life
12.5. How Prokaryotes Control Transcription
Being able to translate a gene into a polypeptide is only part of gene expression. Every cell must also be able to regulate when particular genes are used. Imagine if every instrument in a symphony played at full volume all the time, all the horns blowing full blast and each drum beating as fast and loudly as it could! No symphony plays that way because music is more than noise—it is the controlled expression of sound. In the same way, growth and development are due to the controlled expression of genes, each brought into play at the proper moment to achieve precise and delicate effects.
Control of gene expression is accomplished very differently in prokaryotes than in the cells of complex multicellular organisms. Prokaryotic cells have been shaped by evolution to grow and divide as rapidly as possible, enabling them to exploit transient resources. Proteins in prokaryotes turn over rapidly. This allows them to respond quickly to changes in their external environment by changing patterns of gene expression. In prokaryotes, the primary function of gene control is to adjust the cell’s activities to its immediate environment. Changes in gene expression alter which enzymes are present in the cell in response to the quantity and type of available nutrients and the amount of oxygen present. Almost all of these changes are fully reversible, allowing the cell to adjust its enzyme levels up or down as the environment changes.
How Prokaryotes Turn Genes Off and On
Prokaryotes control the expression of their genes largely by saying when individual genes are to be transcribed. At the beginning of each gene are special regulatory sites that act as points of control. Specific regulatory proteins within the cell bind to these sites, turning transcription of the gene off or on.
For a gene to be transcribed, the RNA polymerase has to bind to a promoter, a specific sequence of nucleotides on the DNA that signals the beginning of a gene. In prokaryotes, gene expression is controlled by either blocking or allowing the RNA polymerase access to the promoter. Genes can be turned off by the binding of a repressor, a protein that binds to the DNA blocking the promoter. Genes can be turned on by the binding of an activator, a protein that makes the promoter more accessible to the RNA polymerase.
Many genes are “negatively” controlled: They are turned off except when needed. In these genes, the regulatory site is located between the place where the RNA polymerase binds to the DNA (the promoter site) and the beginning edge of the gene. When a regulatory protein called a repressor is bound to its regulatory site, called the operator, its presence blocks the movement of the polymerase toward the gene. Imagine if you went to sit down to eat dinner and someone was already sitting in your chair—you could not begin your meal until this person was removed from your chair. In the same way, the polymerase cannot begin transcribing the gene until the repressor protein is removed.
To turn on a gene whose transcription is blocked by a repressor, all that is required is to remove the repressor. Cells do this by binding special “signal” molecules to the repressor protein; the binding causes the repressor protein to contort into a shape that doesn’t fit DNA, and it falls off, removing the barrier to transcription. A specific example demonstrating how repressor proteins work is the set of genes called the lac operon in the bacterium Escherichia coli. An operon is a segment of DNA containing a cluster of genes that are transcribed as a unit. The lac operon, shown in figure 12.12, consists of both polypeptide-encoding genes (labeled genes 1, 2, and 3, which code for enzymes involved in breaking down the sugar lactose) and associated regulatory elements—the operator (the purple segment) and promoter (the orange segment). Transcription is turned off when a repressor molecule binds to the operator such that RNA polymerase cannot bind to the promoter. When E. coli encounters the sugar lactose, a metabolite of lactose called allolactose binds to the repressor protein and induces a twist in its shape that causes it to fall from the DNA. As you can see in figure 12.12h, RNA polymerase is no longer blocked, so it starts to transcribe the genes needed to break down the lactose to get energy.
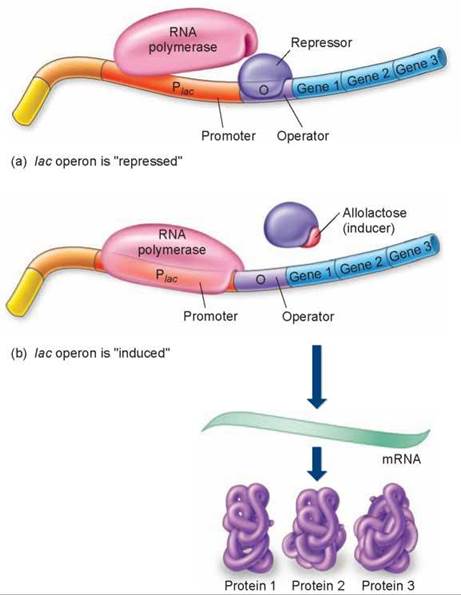
Figure 12.12. How the lac operon works.
