5 Steps to a 5: AP Biology - Mark Anestis 2021
STEP 4 Review the Knowledge You Need to Score High
CHAPTER 10 Molecular Genetics
Exam Weight: 12—16%
IN THIS CHAPTER
Summary: This chapter describes the various processes in cells that take DNA from gene to protein: replication, transcription, posttranscriptional modification, and translation. It also discusses the regulation of these processes before concluding with a discussion about viruses, bacteria, and genetic engineering.

Key Ideas
![]() DNA: adenine-thymine, cytosine-guanine—arranged in a double helix.
DNA: adenine-thymine, cytosine-guanine—arranged in a double helix.
![]() RNA: adenine-uracil, cytosine-guanine—single stranded.
RNA: adenine-uracil, cytosine-guanine—single stranded.
![]() DNA replication occurs during the S-phase in a semi-conservative fashion and in a 5’ to 3’ direction.
DNA replication occurs during the S-phase in a semi-conservative fashion and in a 5’ to 3’ direction.
![]() Types of DNA replication mutations: frameshift, missense, nonsense.
Types of DNA replication mutations: frameshift, missense, nonsense.
![]() Transcription: mRNA is formed from a DNA template.
Transcription: mRNA is formed from a DNA template.
![]() Translation: process by which mRNA specified sequence of amino acids is lined up on a ribosome for protein synthesis.
Translation: process by which mRNA specified sequence of amino acids is lined up on a ribosome for protein synthesis.
![]() Operons act as on-off switches for transcription—allow for production of genes only when needed.
Operons act as on-off switches for transcription—allow for production of genes only when needed.
![]() Types of genetic recombination: transformation, transduction, and conjugation.
Types of genetic recombination: transformation, transduction, and conjugation.
Introduction
Genetics has implications for all of biology. We begin our study of this subject with an introduction to DNA and RNA, followed by a description of the various processes in cells that take DNA from gene to protein: replication, transcription, posttranscriptional modification, translation, and the regulation of all these processes. The genetics of viruses and bacteria follows, and the chapter concludes with a discussion of genetic engineering.
The Central Dogma
The central dogma (Figure 10.1 is a phrase coined by Francis Crick in 1957 to describe the flow of genetic information through a biological system. It consists of DNA ➔ RNA ➔ Proteins and the processes involved in making this flow of information possible. Those processes are replication, transcription, and translation.
IST-1
Heritable information provides for continuity of life.
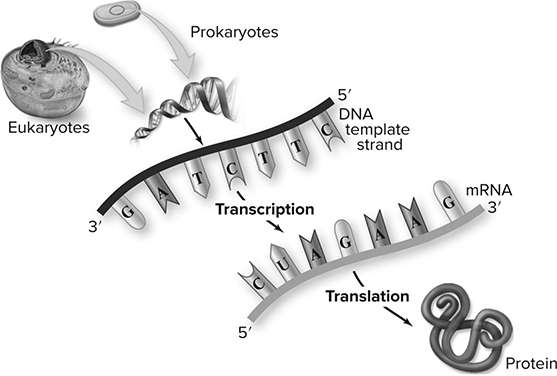
Figure 10.1 The Central Dogma. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
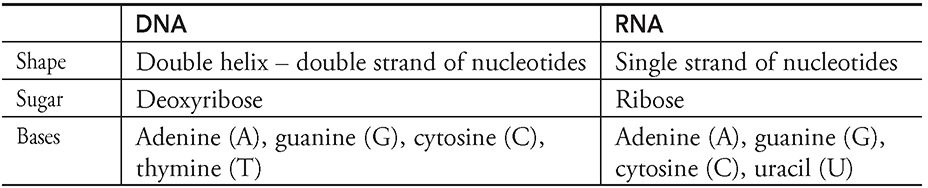
First, take a second to review the structure of DNA and RNA that is located in Chapter 5. Remember DNA is a double helix that is made up of two strands of nucleotides held together by hydrogen bonds while RNA is a single strand of nucleotides. Take a look at the following table:
Replication of DNA
Human cells do not have copy machines to do the dirty work for them. Instead, they use a system called DNA replication to copy DNA molecules from cell to cell. As we discussed in Chapter 8, this process occurs during the S-phase of the cell cycle to ensure that every cell produced during mitosis or meiosis receives the proper amount of DNA.
The mechanism for DNA replication was the source of much debate in the mid-1900s. Some argued that it occurred in what was called a “conservative” (conservative DNA replication) fashion. In this model, the original double helix of DNA does not change at all; it is as if the DNA is placed on a copy machine and an exact duplicate is made. DNA from the parent appears in only one of the two daughter cells. A different model called the semiconservative DNA replication model agrees that the original DNA molecule serves as the template but proposes that before it is copied, the DNA unzips, with each single strand (Figure 10.2 serving as a template for the creation of a new double strand. One strand of DNA from the parent goes to one daughter cell, and the second parent strand to the second daughter cell. A third model, the dispersive DNA replication model, suggested that every daughter strand contains some parental DNA, but it is dispersed among pieces of DNA not of parental origin. Figure 10.2 is a simplistic sketch showing these three main theories. Watson and Crick would not be pleased to see that we did not draw the DNA as a double helix . . . but as long as you realize this is not how the DNA truly looks, the figure serves its purpose.
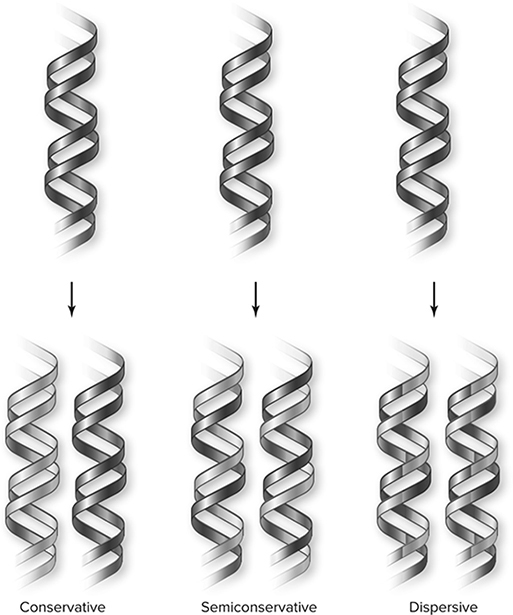
Figure 10.2 Three DNA replication models. The conservative model produces one entirely new molecule and conserves the old. The semiconservative model produces two hybrid molecules of old and new strands. The dispersive model produces hybrid molecules with each strand a mixture of old and new. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
An experiment performed in the 1950s by Meselson and Stahl (Figure 10.3 helped select a winner in the debate about replication mechanisms. The experimenters grew bacteria in a medium containing 15N (a heavier-than-normal form of nitrogen) to create DNA that was denser than normal. The DNA was denser because the bacteria picked up the 15N and incorporated it into their DNA. The bacteria were then transferred to a medium containing normal 14N nitrogen. The DNA was allowed to replicate and produced DNA that was half 15N and half 14N. When the first generation of offspring replicated to form the second generation of offspring, the new DNA produced was of two types—one type that had half 15N and half 14N, and another type that was completely 14N DNA. This gave a hands-down victory to the semi-conservative theory of DNA replication. Let’s take a look at the mechanism of semi-conservative DNA replication.
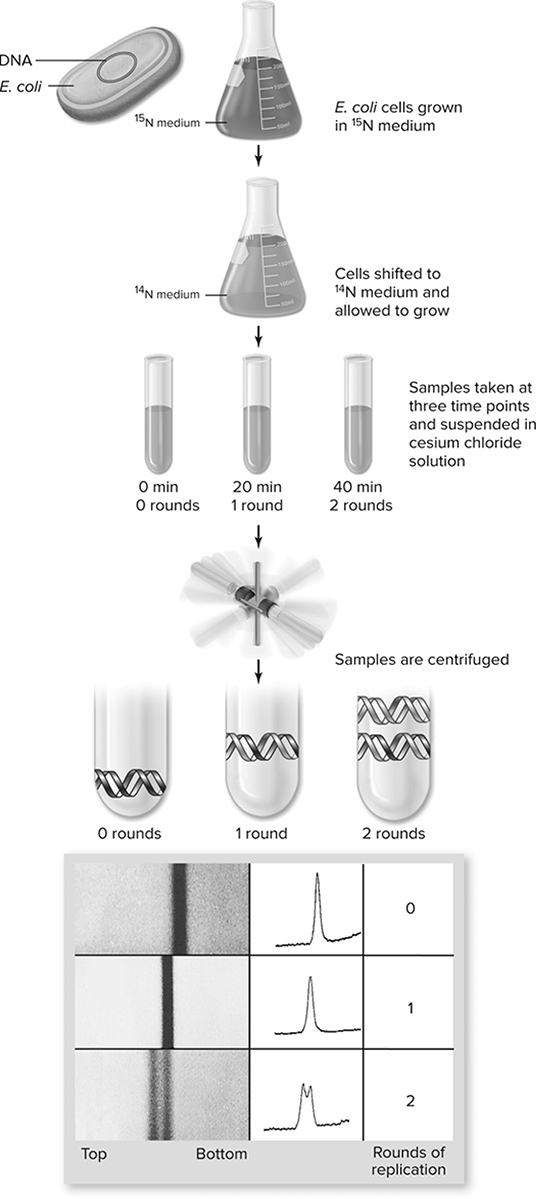
Figure 10.3 The Meselson—Stahl experiment. Bacteria grown in heavy 15N medium are shifted to light 14N medium and grown for two rounds of replication. Samples are taken at time points corresponding to zero, one, and two rounds of replication and centrifuged in cesium chloride to form a gradient. The actual data are shown at the bottom with the interpretation of semiconservative replication shown schematically (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
During the S-phase of the cell cycle, the double-stranded DNA unzips and prepares to replicate. An enzyme called helicase unzips the DNA just like a jacket, breaking the hydrogen bonds between the nucleotides and producing the replication fork (Figure 10.4. Each strand then functions as a template for production of a new double-stranded DNA molecule. Specific regions along each DNA strand serve as primer sites that signal where replication should originate. Primase binds to the primer, and DNA polymerase, the superstar enzyme of this process, attaches to the primer region and adds nucleotides to the growing DNA chain in a 5′-to-3′ direction. DNA polymerase is restricted in that it can only add nucleotides to the 3′ end of a parent strand. This creates a problem because this means that only one of the strands can be produced in a continuous fashion.
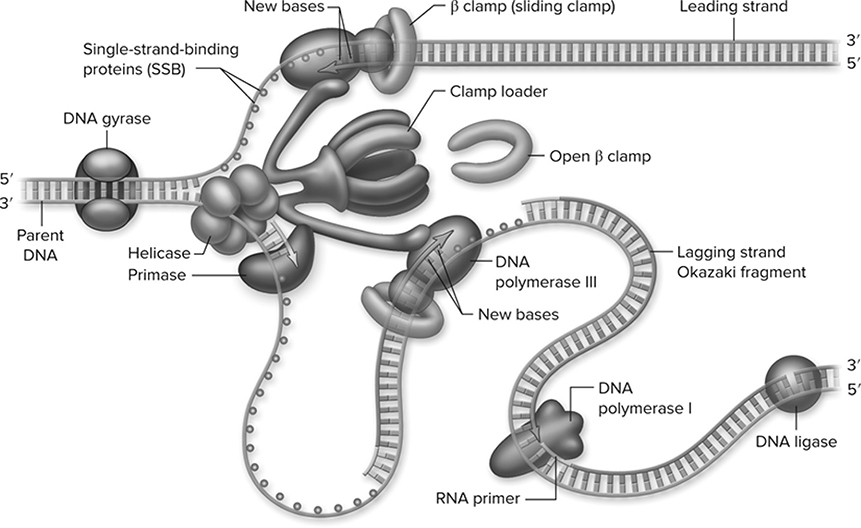
Figure 10.4 The replication fork. A model for the structure of the replication fork with two polymerase III enzymes held together by a large complex of accessory proteins. These include the “clamp loader,” which loads the β subunit sliding clamp periodically on the lagging strand. The polymerase III on the lagging strand periodically releases its template and reassociates along with the β clamp. The loop in the lagging-strand template allows both polymerases to move in the same direction despite DNA being antiparallel. Primase, which makes primers for the lagging-strand fragments, and helicase are also associated with the central complex. Polymerase I removes primers and ligase joins the fragments together. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
This continuous strand is known as the leading strand. The other strand is affectionately known as the lagging strand. You will notice that in the third step of the process in Figure 10.5, the lagging strand consists of tiny pieces called Okazaki fragments, which are later connected by an enzyme called DNA ligase to produce the completed double-stranded daughter DNA molecule. This is the semi-conservative model of DNA replication.
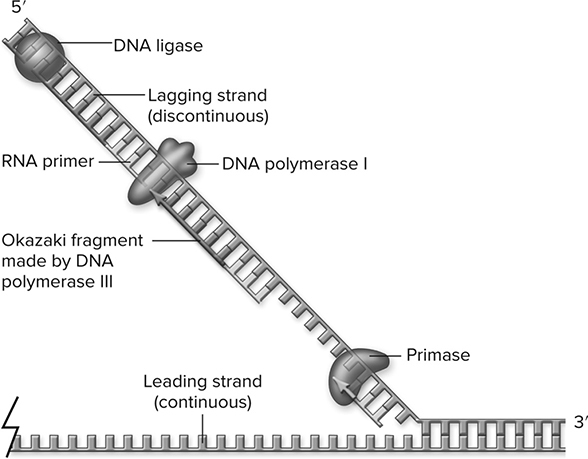
Figure 10.5 Lagging-strand synthesis. The action of primase synthesizes the primers needed by DNA polymerase III (not shown). These primers are removed by DNA polymerase I using its 5′-to-3′ exonuclease activity, then extending the previous Okazaki fragment to replace the RNA. The nick between Okazaki fragments after primer removal is sealed by DNA ligase. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
DNA replication is not a perfect process—mistakes are made. A series of proofreading enzymes function to make sure that the DNA is properly replicated each time. During the first run-through, it is estimated that a nucleotide mismatch is made during replication in 1 out of every 10,000 basepairs. The proofreaders must do a pretty good job since a mismatch error in replication occurs in only one out of every billion nucleotides replicated. DNA polymerase proofreads the newly added base right after it is added on to make sure that it is the correct match. Repair is easy—the polymerase simply removes the incorrect nucleotide, and adds the proper one in its place. This process is known as mismatch repair. Another repair mechanism is excision repair, in which a section of DNA containing an error is cut out and the gap is filled in by DNA polymerase. There are other proteins that assist in the repair process, but their identities are not of major importance. Just be aware that DNA repair exists and is a very efficient process.
Key Enzymes to Know for DNA Replication
Helicase: enzyme that unzips DNA, breaking the hydrogen bonds between the nucleotides and producing the replication fork for replication
Topoisomerase: enzyme that helps in the unwinding or rewinding of DNA.
DNA polymerase: the main enzyme in DNA replication that attaches to primer proteins and adds nucleotides to the growing DNA chain in a 5′-to-3′ chain.
Ligase: enzyme that connects two strands of DNA together by forming a bond between the phosphate group of one strand and the deoxyribose group of another.
RNA polymerase: enzyme that runs transcription and adds the appropriate nucleotides to the 3′ end of the growing strand.
Telomeres
In eukaryotic chromosomes, DNA replication (Figure 10.6 creates an issue for the structure of the chromosome. The leading strand is completely replicated, but the lagging strand is not able to be completed all the way to the end. This occurs because the primer used to start DNA replication on the lagging strand is not replaced. As a result, during each round of replication, the lagging strand template would produce a shorter chromosome (Figure 10.7. This is where telomeres come into play. Telomeres are specialized structures composed of short repeated sequences of DNA that are made by telomerase. They are found on the ends of eukaryotic chromosomes that protect the integrity and length of the chromosomes after each replication.
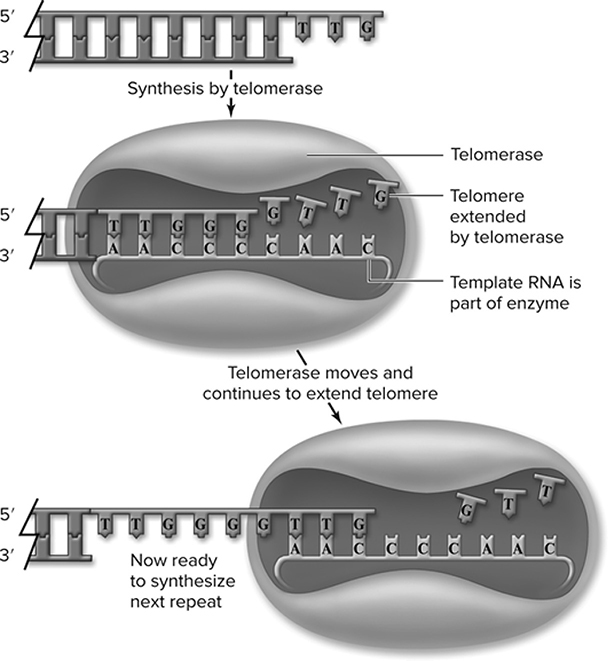
Figure 10.6 Action of telomerase. Telomerase contains an internal RNA that the enzyme uses as a template to extend the DNA of the chromosome end. Multiple rounds of synthesis by telomerase produce repeated sequences. This single strand is completed by normal synthesis using it as a template (not shown). (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
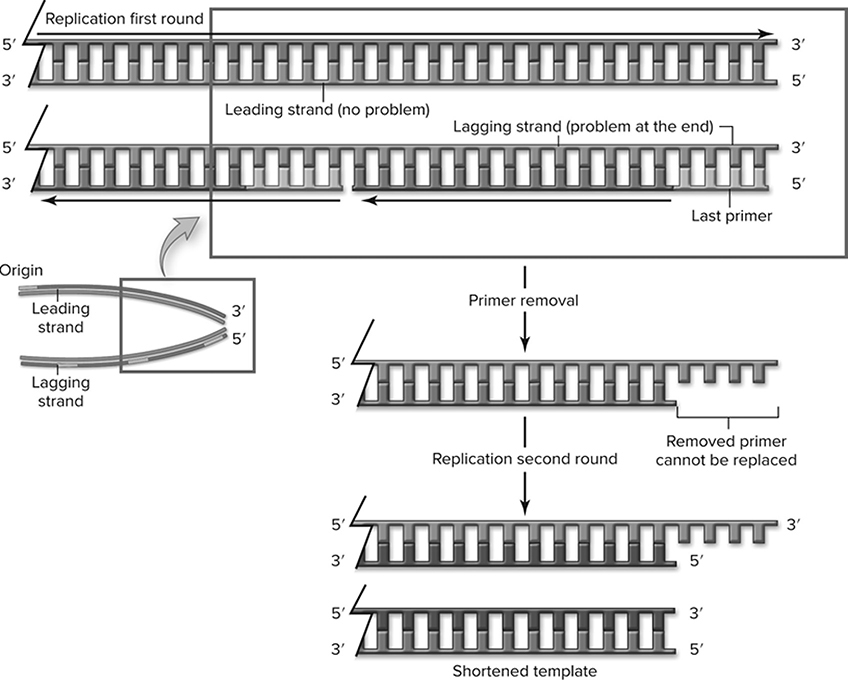
Figure 10.7 Replication of the end of linear DNA. Only one end is shown for simplicity; the problem exists at both ends. The leading strand can be completely replicated, but the lagging strand cannot be -finished. When the last primer is removed, it cannot be replaced. During the next round of replication, when this shortened template is replicated, it will produce a shorter chromosome. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
Transcription of DNA
Up until this point, we have just been discussing DNA replication, which is simply the production of more DNA. In the rest of the chapter, we discuss transcription, translation, and other processes involving DNA. While DNA is the hereditary material responsible for the passage of traits from generation to generation, DNA does not directly produce the proteins that it encodes. DNA must first be transcribed into an intermediary: mRNA. This process is called transcription (Figure 10.8 because both DNA and RNA are built from nucleotides—they speak a similar language. DNA acts as a template for mRNA, which then conveys to the ribosomes the blueprints for producing the protein of interest. Transcription occurs in the nucleus.
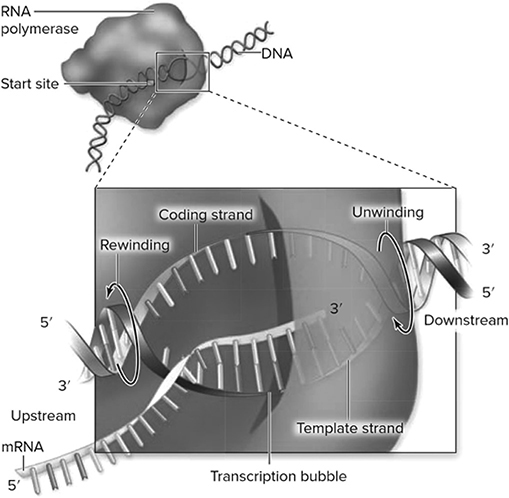
Figure 10.8 Model of a transcription bubble. The DNA duplex is unwound by the RNA polymerase complex, rewinding at the end of the bubble. One of the strands of DNA functions as a template, and nucleotide building blocks are added to the 3′ end of the growing RNA. There is a short region of RNA—DNA hybrid within the bubble. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
NY teacher: “Know the basic principles. They’ll ask you about this process.”
Transcription consists of three steps: initiation, elongation, and termination. The -process begins when RNA polymerase attaches to the promoter region of a DNA strand (initiation). A promoter region is simply a recognition site that shows the polymerase where transcription should begin. The promoter region contains a group of nucleotides known as the TATA box, which is important to the binding of RNA polymerase. As in DNA replication, the polymerase of transcription needs the assistance of helper proteins to find and attach to the promoter region. These helpers are called transcription factors (Figure 10.9. Once bound, the RNA polymerase works its magic by adding the appropriate RNA nucleotide to the 3′ end of the growing strand (elongation). Like DNA polymerase of replication, RNA polymerase adds nucleotides 5′ to 3′. The growing mRNA strand separates from the DNA as it grows longer. A region called the termination site tells the polymerase when transcription should conclude (termination). After reaching this site, the mRNA is released and set free.
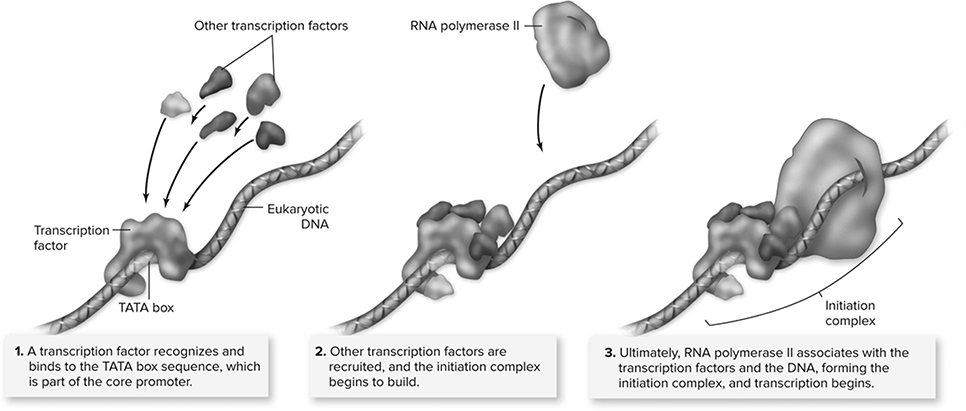
Figure 10.9 Eukaryotic initiation complex. Unlike transcription in prokaryotic cells, in which the RNA polymerase recognizes and binds to the promoter, eukaryotic transcription requires the binding of transcription factors to the promoter before RNA polymerase II binds to the DNA. The association of transcription factors and RNA polymerase II at the promoter is called the initiation complex. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
Polyribosomes
The central dogma differs for eukaryotes and prokaryotes due to the presence of a nucleus in eukaryotes. For prokaryotes that lack a nucleus, transcription is coupled to translation (Figure 10.10. Once the mRNA is produced by transcription, then translation begins before transcription is finished.