(a) The lac operon is shut down ("repressed") when the repressor protein is bound to the operator site. Because promoter and operator sites overlap, RNA polymerase and the repressor cannot bind at the same time. (b) The lac operon is transcribed ("induced") when allolactose binds to the repressor protein changing its shape so that it can no longer sit on the operator site and block polymerase binding.
Because RNA polymerase binds to a specific promoter site on one strand of the DNA double helix, it is necessary that the DNA double helix unzip in the vicinity of this site for the polymerase protein to be able to sit down properly. In many genes, this unzipping cannot take place without the assistance of a regulatory protein called an activator that binds to the DNA in this region and helps it unwind. Just as in the case of the repressor protein described previously, cells can turn genes on and off by binding “signal” molecules to the activator protein. These molecules either prevent the activator from binding to the DNA or enable it to do so. In the lac operon, a protein called catabolite activator protein (CAP) acts as an activator. CAP has to bind a signal molecule, cAMP, before it can associate with the DNA. Once the CAP/cAMP complex forms, as shown in figure 12.13, it binds to the DNA and makes the promoter more accessible to RNA polymerase.
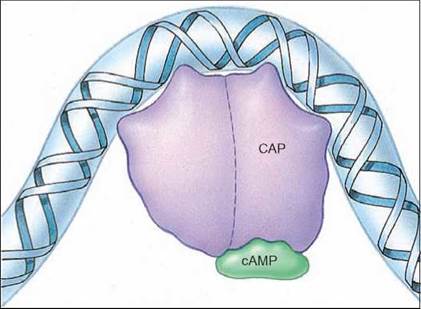
Figure 12.13. How an activator works.
Binding of the catabolite activator protein (CAP)/cAMP complex to DNA causes the DNA to bend around it. This increases the activity of RNA polymerase.
Why bother with activators? Imagine if you had to eat every time you encountered food! Activator proteins enable a cell to cope with this sort of problem. Activators and repressors work together to control transcription. To understand how, let’s consider the lac operon again, now shown in figure 12.14. When a bacterium encounters the sugar lactose, it may already have lots of energy in the form of glucose, as shown in panel 1, and so does not need to break down more lactose. CAP can only bind and activate gene transcription when glucose levels are low. Because RNA polymerase requires the activator to function, the lac operon is not expressed. Also if glucose is present and lactose is absent, not only is the activator CAP unable to bind, but also a repressor blocks the promoter, as shown in panel 2 below and in figure 12.12a. In the absence of both glucose and lactose, cAMP, the “low glucose” signal molecule (the green pie-shaped piece in panels 3 and 4) binds to CAP, and CAP is able to bind to the DNA. However, the repressor is still blocking transcription, as shown in panel 3. Only in the absence of glucose and in the presence of lactose, the repressor is removed, the activator (CAP) is bound, and transcription proceeds, as shown in panel 4.
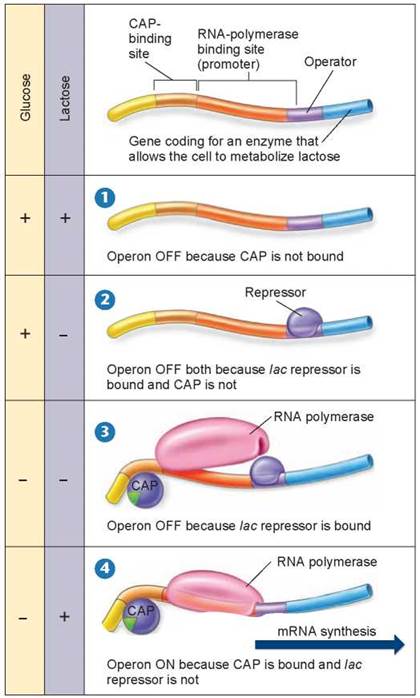
Figure 12.14. Activators and repressors at the lac operon.
Key Learning Outcome 12.5. Cells control the expression of genes by determining when they are transcribed. Some regulatory proteins block the binding of RNA polymerase, and others facilitate it.