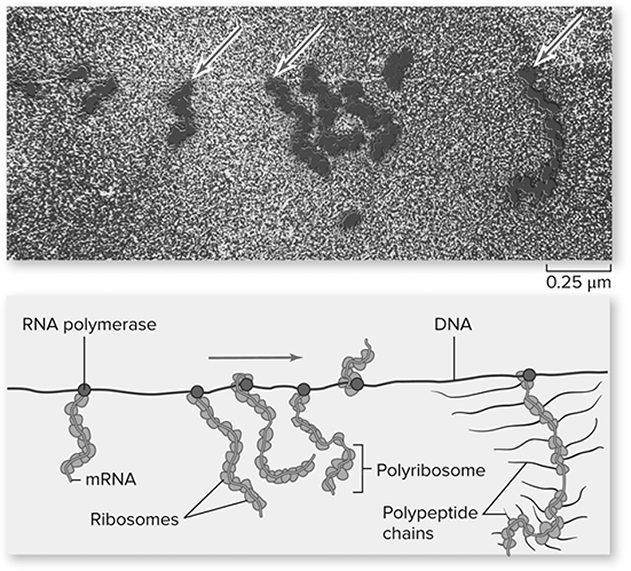
Figure 10.10 Transcription and translation are coupled in prokaryotes. In this micrograph of gene expression in E. coli, translation is occurring during transcription. The arrows point to RNA polymerase enzymes, and ribosomes are attached to the mRNAs extending from the polymerase. Polypeptides being synthesized by ribosomes, which are not visible in the micrograph, have been added to the last mRNA in the drawing. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
RNA Processing
In bacteria, mRNA is ready to rock immediately after it is released from the DNA. In eukaryotes, this is not the case. The mRNA produced after transcription must be modified before it can leave the nucleus and lead the formation of proteins on the ribosomes. The 5′ and the 3′ ends of the newly produced mRNA molecule are touched up (Figure 10.11. The 5′ end is given a guanine cap, which serves to protect the RNA and also helps in attachment to the ribosome later on. The 3′ end is given something called a polyadenine tail, which may help ease the movement from the nucleus to the cytoplasm.
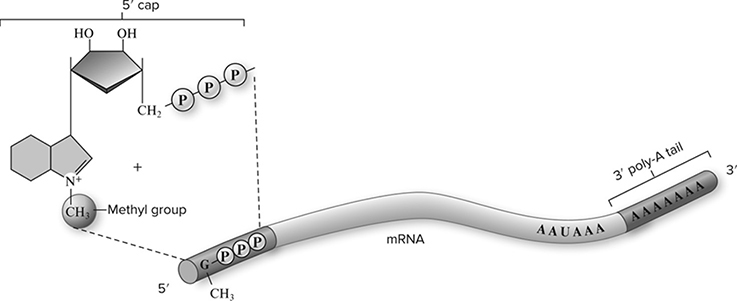
Figure 10.11 Posttranscriptional modifications to 5′ and 3′ ends. Eukaryotic mRNA molecules are modified in the nucleus with the addition of a methylated GTP to the 5′ end of the transcript, called the 5′ cap, and a long chain of adenine residues to the 3′ end of the transcript, called the 3′ poly-A tail. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
Along with these changes, the introns (noncoding regions produced during transcription) are cut out of the mRNA, and the remaining exons (coding regions) are glued back together to produce the mRNA that is translated into a protein (Figure 10.12. This is called RNA splicing. We admit that it does seem strange and inefficient that the DNA would contain so many regions that are not used in the production of the gene, but perhaps there is a method to the madness. It is hypothesized that introns exist to provide flexibility to the genome. They could allow an organism to make different proteins from the same gene; the only difference is which introns get spliced out from one to the other. It is also possible that this whole splicing process plays a role in allowing the movement of mRNA from the nucleus to the cytoplasm.
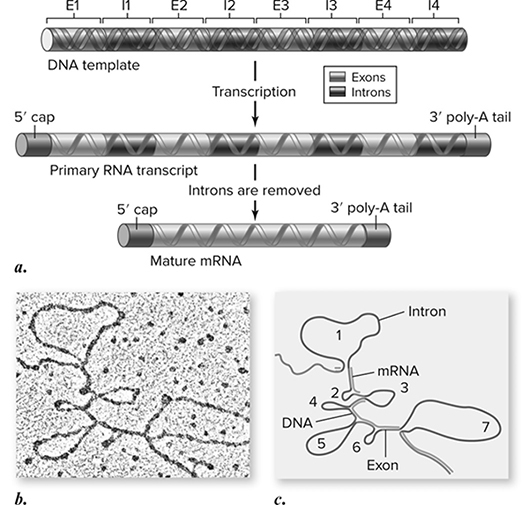
Figure 10.12 Eukaryotic genes contain introns and exons. a. Eukaryotic genes contain sequences that form the coding sequence called exons and intervening sequences called introns. b. An electron micrograph showing hybrids formed with the mRNA and the DNA of the ovalbumin gene, which has seven introns. Introns within the DNA sequence have no corresponding sequence in the mRNA and thus appear as seven loops. c. A schematic drawing of the micrograph. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
Translation of RNA
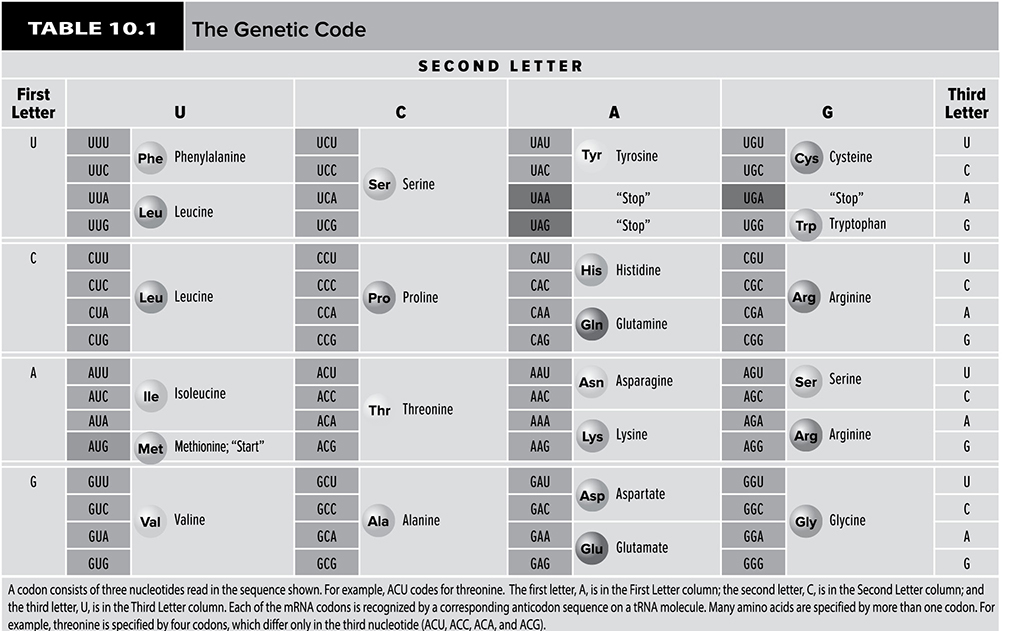
Now that the mRNA has escaped from the nucleus, it is ready to help direct the construction of proteins. This process occurs in the cytoplasm, and the site of protein synthesis is the ribosome. As mentioned in Chapter 5, proteins are made of amino acids. Each protein has a distinct and particular amino acid order. Therefore, there must be some system used by the cell to convert the sequences of nucleotides that make up an mRNA molecule into the sequence of amino acids that make up a particular protein. The cell carries out this conversion from nucleotides to amino acids through the use of the genetic code. An mRNA molecule is divided into a series of codons that make up the code. Each codon is a triplet of nucleotides that codes for a particular amino acid. There are 20 different amino acids, and 64 different combinations of codons. This means that some amino acids are coded for by more than one codon. For example, the codons GCU, GCC, GCA, and GCG all call for the addition of the amino acid alanine during protein creation. Of these 64 possibilities, one is a start codon, AUG, which establishes the reading frame for protein formation. Also among these 64 codons (Table 10.1) are three stop codons: UGA, UAA, and UAG. When the protein formation machinery hits these codons, the production of a protein stops.
Before we go through the steps of protein synthesis, we would like to introduce to you the other players involved in the process. We have already spoken about mRNA, but we should meet the host of the entire shindig, the ribosomes, which are made up of a large and a small subunit. A huge percentage of a ribosome is built out of the second type of RNA mentioned earlier, rRNA. Two other important parts of a ribosome that we will discuss in more detail later are the A site and the P site, which are tRNA attachment sites. The job of tRNA is to carry amino acids to the ribosomes. The mRNA molecule that is involved in the formation of a protein consists of a series of codons. Each tRNA has, at its attachment site, a region called the anticodon, which is a three-nucleotide sequence that is perfectly complementary to a particular codon. For example, a codon that is AUU has an anticodon that reads UAA in the same direction. Each tRNA molecule carries an amino acid that is coded for by the codon that its anticodon matches up with. Once the tRNA’s amino acid has been incorporated into the growing protein, the tRNA leaves the site to pick up another amino acid just in case its services are needed again at the ribosome. An enzyme known as aminoacyl tRNA synthetase (Figure 10.13 makes sure that each tRNA molecule picks up the appropriate amino acid for its anticodon.
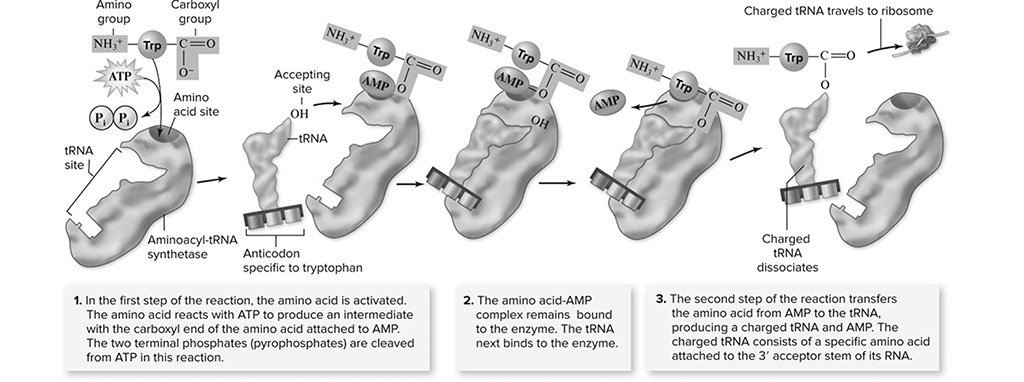
Figure 10.13 The Binding of amino acids to specific tRNA molecules. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
Uh-oh . . . there is a potential problem here. There are fewer than 50 different types of tRNA molecules. But there are more codons than that. Oh, dear . . . but wait! This is not a problem because some tRNA are able to match with more than one codon. How can this be? This works thanks to a phenomenon known as wobble, where a uracil in the third position of an anticodon can pair with A or G instead of just A. There are some tRNA molecules that have an altered form of adenine, called inosine (I), in the third position of the anticodon. This nitrogenous base is able to bind with U, C, or A. Wobble allows the 45 tRNA molecules to service all the different types of codons seen in mRNA molecules.
We have met all the important players in the translation process (see also Figure 10.14), which begins when an mRNA attaches to a small ribosomal subunit. The first codon for this process is always AUG. This attracts a tRNA molecule carrying methionine to attach to the AUG codon. When this occurs, the large subunit of the ribosome, containing the A site and the P site, binds to the complex. The elongation of the protein is ready to begin. The P site is the host for the tRNA carrying the growing protein, while the A site is where the tRNA carrying the next amino acid sits. Think of the A site as the on-deck circle of a baseball field, and the p site as the batter’s box. So, AUG is the first codon bound, and in the P site is the tRNA carrying the methionine. The next codon in the sequence determines which tRNA binds next, and that tRNA molecule sits in the A site of the ribosome. An enzyme helps a peptide bond form (Figure 10.15 between the amino acid on the A site tRNA and the amino acid on the P site tRNA. After this happens, the amino acid from the P site moves to the A site, setting the stage for the tRNA in the P site to leave the ribosome. Now a step called translocation occurs (Figure 10.16. During this step, the ribosome moves along the mRNA in such a way that the A site becomes the P site and the next tRNA comes into the new A site carrying the next amino acid. This process continues until the stop codon is reached (Figure 10.17, causing the completed protein to leave the ribosome.
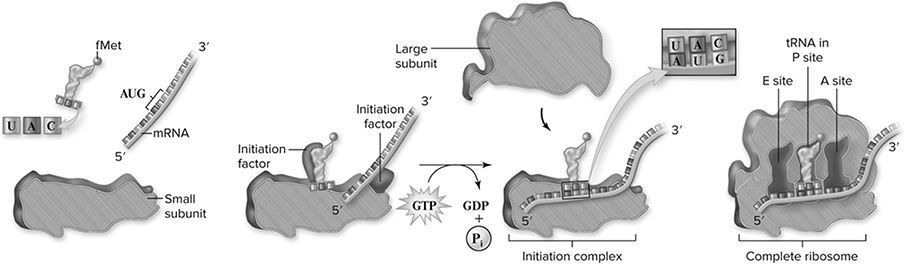
Figure 10.14 Initiation of translation. In prokaryotes, initiation factors play key roles in positioning the small ribosomal subunit, the initiator tRNAfMet, and the mRNA. When the tRNAfMet is positioned over the first AUG codon of the mRNA, the large ribosomal subunit binds, forming the E, P, and A sites where successive tRNA molecules bind to the ribosomes, and polypeptide synthesis begins. Ribosomal subunits are shown as a cutaway sectioned through the middle. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
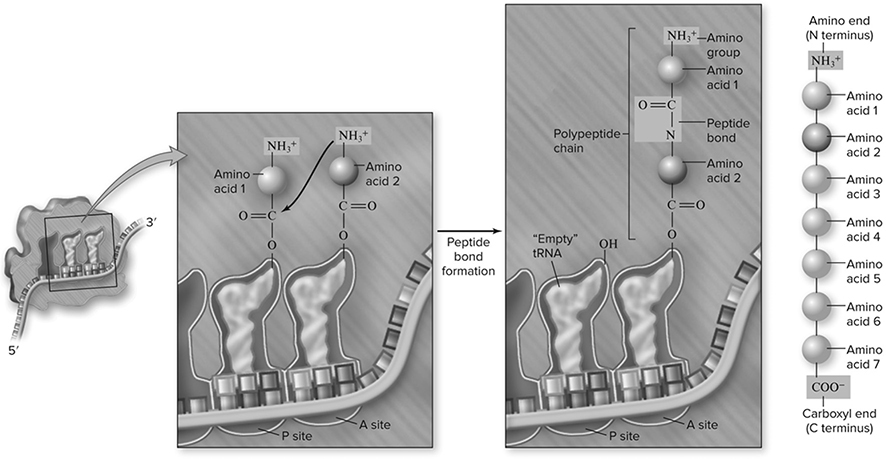
Figure 10.15 Peptide bond formation. Peptide bonds are formed between a “new” charged tRNA in the A site and the growing chain attached to the tRNA in the P site. The bond forms between the amino group of the new amino acid and the carboxyl group of the growing chain. This breaks the bond between the growing chain and its tRNA, transferring it to the A site as the new amino acid remains attached to its tRNA. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
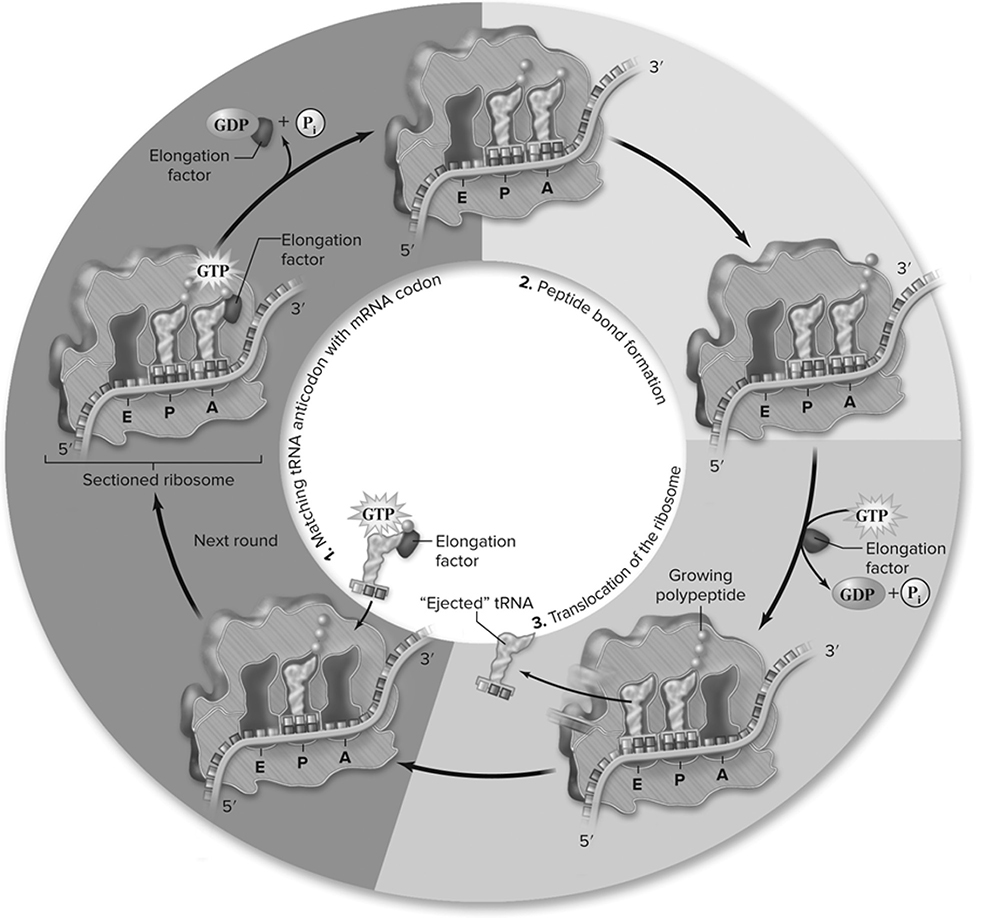
Figure 10.16 Elongation cycle. Numbering of the cycle corresponds to the numbering in the text. The cycle begins when a new charged tRNA with anticodon matching the codon of the mRNA in the A site arrives with EF-Tu. The EF-Tu hydrolyzes GTP and dissociates from the ribosome. A peptide bond is formed between the amino acid in the A site and the growing chain in the P site, transferring the growing chain to the A site, and leaving the tRNA in the P site empty. Ribosome translocation requires another elongation factor and GTP hydrolysis. This moves the tRNA in the A site into the P site, the next codon in the mRNA into the A site, and the empty tRNA into the E site. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
Gene Regulation
Let’s cover some vocabulary before diving into this section:
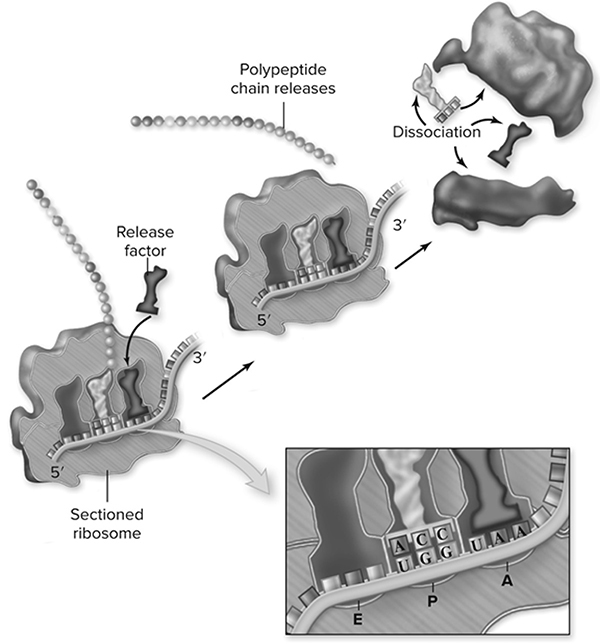
Figure 10.17 Termination of protein synthesis. There is no tRNA with an anticodon complementary to any of the three termination signal codons. When a ribosome encounters a termination codon, it stops translocating. A specific protein release factor facilitates the release of the polypeptide chain by breaking the covalent bond that links the polypeptide to the P site tRNA. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
IST-2
Differences in the expression of genes account for some of the phenotypic differences between organisms.
Promoter region: a base sequence that signals the start site for gene transcription; this is where RNA polymerase binds to begin the process.
Operator: a short sequence near the promoter that assists in transcription by interacting with regulatory proteins (transcription factors).
Operon: a promoter/operator pair that services multiple genes; the lac operon is a well-known example (Figure 10.18.
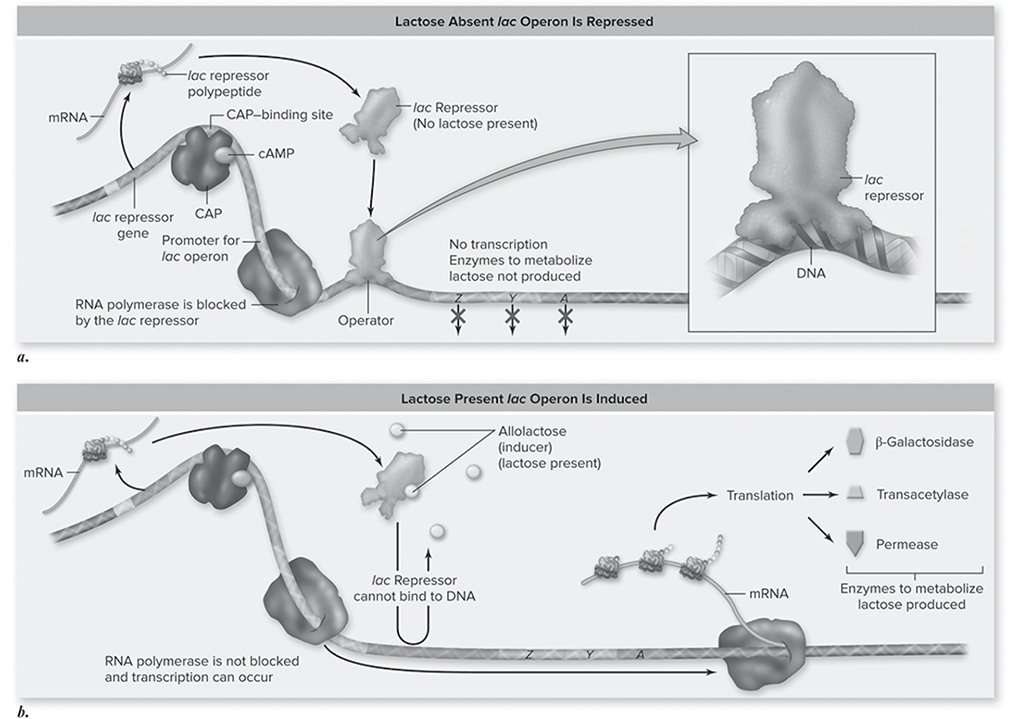
Figure 10.18 Induction of the lac operon. a. In the absence of lactose the lac repressor binds to DNA at the operator site, thus preventing transcription of the operon. When the repressor protein is bound to the operator site, the lac operon is shut down (repressed). b. The lac operon is transcribed (induced) when CAP is bound and when the repressor is not bound. Allolactose binding to the repressor alters the repressor’s shape so it cannot bind to the operator site and block RNA polymerase activity. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
Repressor: protein that prevents the binding of RNA polymerase to the promoter site.
CT teacher:“Be able to write about operons.”
Enhancer: DNA region, also known as a “regulator,” that is located thousands of bases away from the promoter; it influences transcription by interacting with specific transcription factors.
Inducer: a molecule that binds to and inactivates a repressor (e.g., lactose for the lac operon).
Prokaryotic Gene Regulation
The control of gene expression is vital to the proper and efficient functioning of an organism. In bacteria, operons are a major method of gene expression control. The lactose operon services a series of three genes involved in the process of lactose metabolism. This contains the genes that help the bacteria digest lactose. It makes sense for bacteria to produce these genes only if lactose is present. Otherwise, why waste the energy on unneeded enzymes? This is where operons come into play—in the absence of lactose, a repressor binds to the promoter region and prevents transcription from occurring. When lactose is present, there is a binding site on the repressor where lactose attaches, causing the repressor to let go of the promoter region. RNA polymerase is then free to bind to that site and initiate transcription of the genes. When the lactose is gone, the repressor again becomes free to bind to the promoter, halting the process.
Eukaryotic Gene Regulation
Because gene expression in eukaryotes involves more steps, there are more places where gene control can occur. Here are the different ways that gene expression occurs in eukaryotes (Figures 10.19—10.21):
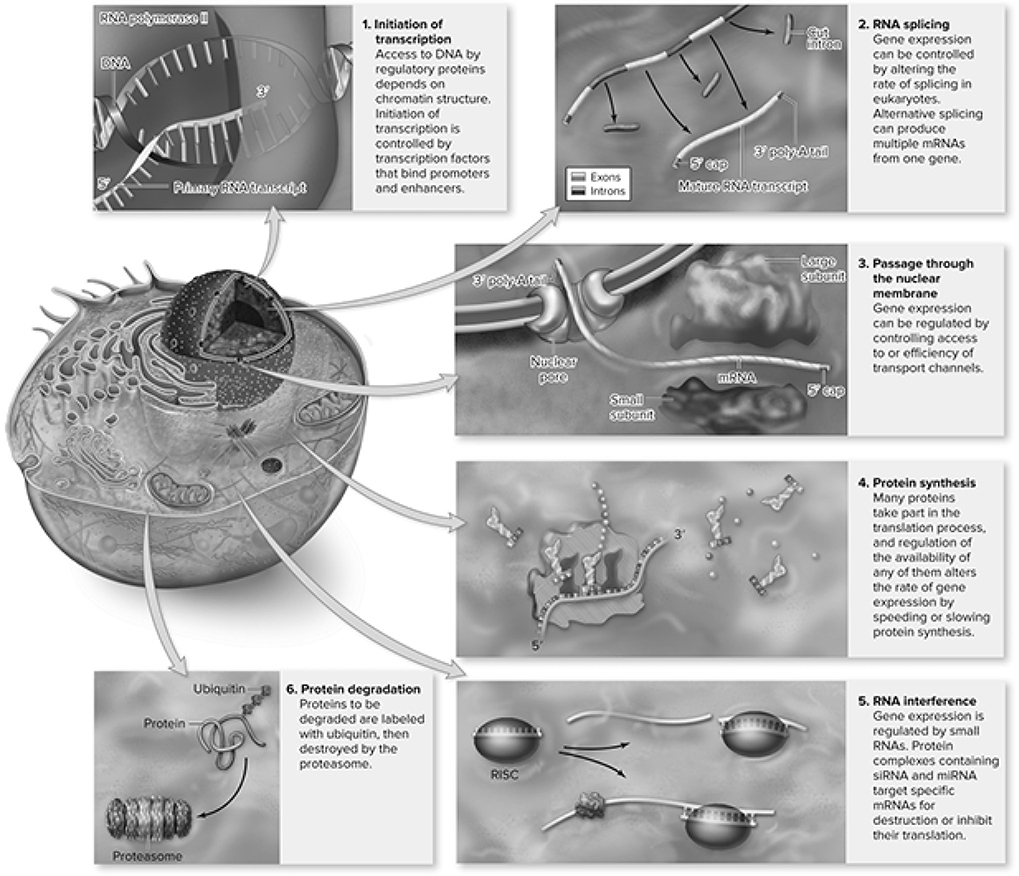
Figure 10.19 Mechanisms for control of gene expression in eukaryotes. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
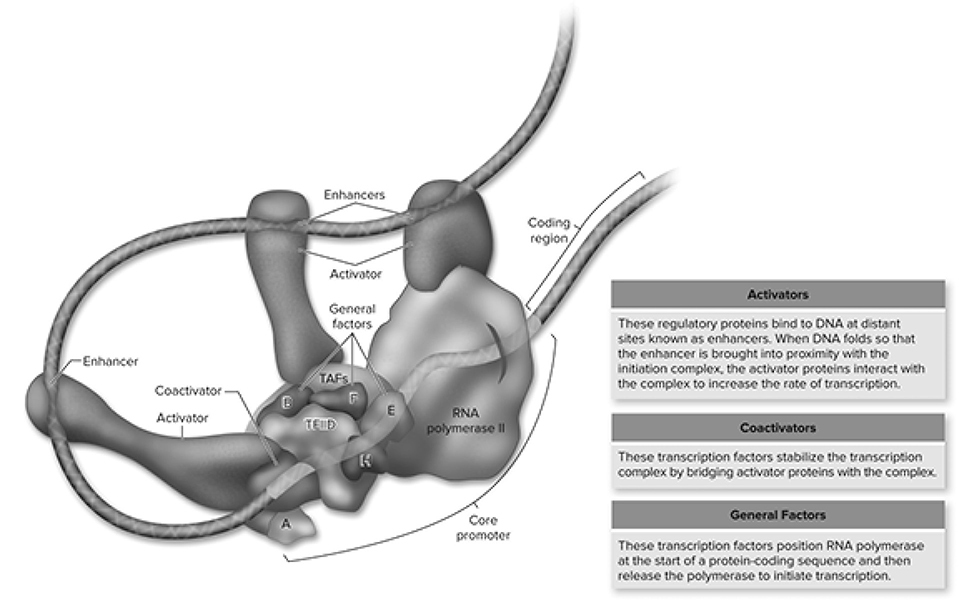
Figure 10.20 Interactions of various factors within the transcription complex. All specific transcription factors bind to enhancer sequences that may be distant from the promoter. These proteins can then interact with the initiation complex by DNA looping to bring the factors into proximity with the initiation complex. As detailed in the text, some transcription factors, called activators, can directly interact with the RNA polymerase II or the initiation complex, whereas others require additional coactivators. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
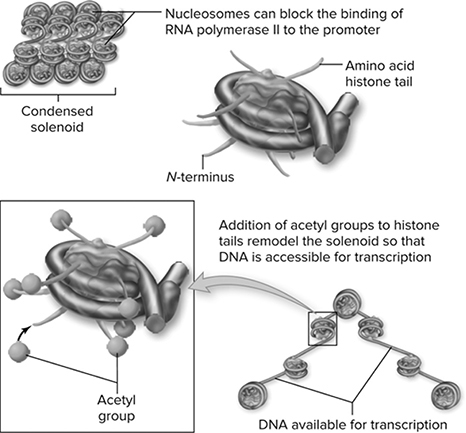
Figure 10.21 Histone modification affects chromatin structure. DNA in eukaryotes is organized first into nucleosomes and then into higher-order chromatin structures. The histones that make up the nucleosome core have amino tails that protrude. These amino tails can be modified by the addition of acetyl groups. The acetylation alters the structure of chromatin, making it accessible to the transcription apparatus. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
Cell Specialization
The differences in the expression of genes controlled by the cell lead to different phenotypes and cells in an organism. Transcription factors play a vital role in the regulation of transcription, determining which genes are active in each cell of your body. These factors are needed to assemble a transcription apparatus at the promoter region with RNA polymerase during transcription and can enhance the sequence leading to the transcription of a particular gene. While some transcription factors act as enhancers, others act as repressors by binding to the DNA and blocking transcription.
The regulation of eukaryotic gene expression leading to cell specialization continues with the packaging of the DNA into chromatin. The methylation of DNA bases in the chromatin correlates with genes being “turned off” while the acetylation of DNA bases in the chromatin correlates with the genes being “turned on.”
Cell Differentiation
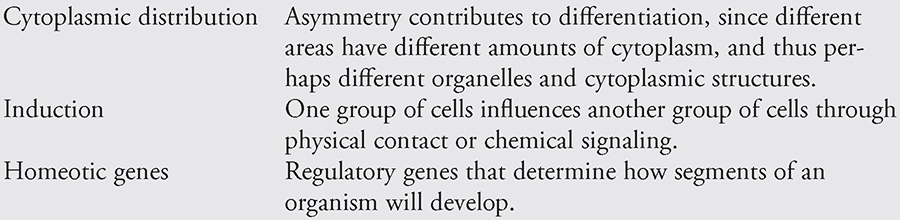
How do the various cells of the developing embryo differentiate into cells with different functions if they come from the same parent cell? As mentioned earlier, not every cell receives the same amount of cytoplasm during the cleavage divisions. It is thought that this asymmetric distribution of cytoplasm plays a role in the differentiation of the daughter cells. Cells containing different organelles or other cytoplasmic components are able to perform different functions. Two other factors, induction and homeotic genes, contribute to cellular differentiation.
Induction is the influence of one group of cells on the development of another through physical contact or chemical signaling. Just in case you are asked to write an essay on induction, it is good to know a bit about the experiments of the German embryologist Hans Spemann. His experiments revealed that the notochord induces cells of the dorsal ectoderm to develop into the neural plate. When cells from the notochord of an embryo are transplanted to a different place near the ectoderm, the neural plate will develop in the new location. The cells from the notochord region act as “project directors,” telling the ectoderm where to produce the neural tube and central nervous system.
Homeotic genes regulate or “direct” the body plan of organisms. For example, a fly’s homeotic genes help determine how its segments will develop and which appendages should grow from each segment. Scientists interfering with the development of these poor creatures have found that mutations in these genes can lead to the production of too many wings, legs in the wrong place, and other unfortunate abnormalities. The DNA sequence of a homeotic gene that tells the cell where to put things is called the homeobox. It is similar from organism to organism and has been found to exist in a variety of organisms—birds, humans, fish, and frogs.

Factors in Cellular Differentiation

Mutations
A mutation is a heritable change in the genes of an organism. These heritable changes can result in changes to the phenotype of an organism or be silent and not affect the phenotype of an organism. These alterations of the DNA sequences in organisms contribute to variation in a population and can be subject to natural selection.
IST-2
Differences in the expression of genes account for some of the phenotypic differences between organisms.
Point mutations, which alter a single base, can be a substitution of another base, a deletion of a base(s) or an addition of a base(s). It is important that you can -identify the different types of mutations. The table and figure below show each type of -mutation.
Here is a short list of mutation types (Figure 10.22 that you should know:
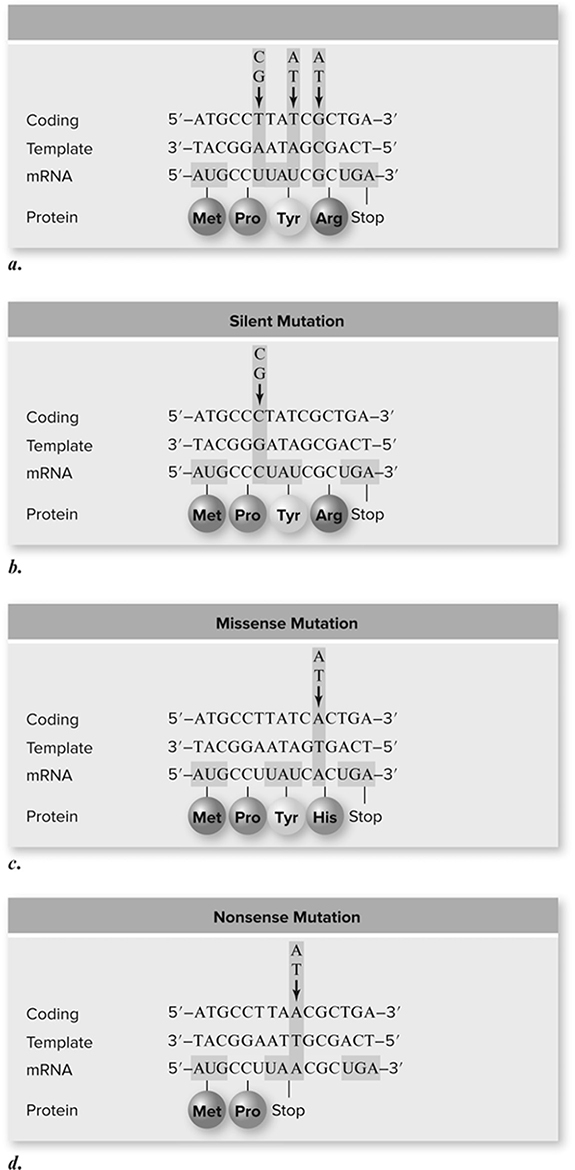
Figure 10.22 Types of point mutations. a. A hypothetical gene is shown with encoded mRNA and protein. Arrows above the gene indicate sites of mutations described in the rest of the figure. b. Silent mutation. Due to degeneracy in the genetic code, base substitution does not always change an amino acid. This usually involves the third position of a codon, in this case a T/A to C/G mutation. c. Missense mutation. The G/C to A/T mutation changes the amino acid encoded from arginine to histidine. d. Nonsense mutation. The T/A to A/T mutation produces a UAA stop codon in the mRNA. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
1. Frameshift mutations. Deletion or addition of DNA nucleotides that does not add or remove a multiple of three nucleotides. mRNA is produced on a DNA template and is read in bunches of three called codons, which tell the protein synthesis machinery which amino acid to add to the growing protein chain. If the mRNA reads: THE FAT CAT ATE HER HAT, and the F is removed because of an error somewhere, the frame has now shifted to read THE ATC ATA THE ERH AT . . . (gibberish). This kind of mutation usually produces a nonfunctional protein unless it occurs late in protein production.
2. Missense mutation. Substitution of the wrong nucleotides into the DNA sequence. These substitutions still result in the addition of amino acids to the growing protein chain during translation, but they can sometimes lead to the addition of incorrect amino acids to the chain. It could cause no problem at all, or it could cause a big problem as in sickle cell anemia, in which a single amino acid error caused by a substitution mutation leads to a disease that wreaks havoc on the body as a whole.
IST-4
The processing of genetic information is imperfect and is a source of genetic variation.
3. Nonsense mutation. Substitution of the wrong nucleotides into the DNA sequence. These substitutions lead to premature stoppage of protein synthesis by the early placement of a stop codon, which tells the protein synthesis machinery to grind to a halt. The stop codons are UAA, UAG, and UGA. This type of mutation usually leads to a nonfunctional protein.
4. Thymine dimers. Result of too much exposure to UV (ultraviolet) light. Thymine nucleotides located adjacent to one another on the DNA strand bind together when this exposure occurs. This can negatively affect replication of DNA and help cause further mutations.
Sickle cell anemia represents a prime example of a point mutation leading to a phenotypic change in humans. Sickle cell is caused by a mutation of the fourth codon in the gene for hemoglobin. The substitution of glycine (a polar amino acid) to valine (a nonpolar amino acid) causes the shape of the hemoglobin protein to change, distorting the shape of the red blood cells. Figure 10.23 shows the mutation leading to the altered protein.
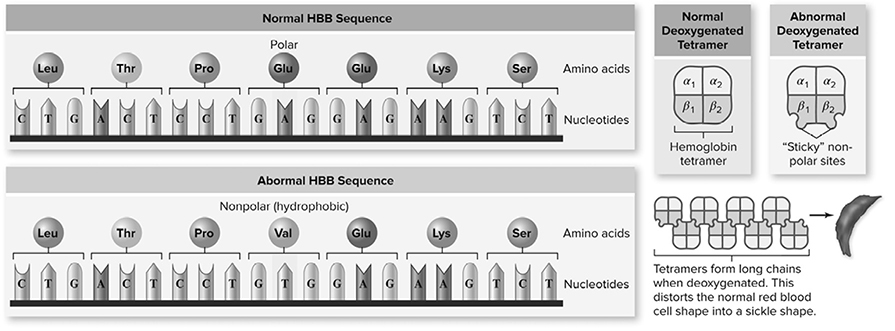
Figure 10.23 Sickle cell anemia is caused by an altered protein. Hemoglobin is composed of a tetramer of two α-globin and two β-globin chains. The sickle cell allele of the β-globin gene contains a single base change resulting in the substitution of Val for Glu. This creates a hydrophobic region on the surface of the protein that is “sticky,” leading to their association into long chains that distort the shape of the red blood cells. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
Chromosomal mutations can change the structure of chromosomes and lead to many -different disorders found in humans. The following figure shows the four types of chromosomal mutations: deletion, duplication, inversion, and reciprocal translocation (Figure 10.24.
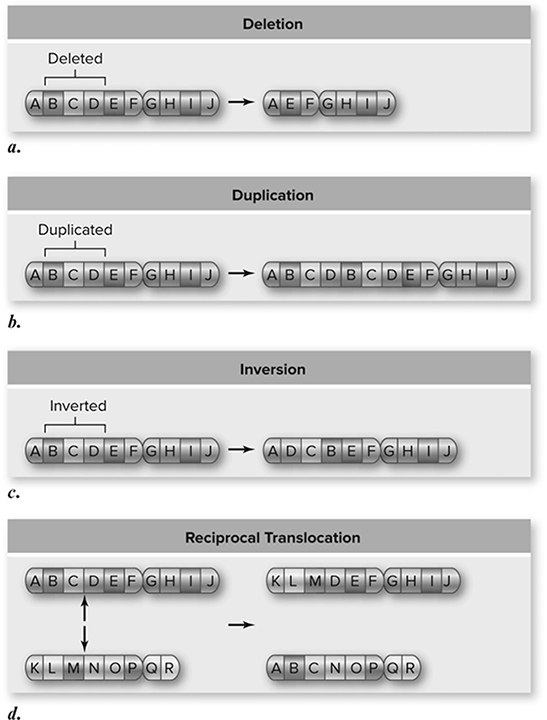
Figure 10.24 Chromosomal mutations. Larger-scale changes in chromosomes are also possible. Material can be deleted (a), duplicated (b), and inverted (c). Translocations occur when one chromosome is broken and becomes part of another chromosome. This often occurs where both chromosomes are broken and exchange material, an event called a reciprocal translocation (d). (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
The Genetics of Viruses
A virus is a parasitic infectious agent that is unable to survive outside of a host organism. Viruses do not contain enzymes for metabolism, and they do not contain ribosomes for protein synthesis. They are completely dependent on their host. Once a virus infects a cell, it takes over the cell’s machinery and uses it to produce whatever it needs to survive and reproduce. How a virus acts after it enters a cell depends on what type of virus it is. Classification of viruses is based on many factors:
Genetic material: DNA, RNA, protein, etc.?
Capsid: type of capsid?
Viral envelope: present or absent?
Host range: what type of cells does it affect?
All viruses have a genome (DNA or RNA) and a protein coat (capsid). A capsid is a protein shell that surrounds the genetic material. Some viruses are surrounded by a structure called a viral envelope, which not only protects the virus but also helps the virus attach to the cells that it prefers to infect. The viral envelope is produced in the endoplasmic reticulum (ER) of the infected cell and contains some elements from the host cell and some from the virus. Each virus has a host range, which is the range of cells that the virus is able to infect. For example, HIV infects the T cells of our body, and bacteriophages infect only bacteria.
A special type of virus that merits discussion is one called a retrovirus. This is an RNA virus that carries an enzyme called reverse transcriptase. Once in the cytoplasm of the cell, the RNA virus uses this enzyme and “reverse transcribes” its genetic information from RNA into DNA, which then enters the nucleus of the cell. In the nucleus, the newly transcribed DNA incorporates into the host DNA and is transcribed into RNA when the host cell undergoes normal transcription. The mRNA produced from this process gives rise to new retrovirus offspring, which can then leave the cell in a lytic pathway. A well-known example of a retrovirus is the HIV virus of AIDS.
Once inside the cell, a DNA virus can take one of two pathways—a lytic or a lysogenic pathway. In a lytic cycle, the cell actually produces many viral offspring, which are released from the cell—killing the host cell in the process. In a lysogenic cycle, the virus falls dormant and incorporates its DNA into the host DNA as an entity called a provirus. The viral DNA is quietly reproduced by the cell every time the cell reproduces itself, and this allows the virus to stay alive from generation to generation without killing the host cell. Viruses in the lysogenic cycle can sometimes separate out from the host DNA and enter the lytic cycle (like a bear awaking from hibernation).
Viruses come in many shapes and sizes. Although many viruses are large, viroids are plant viruses that are only a few hundred nucleotides in length, showing that size is not the only factor in viral success. Another type of infectious agent you should be familiar with is a prion—an incorrectly folded form of a brain cell protein that works its magic by converting other normal host proteins into misshapen proteins. An example of a prion disease that has been getting plenty of press coverage is “mad cow” disease. Prion diseases are degenerative diseases that tend to cause brain dysfunction—dementia, muscular control problems, and loss of balance.
The Genetics of Bacteria
Bacteria are prokaryotic cells that consist of one double-stranded circular DNA molecule. Present in the cells of many bacteria are extra circles of DNA called plasmids, which contain just a few genes and have been useful in genetic engineering. Plasmids replicate independently of the main chromosome. Bacterial cells reproduce in an asexual fashion, undergoing binary fission. Quite simply, the cell replicates its DNA and then physically pinches in half, producing a daughter cell that is identical to the parent cell. From this description of binary fission, it seems unlikely that there could be variation among bacterial cells. This is not the case, thanks to mutation and genetic recombination. As in humans, DNA mutation in bacteria occurs very rarely, but some bacteria replicate so quickly that these mutations can have a pronounced effect on their variability.
Transformation
An experiment performed by Griffith in 1928 provides a fantastic example of transformation—the uptake of foreign DNA from the surrounding environment. Transformation occurs through the use of proteins on the surface of cells that snag pieces of DNA from around the cell that are from closely related species. This particular experiment involved a bacteria known as Streptococcus pneumoniae, which existed as either a rough strain (R), which is nonvirulent, or as a smooth strain (S), which is virulent. A virulent strain is one that can lead to contraction of an illness. The experimenters exposed mice to different forms of the bacteria. Mice given live S bacteria died. Mice given live R bacteria survived. Mice given heat-killed S bacteria survived. Mice given heat-killed S bacteria combined with live R bacteria died. This was the kicker . . . all the other results to this point were expected. Those exposed to heat-killed S combined with live R bacteria contracted the disease because the live R bacteria underwent transformation. Some of the R bacteria picked up the portion of the heat-killed S bacteria’s DNA, which contained the instructions on how to make the vital component necessary for successful disease transmission. These R bacteria became virulent.
Transduction
To understand transduction, you first need to be introduced to something called a phage a virus that infects bacteria. The mechanism by which a phage (otherwise known as bacteriophage) infects a cell reminds me of a syringe. A phage contains within its capsid the DNA that it is attempting to deliver. A phage latches onto the surface of a cell and, like a syringe, fires its DNA through the membrane and into the cell. Transduction is the movement of genes from one cell to another by phages. The two main forms of transduction you should be familiar with are generalized and specialized transduction.
Generalized Transduction Imagine that a phage virus infects and takes over a bacterial cell that contains a functional gene for resistance to penicillin. Occasionally during the creation of new phage viruses, pieces of host DNA instead of viral DNA are accidentally put into a phage. When the cell lyses, expelling the newly formed viral particles, the phage containing the host DNA may latch onto another cell, injecting the host DNA from one cell into another bacterial cell. If the phage attaches to a cell that contains a nonfunctional gene for resistance to penicillin, the effects of this transduction process can be observed. After injecting the host DNA containing the functional penicillin resistance gene, crossover could occur between the comparable gene regions, switching the nonfunctional gene with the functional gene. This would create a new cell that is resistant to penicillin.
Specialized Transduction This type of transduction involves a virus that is in the lysogenic cycle, resting quietly along with the other DNA of the host cell. Occasionally when a lysogenic virus switches cycles and becomes lytic, it may bring with it a piece of the host DNA as it pulls out of the host chromosome. Imagine that the host DNA it brought with it contains a functional gene for resistance to penicillin. This virus, now in the lytic cycle, will produce numerous copies of new viral offspring that contain this resistance gene from the host cell. If the new phage offspring attaches to a cell that is not penicillin resistant and injects its DNA and crossover occurs, specialized transduction will have occurred.
Conjugation
This is the raciest of the genetic recombinations that we will cover . . . the bacterial version of sex. It is the transfer of DNA between two bacterial cells connected by appendages called sex pili. Movement of DNA between two cells occurs across a cytoplasmic connection between the two cells and requires the presence of an F-plasmid, which contains the genes necessary for the production of a sex pilus.
Biotechnology
DNA technology is advancing at a rapid rate, and you need to have a basic understanding of the most common laboratory techniques for the AP Biology exam.
IST-1
Heritable information provides for continuity of life.
Restriction enzymes are enzymes that cut DNA at specific nucleotide sequences. When added to a solution containing DNA, the enzymes cut the DNA wherever the enzyme’s particular sequence appears. This creates DNA fragments with single-stranded ends called “sticky-ends,” which find and reconnect with other DNA fragments containing the same ends (with the assistance of DNA ligase). Sticky ends allow DNA pieces from different sources to be connected, creating recombinant DNA. Another concept important to genetic engineering is the vector, which moves DNA from one source to another. Plasmids can be removed from bacterial cells and used as vectors by cutting the DNA of interest and the DNA of the plasmid with the same restriction enzyme to create DNA with similar sticky ends. The DNA can be attached to the plasmid, creating a vector that can be used to transport DNA.
Gel Electrophoresis
This technique is used to separate and examine DNA fragments (Figure 10.25. The DNA is cut with our new friends, the restriction enzymes, and then separated by electrophoresis. The pieces of DNA are separated on the basis of size with the help of an electric charge. DNA is added to the wells at the negative end of the gel. When the electric current is turned on, the migration begins. Smaller pieces travel farther along the gel, and larger pieces do not travel as far. The bigger you are, the harder it is to move. This technique can be used to sequence DNA and determine the order in which the nucleotides appear. It can be used in a procedure known as Southern blotting (after Edwin M. Southern, a British biologist) to determine if a particular sequence of nucleotides is present in a sample of DNA. Electrophoresis is used in forensics to match DNA found at the crime scene with DNA of suspects. This requires the use of pieces of DNA called restriction fragment length polymorphisms (RFLPs). DNA is specific to each individual, and when it is mixed with restriction enzymes, different combinations of RFLPs will be obtained from person to person. Electrophoresis separates DNA samples from the suspect and whatever sample is found at the scene of the crime. The two are compared, and if the RFLPs match, there is a high degree of certainty that the DNA sample came from the suspect.
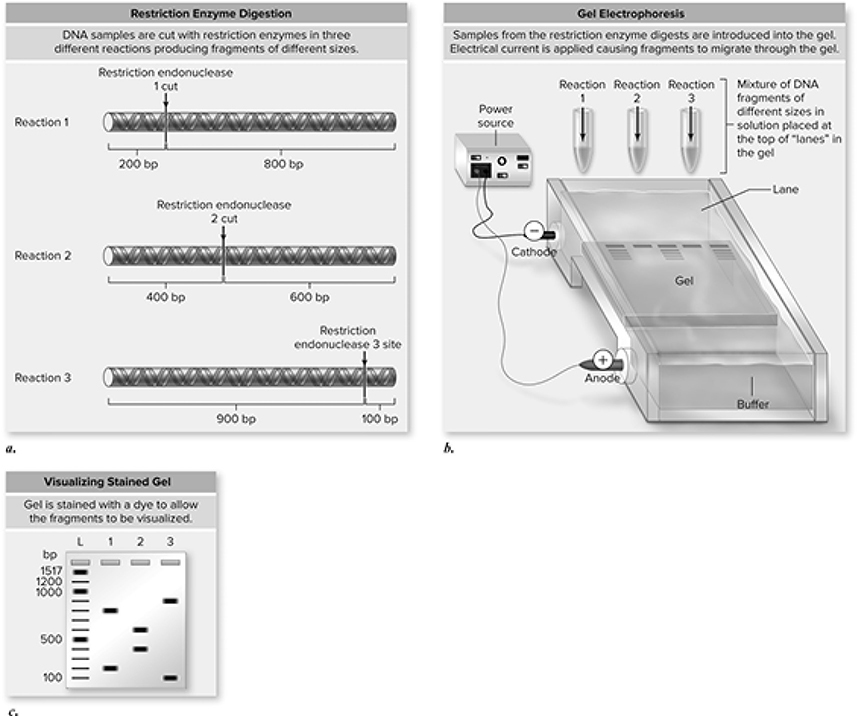
Figure 10.25 Gel electrophoresis separates DNA fragments based on size. a. Three restriction enzymes are used to cut DNA into specific pieces, depending on each enzyme’s recognition sequence. b. The fragments are loaded into a gel (agarose or polyacrylamide), and an electrical current is applied. The DNA fragments migrate through the gel based on size, with larger fragments moving more slowly. c. This results in a pattern of fragments separated based on size, with the smaller fragments migrating farther than larger ones. A series of fragments of known sizes produces a ladder so that sizes of fragments of unkown size can be estimated (bp = base-pairs; L = ladder; 1, 2, 3 = fragments from piece of DNA cut with restriction endonucleases 1, 2, and 3, respectively). (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
Cloning
Sometimes it is desirable to obtain large quantities of a gene of interest, such as insulin for the treatment of diabetes. The process of cloning involves many of the steps we just -mentioned. Plasmids used for cloning often contain two important genes—one that provides resistance to an antibiotic, and one that gives the bacteria the ability to metabolize some sugar. In this case, we will use a galactose hydrolyzing gene and a gene for ampicillin resistance. The plasmid and DNA of interest are both cut with the same restriction enzyme. The restriction site for this enzyme is right in the middle of the galactose gene of the plasmid. When the sticky ends are created, the DNA of interest and the plasmid molecules are mixed and join together. Not every combination made here is what the scientist is looking for. The recombinant plasmids produced are transformed into bacterial cells. This is where the two specific genes for the plasmid come into play. The transformed cells are allowed to reproduce and are placed on a medium containing ampicillin. Cells that have taken in the ampicillin resistance gene will survive, while those that have not will perish. The medium also contains a special sugar that is broken down by the galactose enzyme present in the vector to form a colored product. The cells containing the gene of interest will remain white since the galactose gene has been interrupted and rendered nonfunctional. This allows the experimenter to isolate cells that contain the desired product. Now, it is time for us to quit cloning around and move onto another genetic engineering technique.
Polymerase Chain Reaction
Think of this technique as a high-speed copy machine. It is used to produce large quantities of a particular sequence of DNA in a very short amount of time. If the cloning reaction is the 747 of copying DNA, then polymerase chain reaction (PCR) is the Concorde. This process begins with double-stranded DNA containing the gene of interest. DNA polymerase, the superstar enzyme of DNA replication, is added to the mixture along with a huge number of nucleotides and primers specific for the sequence of interest, which help initiate the synthesis of DNA. PCR begins by heating the DNA to split the strands, followed by the cooling of the strands to allow the primers to bind to the sequence of interest. DNA polymerase then steps up to the plate and produces the rest of the DNA molecule by adding the nucleotides to the growing DNA strand. Each cycle concludes having doubled the amount of DNA present at the beginning of the cycle. The cycle is repeated over and over, every few minutes, until a huge amount of DNA has been created. PCR is used in many ways, such as to detect the presence of viruses like HIV in cells, diagnose genetic disorders, and amplify trace amounts of DNA found at crime scenes (Figure 10.26a and b).
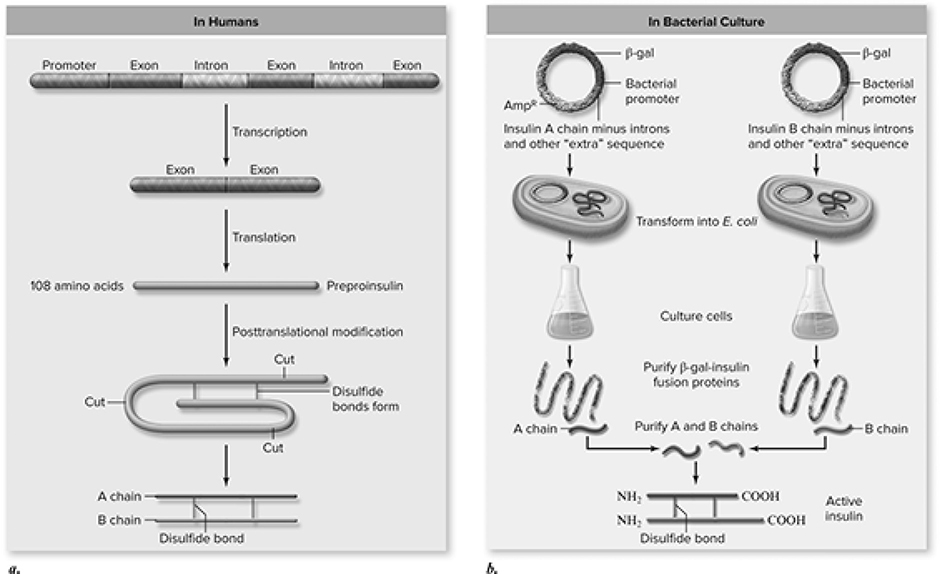
Figure 10.26 Making insulin in genetically engineered E. coli. a. In human cells one preproinsulin polypeptide is processed posttranslationally into insulin chains A and B that associate via disulfide bonds to form mature insulin. b. Two cDNAs corresponding to the gene sequences for insulin chain A and chain B are cloned into plasmids and introduced into different E. coli. Cultures of the two types of E. coli express either chain A or chain B. Chains A and B are purified from the different E. coli and, when mixed, associate into active mature insulin. (Reproduced with permission from Raven P, Johnson G, Mason K, Losos J, Duncan T; Biology, 12th ed. New York: McGraw Hill; 2020)
DNA Sequencing
In 2003, the international science community completed sequencing of the human genome. Using cutting edge techniques, scientists were able to determine the sequence of nucleotide bases for a human’s DNA. This discovery has led to new techniques and technologies that allow for the sequencing of small pieces of DNA to entire genomes of organisms.
![]() Review Questions
Review Questions
1. Which of the following statements is incorrect?
A. Messenger RNA must be processed before it can leave the nucleus of a eukaryotic cell.
B. A virus in the lysogenic cycle does not kill its host cell, whereas a virus in the lytic cycle destroys its host cell.
C. DNA polymerase is restricted in that it can add nucleotides only in a 5′-to-3′ direction.
D. During translation, the A site holds the tRNA carrying the growing protein, while the P site holds the tRNA carrying the next amino acid.
2. The process of transcription results in the formation of
A. DNA.
B. proteins.
C. lipids.
D. RNA.
3. Which of the following codons signals the beginning of the translation process?
A. AGU
B. UGA
C. AUG
D. AGG
4. Which of the following is an improper pairing of DNA or RNA nucleotides?
A. Thymine-adenine
B. Guanine-thymine
C. Uracil-adenine
D. Guanine-cytosine
5. Which of the following is responsible for the type of diseases that includes “mad cow” disease?
A. Viroids
B. Plasmids
C. Prions
D. Provirus
6. Which of the following is the correct sequence of events that must occur for trans-lation to begin?
A. Transfer RNA binds to the small ribosomal subunit, which leads to the attachment of the large ribosomal subunit. This signals to the mRNA molecule that it should now bind, with its first codon in the correct site, to the protein synthesis machinery, and translation begins.
B. Messenger RNA attaches to the small ribosomal subunit, with its first codon in the correct site, thus attracting a tRNA molecule to attach to the codon. This signals to the large subunit that it should now bind to the protein synthesis machinery, and translation can begin.
C. Messenger RNA attaches to the large ribosomal subunit with its first codon in the correct site, attracting a tRNA molecule to attach to the codon. This signals to the small subunit that it should now bind to the protein synthesis machinery, and translation can begin.
D. Transfer RNA binds to the large ribosomal subunit, which leads to the attachment of the small ribosomal subunit. This signals to the mRNA molecule that it should now bind with its first codon in the correct site to the protein synthesis machinery, and translation begins.
7. All the following are players involved in the control of gene expression except
A. episomes.
B. repressors.
C. operons.
D. methylation.
8. Which of the following does not occur during RNA processing in the nucleus of eukaryotes?
A. The removal of introns from the RNA molecule
B. The addition of a string of adenine nucleotides to the 3′ end of the RNA molecule
C. The addition of a guanine cap to the 5′ end of the RNA molecule
D. The addition of methyl groups to certain nucleotides of the RNA molecules
9. Which of the following statements is not true of a tRNA molecule?
A. The job of transfer RNA is to carry amino acids to the ribosomes.
B. At the attachment site of each tRNA, there is a region called the anticodon, which is a three-nucleotide sequence that is perfectly complementary to a particular codon.
C. Each tRNA molecule has a short lifespan and is used only once during translation.
D. The enzyme responsible for ensuring that a tRNA molecule is carrying the appropriate amino acid is aminoacyl tRNA synthase.
For questions 10 and 11, please use the following gel:
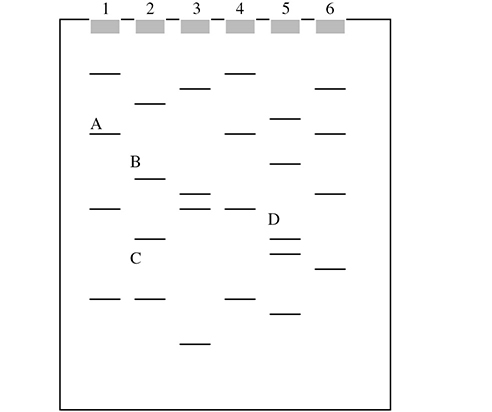
10. Which of the DNA pieces in the gel is smallest in size?
A. A
B. B
C. C
D. D
11. If well 1 is DNA from a crime scene, which individual should contact a lawyer?
A. Person 2
B. Person 3
C. Person 4
D. Person 5
![]() Answers and Explanations
Answers and Explanations
1. D—During translation, the P site holds the tRNA carrying the growing protein, while the A site holds the tRNA carrying the next amino acid. When translation begins, the first codon bound is the AUG codon, and in the P site is the tRNA with the methionine. The next codon in the sequence determines which tRNA binds next, and the appropriate tRNA molecule sits in the A site of the ribosome. A peptide bond forms between the amino acid on the A site tRNA and the amino acid on the P site tRNA. The amino acid from the P site then moves to the A site, allowing the tRNA in the P site to leave the ribosome. Next, the ribosome moves along the mRNA in such a way that the A site is now the P site and the next tRNA comes into the A site carrying the next amino acid. Answer choices A, B, and C are all true.
2. D—The process of transcription leads to the production of RNA. RNA is not immediately ready to leave the nucleus after it is produced. It must first be processed, during which a 3′ poly-A tail and a 5′ cap are added and the introns are spliced from the RNA molecule. After this process, the RNA is free to leave the nucleus and lead the production of proteins.
3. C—AGG codes for the amino acid arginine. AGU codes for the amino acid serine. UGA and UAG are stop codons, which signal the end of the translation process. AUG is the start codon, which also codes for methionine.
4. B—Guanine does not pair with thymine in DNA or RNA. Watson and Crick discovered that adenine pairs with thymine (A=T) held together by two hydrogen bonds and guanine pairs with cytosine (C≡G) held together by three hydrogen bonds. One way that RNA differs from DNA is that it contains uracil instead of thymine. But in RNA, guanine still pairs with cytosine and adenine instead pairs with uracil. Watson and Crick also discovered that for the structure of DNA they discovered to be true, a purine must always be paired with a pyrimidine. Adenine and guanine are the purines, and thymine and cytosine are the pyrimidines.
5. C—Prions are the culprit for mad cow disease. Viroids are tiny viruses that infect plants. Plasmids are small circles of DNA in bacteria that are separate from the main chromosome. They are self-replicating and are vital to the process of genetic engineering. A provirus is that which is formed during the lysogenic cycle of a virus when it falls dormant and incorporates its DNA into the host DNA. A retrovirus is an RNA virus that carries an enzyme called reverse transcriptase. A classic example of a retrovirus is HIV.
6. B—Translation begins when the mRNA attaches to the small ribosomal subunit. The first codon for this process is always AUG. This attracts a tRNA molecule carrying methionine to attach to the AUG codon. When this occurs, the large subunit of the ribosome, containing the A site and the P site, binds to the complex. The elongation of the protein is ready to begin after the complex has been properly constructed. Answers A, C, and D are all in the incorrect order.
7. A—Episomes are not involved in gene expression regulation. Episomes are plasmids that can be incor-porated into a bacterial chromosome. Repressors are regulatory proteins involved in gene regulation. They work by preventing transcription by binding to the promoter region. Operons are a promoter-operator pair that controls a group of genes, such as the lac operon. Methylation is involved in gene regulation. Barr bodies, discussed in Chapter 9, are found to contain a very high level of methylated DNA. Methyl groups have been associated with inactive DNA that does not undergo transcription. Hormones can affect transcription by acting directly on the transcription machinery in the nucleus of cells.
8. D—The mRNA produced after transcription must be modified before it can leave the nucleus and lead the translation of proteins in the ribosomes. Introns are cut out of the mRNA, and the remaining exons are ligated back together to produce the mRNA ready to be translated into a protein. Also, the 5′ end is given a guanine cap, which serves to protect the RNA and also helps the mRNA attach to the ribosome. The 3′ end is given the poly-A tail, which may help ease the movement from the nucleus to the cytoplasm. Methylation does not occur during posttranscriptional modification—it is a means of gene expression control.
9. C—tRNA does not have a short lifespan. Each tRNA molecule is released and re-cycled to bring more amino acids to the ribosomes to aid in translation. It is like a taxicab constantly picking up new passengers to deliver from place to place. Answer choices A, B, and D are all true.
10. C—Gel electrophoresis separates DNA fragments on the basis of size—the smaller you are, the farther you go. Because C went the farthest in this gel, this must be the smallest of the four selected DNA pieces. Of the four labeled, piece A must be the largest because it moved the least.
11. C—Person 4 should contact a lawyer. The DNA from the crime scene seems to match the DNA fingerprint from person 4. Electrophoresis is a very useful tool in forensics and can very accurately match DNA found at crime scenes with potential suspects.
![]() Rapid Review
Rapid Review
Briefly review the following terms:
DNA: contains A and G (purines), C and T (pyrimidines), arranged in a double helix of two strands held together by hydrogen bonds (A with T, and C with G).
RNA: contains A and G (purines), C and U (pyrimidines), single stranded. There are three types: mRNA (blueprints for proteins), tRNA (brings acids to ribosomes), and rRNA (make up ribosomes).
DNA replication: occurs during S-phase, semi-conservative, built in 5′-to-3′ direction. Helicase unzips the double strand, DNA polymerase comes in and adds on the nucleotides. Proofreading enzymes minimize errors of process.
Frameshift mutation: deletion or addition of nucleotides (not a multiple of 3); shifts reading frame.
Missense mutation: substitution of wrong nucleotide into DNA (e.g., sickle cell anemia); still produces a protein.
Nonsense mutation: substitution of wrong nucleotide into DNA that produces an early stop codon.
Transcription: process by which mRNA is synthesized on a DNA template.
RNA processing: introns (noncoding) are spliced out, exons (coding) glued together: 3′ poly-A tail, 5′ G cap.
Translation: process by which the mRNA specified sequence of amino acids is lined up on a ribosome for protein synthesis.
Codon: triplet of nucleotides that codes for a particular amino acid: start codon = AUG; stop codon = UGA, UAA, UAG. (For specifics on translation, please flip to text for a good description.)
Promoter: base sequence that signals start site for transcription.
Repressor: protein that prevents the binding of RNA polymerase to promoter site.
Inducer: molecule that binds to and inactivates a repressor.
Operator: short sequence near the promoter that assists in transcription by interacting with transcription factors.
Operon: on/off switch for transcription. Allows for production of genes only when needed. Remember the lac operon—lactose is the inducer, when present, transcription on; when absent, it is off.
Viruses: parasitic infectious agent unable to survive outside the host; can contain DNA or RNA, or have a viral envelope (protective coat).
✵ Lytic cycle: one in which the virus is actively reproducing and kills the host cell.
✵ Lysogenic cycle: one in which the virus lies dormant within the DNA of the host cell.
Retrovirus: RNA virus that carries with it reverse transcriptase (HIV).
Prion: virus that converts host brain proteins into misshapen proteins (mad cow disease).
Viroids: tiny plant viruses.
Phage: virus that infects bacteria.
Bacteria: prokaryotic cells; consist of one double-stranded circular DNA molecule; reproduce by binary fission (e.g., plasmid—extra circle of DNA present in bacteria that replicate independently of main chromosome).
Genetic Recombination
Transformation: uptake of foreign DNA from the surrounding environment (smooth vs. rough pneumococcus).
Transduction: movement of genes from one cell to another by phages, which are incorporated by crossover.
✵ Generalized: lytic cycle accidently places host DNA into a phage, which is brought to another cell.
✵ Specialized: virus leaving lysogenic cycle brings host DNA with it into phage.
Conjugation: transfer of DNA between two bacterial cells connected by sex pili.
Genetic Engineering
Restriction enzymes: enzymes that cut DNA at particular sequences, creating sticky ends.
Vector: mover of DNA from one source to another (plasmids are good vectors).
Cloning: somewhat slow process by which a desired sequence of DNA is copied numerous times.
Gel electrophoresis: technique used to separate DNA according to size (small = faster). DNA moves from: − to +.
Polymerase chain reaction (PCR): produces large quantities of sequence in short amount of time.